Systems Analysis of Drug-Induced Receptor Tyrosine Kinase Reprogramming Following Targeted Mono- and Combination Anti-Cancer Therapy
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Comprehensive Genome Methylation Analysis in Bladder Cancer: Identification and Validation of Novel Methylated Genes and Application of These As Urinary Tumor Markers
Published OnlineFirst July 25, 2011; DOI: 10.1158/1078-0432.CCR-10-2659 Clinical Cancer Human Cancer Biology Research Comprehensive Genome Methylation Analysis in Bladder Cancer: Identification and Validation of Novel Methylated Genes and Application of These as Urinary Tumor Markers Thomas Reinert1, Charlotte Modin1, Francisco M. Castano1, Philippe Lamy1, Tomasz K. Wojdacz4, Lise Lotte Hansen4, Carsten Wiuf3, Michael Borre2, Lars Dyrskjøt1, and Torben F. Ørntoft1 Abstract Purpose: Epigenetic alterations are common and can now be addressed in a parallel fashion. We investigated the methylation in bladder cancer with respect to location in genome, consistency, variation in metachronous tumors, impact on transcripts, chromosomal location, and usefulness as urinary markers. Experimental Design: A microarray assay was utilized to analyze methylation in 56 samples. Inde- pendent validation was conducted in 63 samples by a PCR-based method and bisulfite sequencing. The methylation levels in 174 urine specimens were quantified. Transcript levels were analyzed using expression microarrays and pathways were analyzed using dedicated software. Results: Global methylation patterns were established within and outside CpG islands. We validated methylation of the eight tumor markers genes ZNF154 (P < 0.0001), HOXA9 (P < 0.0001), POU4F2 (P < 0.0001), EOMES (P ¼ 0.0005), ACOT11 (P ¼ 0.0001), PCDHGA12 (P ¼ 0.0001), CA3 (P ¼ 0.0002), and PTGDR (P ¼ 0.0110), the candidate marker of disease progression TBX4 (P < 0.04), and other genes with stage-specific methylation. The methylation of metachronous tumors was stable and targeted to certain pathways. The correlation to expression was not stringent. Chromosome 21 showed most differential methylation (P < 0.0001) and specifically hypomethylation of keratins, which together with keratin-like proteins were epigenetically regulated. -

Prediction of Host-Virus Interaction Networks
Prediction of host-virus interaction networks Janet M. Doolittle A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Curriculum of Bioinformatics and Computational Biology. Chapel Hill 2012 Approved by: Tim Elston, PhD Shawn Gomez, Eng.Sc.D. Klaus Hahn, PhD Keith Burridge, PhD Anne Taylor, PhD Jennifer Webster-Cyriaque, PhD ©2012 Janet M. Doolittle ALL RIGHTS RESERVED ii ABSTRACT JANET DOOLITTLE: Prediction of host-virus interaction networks (Under the direction of Shawn Gomez) As with other viral pathogens, HIV-1 and dengue virus (DENV) are dependent on their hosts for the bulk of the functions necessary for viral survival and replication. Thus, successful infection depends on the pathogen's ability to manipulate the biological pathways and processes of the organism it infects, while avoiding elimination by the immune system. Protein-protein interactions are one avenue through which viruses can connect with and exploit these host cellular pathways and processes. We developed a computational approach to predict interactions between HIV and human proteins based on structural similarity of 9 HIV-1 proteins to human proteins having known interactions. Using functional data from RNAi studies as a filter, we generated over 2,000 interaction predictions between HIV proteins and 406 unique human proteins. Additional filtering based on Gene Ontology cellular component annotation reduced the number of predictions to 502 interactions involving 137 human proteins. We find numerous known interactions as well as novel interactions showing significant functional relevance based on supporting Gene Ontology and literature evidence. -

Mai Muudatuntuu Ti on Man Mini
MAIMUUDATUNTUU US009809854B2 TI ON MAN MINI (12 ) United States Patent ( 10 ) Patent No. : US 9 ,809 ,854 B2 Crow et al. (45 ) Date of Patent : Nov . 7 , 2017 Whitehead et al. (2005 ) Variation in tissue - specific gene expression ( 54 ) BIOMARKERS FOR DISEASE ACTIVITY among natural populations. Genome Biology, 6 :R13 . * AND CLINICAL MANIFESTATIONS Villanueva et al. ( 2011 ) Netting Neutrophils Induce Endothelial SYSTEMIC LUPUS ERYTHEMATOSUS Damage , Infiltrate Tissues, and Expose Immunostimulatory Mol ecules in Systemic Lupus Erythematosus . The Journal of Immunol @(71 ) Applicant: NEW YORK SOCIETY FOR THE ogy , 187 : 538 - 552 . * RUPTURED AND CRIPPLED Bijl et al. (2001 ) Fas expression on peripheral blood lymphocytes in MAINTAINING THE HOSPITAL , systemic lupus erythematosus ( SLE ) : relation to lymphocyte acti vation and disease activity . Lupus, 10 :866 - 872 . * New York , NY (US ) Crow et al . (2003 ) Microarray analysis of gene expression in lupus. Arthritis Research and Therapy , 5 :279 - 287 . * @(72 ) Inventors : Mary K . Crow , New York , NY (US ) ; Baechler et al . ( 2003 ) Interferon - inducible gene expression signa Mikhail Olferiev , Mount Kisco , NY ture in peripheral blood cells of patients with severe lupus . PNAS , (US ) 100 ( 5 ) : 2610 - 2615. * GeneCards database entry for IFIT3 ( obtained from < http : / /www . ( 73 ) Assignee : NEW YORK SOCIETY FOR THE genecards. org /cgi - bin / carddisp .pl ? gene = IFIT3 > on May 26 , 2016 , RUPTURED AND CRIPPLED 15 pages ) . * Navarra et al. (2011 ) Efficacy and safety of belimumab in patients MAINTAINING THE HOSPITAL with active systemic lupus erythematosus : a randomised , placebo FOR SPECIAL SURGERY , New controlled , phase 3 trial . The Lancet , 377 :721 - 731. * York , NY (US ) Abramson et al . ( 1983 ) Arthritis Rheum . -

RNA-Seq Analysis of Seasonal and Individual Variation in Blood Transcriptomes of Healthy Managed Bottlenose Dolphins Jeanine S
Morey et al. BMC Genomics (2016) 17:720 DOI 10.1186/s12864-016-3020-8 RESEARCH ARTICLE Open Access RNA-Seq analysis of seasonal and individual variation in blood transcriptomes of healthy managed bottlenose dolphins Jeanine S. Morey1*, Marion G. Neely1, Denise Lunardi2, Paul E. Anderson3, Lori H. Schwacke1, Michelle Campbell4 and Frances M. Van Dolah1,5* Abstract Background: The blood transcriptome can reflect both systemic exposures and pathological changes in other organs of the body because immune cells recirculate through the blood, lymphoid tissues, and affected sites. In human and veterinary medicine, blood transcriptome analysis has been used successfully to identify markers of disease or pathological conditions, but can be confounded by large seasonal changes in expression. In comparison, the use of transcriptomic based analyses in wildlife has been limited. Here we report a longitudinal study of four managed bottlenose dolphins located in Waikoloa, Hawaii, serially sampled (approximately monthly) over the course of 1 year to establish baseline information on the content and variation of the dolphin blood transcriptome. Results: Illumina based RNA-seq analyses were carried out using both the Ensembl dolphin genome and a de novo blood transcriptome as guides. Overall, the blood transcriptome encompassed a wide array of cellular functions and processes and was relatively stable within and between animals over the course of 1 year. Principal components analysis revealed moderate clustering by sex associated with the variation among global gene expression profiles (PC1, 22 % of variance). Limited seasonal change was observed, with < 2.5 % of genes differentially expressed between winter and summer months (FDR < 0.05). -

Robles JTO Supplemental Digital Content 1
Supplementary Materials An Integrated Prognostic Classifier for Stage I Lung Adenocarcinoma based on mRNA, microRNA and DNA Methylation Biomarkers Ana I. Robles1, Eri Arai2, Ewy A. Mathé1, Hirokazu Okayama1, Aaron Schetter1, Derek Brown1, David Petersen3, Elise D. Bowman1, Rintaro Noro1, Judith A. Welsh1, Daniel C. Edelman3, Holly S. Stevenson3, Yonghong Wang3, Naoto Tsuchiya4, Takashi Kohno4, Vidar Skaug5, Steen Mollerup5, Aage Haugen5, Paul S. Meltzer3, Jun Yokota6, Yae Kanai2 and Curtis C. Harris1 Affiliations: 1Laboratory of Human Carcinogenesis, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 2Division of Molecular Pathology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 3Genetics Branch, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 4Division of Genome Biology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 5Department of Chemical and Biological Working Environment, National Institute of Occupational Health, NO-0033 Oslo, Norway. 6Genomics and Epigenomics of Cancer Prediction Program, Institute of Predictive and Personalized Medicine of Cancer (IMPPC), 08916 Badalona (Barcelona), Spain. List of Supplementary Materials Supplementary Materials and Methods Fig. S1. Hierarchical clustering of based on CpG sites differentially-methylated in Stage I ADC compared to non-tumor adjacent tissues. Fig. S2. Confirmatory pyrosequencing analysis of DNA methylation at the HOXA9 locus in Stage I ADC from a subset of the NCI microarray cohort. 1 Fig. S3. Methylation Beta-values for HOXA9 probe cg26521404 in Stage I ADC samples from Japan. Fig. S4. Kaplan-Meier analysis of HOXA9 promoter methylation in a published cohort of Stage I lung ADC (J Clin Oncol 2013;31(32):4140-7). Fig. S5. Kaplan-Meier analysis of a combined prognostic biomarker in Stage I lung ADC. -

WO 2015/179918 Al 3 December 2015 (03.12.2015) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2015/179918 Al 3 December 2015 (03.12.2015) P O P C T (51) International Patent Classification: (81) Designated States (unless otherwise indicated, for every C07K 19/00 (2006.01) A61K 39/395 (2006.01) kind of national protection available): AE, AG, AL, AM, C07K 16/46 (2006.01) A61P 3/10 (2006.01) AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, A61K 38/20 (2006.01) A61P 5/50 (2006.01) BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, (21) International Application Number: HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, PCT/AU20 15/050283 KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, (22) International Filing Date: MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, 27 May 2015 (27.05.2015) PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, (25) Filing Language: English TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (26) Publication Language: English (84) Designated States (unless otherwise indicated, for every (30) Priority Data: kind of regional protection available): ARIPO (BW, GH, 62/003,492 27 May 2014 (27.05.2014) US GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, TZ, UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, (71) Applicant: THE UNIVERSITY OF QUEENSLAND TJ, TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, [AU/AU]; The University of Queensland, St Lucia, DK, EE, ES, FI, FR, GB, GR, HR, HU, IE, IS, IT, LT, LU, Queensland 4072 (AU). -

Global Patterns of Changes in the Gene Expression Associated with Genesis of Cancer a Dissertation Submitted in Partial Fulfillm
Global Patterns Of Changes In The Gene Expression Associated With Genesis Of Cancer A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy at George Mason University By Ganiraju Manyam Master of Science IIIT-Hyderabad, 2004 Bachelor of Engineering Bharatiar University, 2002 Director: Dr. Ancha Baranova, Associate Professor Department of Molecular & Microbiology Fall Semester 2009 George Mason University Fairfax, VA Copyright: 2009 Ganiraju Manyam All Rights Reserved ii DEDICATION To my parents Pattabhi Ramanna and Veera Venkata Satyavathi who introduced me to the joy of learning. To friends, family and colleagues who have contributed in work, thought, and support to this project. iii ACKNOWLEDGEMENTS I would like to thank my advisor, Dr. Ancha Baranova, whose tolerance, patience, guidance and encouragement helped me throughout the study. This dissertation would not have been possible without her ever ending support. She is very sincere and generous with her knowledge, availability, compassion, wisdom and feedback. I would also like to thank Dr. Vikas Chandhoke for funding my research generously during my doctoral study at George Mason University. Special thanks go to Dr. Patrick Gillevet, Dr. Alessandro Giuliani, Dr. Maria Stepanova who devoted their time to provide me with their valuable contributions and guidance to formulate this project. Thanks to the faculty of Molecular and Micro Biology (MMB) department, Dr. Jim Willett and Dr. Monique Vanhoek in embedding valuable thoughts to this dissertation by being in my dissertation committee. I would also like to thank the present and previous doctoral program directors, Dr. Daniel Cox and Dr. Geraldine Grant, for facilitating, allowing, and encouraging me to work in this project. -

C M Gabe Thesis Student200902650 2 Pdfx
Production and purification of recombinant amelogenins for investigating proteopathic mechanisms associated with amelogenesis imperfecta Claire Madialin Gabe Submitted in accordance with the requirements for the degree of Doctorate of Philosophy The University of Leeds Faculty of Medicine and Health School of Dentistry November 2018 - ii - Intellectual Property and Publication Statements The candidate confirms that the work submitted is his/her own, except where work which has formed part of jointly-authored publications has been included. The contribution of the candidate and the other authors to this work has been explicitly indicated below. The candidate confirms that appropriate credit has been given within the thesis where reference has been made to the work of others. The published work is integrated in the Materials and Methods, Results and Discussion sections of the thesis. The method Sections 2.1.1.1 (expression of recombinant amelogenin), 2.1.1.4 and 2.1.2.3 (polyacrylamide gel electrophoreses), 2.1.2.1 (nickel column chromatography), and the Figures 41, 42, 43 and 73 are based on work from the jointly authored publication: GABE, C. M., BROOKES, S. J. & KIRKHAM, J. 2017. Preparative SDS PAGE as an Alternative to His-Tag Purification of Recombinant Amelogenin. Frontiers in Physiology, 8, 424. All three authors Miss Claire Madialin Gabe (CMG), Dr. Seven Joseph Brookes (SJB) and Prof. Jennifer Kirkham (JK) contributed to the design of the experiments. CMG carried out the practical work (data acquisition). All three authors CMG, SJB and JK contributed to drafting the manuscript and approved the final submitted version. This copy has been supplied on the understanding that it is copyright material and that no quotation from the thesis may be published without proper acknowledgement. -
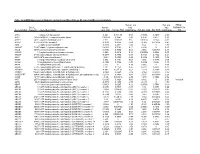
Mrna Expression in Human Leiomyoma and Eker Rats As Measured by Microarray Analysis
Table 3S: mRNA Expression in Human Leiomyoma and Eker Rats as Measured by Microarray Analysis Human_avg Rat_avg_ PENG_ Entrez. Human_ log2_ log2_ RAPAMYCIN Gene.Symbol Gene.ID Gene Description avg_tstat Human_FDR foldChange Rat_avg_tstat Rat_FDR foldChange _DN A1BG 1 alpha-1-B glycoprotein 4.982 9.52E-05 0.68 -0.8346 0.4639 -0.38 A1CF 29974 APOBEC1 complementation factor -0.08024 0.9541 -0.02 0.9141 0.421 0.10 A2BP1 54715 ataxin 2-binding protein 1 2.811 0.01093 0.65 0.07114 0.954 -0.01 A2LD1 87769 AIG2-like domain 1 -0.3033 0.8056 -0.09 -3.365 0.005704 -0.42 A2M 2 alpha-2-macroglobulin -0.8113 0.4691 -0.03 6.02 0 1.75 A4GALT 53947 alpha 1,4-galactosyltransferase 0.4383 0.7128 0.11 6.304 0 2.30 AACS 65985 acetoacetyl-CoA synthetase 0.3595 0.7664 0.03 3.534 0.00388 0.38 AADAC 13 arylacetamide deacetylase (esterase) 0.569 0.6216 0.16 0.005588 0.9968 0.00 AADAT 51166 aminoadipate aminotransferase -0.9577 0.3876 -0.11 0.8123 0.4752 0.24 AAK1 22848 AP2 associated kinase 1 -1.261 0.2505 -0.25 0.8232 0.4689 0.12 AAMP 14 angio-associated, migratory cell protein 0.873 0.4351 0.07 1.656 0.1476 0.06 AANAT 15 arylalkylamine N-acetyltransferase -0.3998 0.7394 -0.08 0.8486 0.456 0.18 AARS 16 alanyl-tRNA synthetase 5.517 0 0.34 8.616 0 0.69 AARS2 57505 alanyl-tRNA synthetase 2, mitochondrial (putative) 1.701 0.1158 0.35 0.5011 0.6622 0.07 AARSD1 80755 alanyl-tRNA synthetase domain containing 1 4.403 9.52E-05 0.52 1.279 0.2609 0.13 AASDH 132949 aminoadipate-semialdehyde dehydrogenase -0.8921 0.4247 -0.12 -2.564 0.02993 -0.32 AASDHPPT 60496 aminoadipate-semialdehyde -

Investigation of Differential Gene Expression in Parkinson's Disease Patients: a Whole Transcriptome Approach
Investigation of differential gene expression in Parkinson's disease patients: A whole transcriptome approach Genevie Borrageiro Thesis presented in partial fulfilment of the requirements for the degree of Master of Science (Human Genetics) in the Faculty of Medicine and Health Sciences at Stellenbosch University Supervisor: Professor Soraya Bardien Faculty of Medicine and Health Sciences Department of Biomedical Sciences Co-Supervisor: Professor Sian Hemmings Faculty of Medicine and Health Sciences Department of Biomedical Sciences December 2016 Stellenbosch University https://scholar.sun.ac.za DECLARATION By submitting this thesis/dissertation, I declare that the entirety of the work contained therein is my own, original work, that I am the sole author thereof (save to the extent explicitly otherwise stated), that reproduction and publication thereof by Stellenbosch University will not infringe any third party rights and that I have not previously in its entirety or in part submitted it for obtaining any qualification. Signature: …………………… Date: ........................................ Copyright © 2016 Stellenbosch University All rights reserved Page | i Stellenbosch University https://scholar.sun.ac.za Abstract Parkinson’s disease (PD) is the most common neurodegenerative movement disorder and is characterized by the loss of dopaminergic neurons in the substantia nigra pars compacta. Dopaminergic neuronal loss results in motor symptoms such as resting tremor, bradykinesia, rigidity and postural instability. Although several PD-causing genes have been identified, the process(es) that lead to progressive neuronal loss is poorly understood. Therefore, PD research efforts have turned to genome- wide next generation sequencing technologies to provide clues to the etiology. The focus of this study was to investigate the entire transcriptome in South African patients with PD (particularly in the under- studied mixed ancestry population) to identify biological pathways that may shed light on the pathobiology of PD. -

A Non-Random Walk Through the Genome Comment Brian Oliver* and Tom Misteli†
Review A non-random walk through the genome comment Brian Oliver* and Tom Misteli† Addresses: *National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Department of Health and Human Services, Bethesda, MD 20892, USA. †National Cancer Institute, National Institutes of Health, Department of Health and Human Services, Bethesda, MD 20892, USA. Correspondence: Brian Oliver. E-mail: [email protected]. Tom Misteli. E-mail: [email protected] reviews Published: 31 March 2005 Genome Biology 2005, 6:214 (doi:10.1186/gb-2005-6-4-214) The electronic version of this article is the complete one and can be found online at http://genomebiology.com/2005/6/4/214 © 2005 BioMed Central Ltd reports Abstract Recent publications on a wide range of eukaryotes indicate that genes showing particular expression patterns are not randomly distributed in the genome but are clustered into contiguous regions that we call neighborhoods. It seems probable that this organization is related to chromatin and the structure of the nucleus. deposited research Consideration of gene regulation as an interaction between the lingering questions about the best methods for finding them basal transcription machinery, regulators, and a segment of and about how to avoid being tricked by spurious or artifac- naked DNA containing a gene has been an extremely successful tual patterns, the body of available evidence leaves little model for understanding how genetic information is read [1]. It doubt that they exist. With a few exceptions from classical is impossible to separate this model from the great success of molecular biology, the regulatory mechanisms underlying modern molecular genetics. -
Abnormal Gene Expression in Regular and Aggregated Somatic Cell Nuclear Transfer Placentas Bo-Woong Sim1,2, Chae-Won Park1, Myung-Hwa Kang3 and Kwan-Sik Min1*
Sim et al. BMC Biotechnology (2017) 17:34 DOI 10.1186/s12896-017-0355-4 RESEARCHARTICLE Open Access Abnormal gene expression in regular and aggregated somatic cell nuclear transfer placentas Bo-Woong Sim1,2, Chae-Won Park1, Myung-Hwa Kang3 and Kwan-Sik Min1* Abstract Background: Placental defects in somatic cell nuclear transfer (SCNT) are a major cause of complications during pregnancy. One of the most critical factors for the success of SCNT is the successful epigenetic reprogramming of donor cells. Recently, it was reported that the placental weight in mice cloned with the aggregated SCNT method was significantly reduced. Here, we examine the profile of abnormal gene expression using microarray technology in both regular SCNT and aggregated SCNT placentas as well as in vivo fertilization placentas. One SCNT embryo was aggregated with two 2 to 4 -cell stage tetraploid embryos from B6D2F1 mice (C57BL/6 × DBA/2). Results: In SCNT placentas, 206 (1.6%) of the 12,816 genes probed were either up-regulated or down-regulated by more than two-fold. However, 52 genes (0.4%) showed differential expression in aggregated SCNT placentas compared to that in controls. In comparison of both types of SCNT placentas with the controls, 33 (92%) out of 36 genes were found to be up-regulated (>2-fold) in SCNT placentas. Among 36 genes, 13 (36%) genes were up- regulated in the aggregated SCNT placentas. Eighty-five genes were down-regulated in SCNT placentas compared with the controls. However, only 9 (about 10.5%) genes were down-regulated in the aggregated SCNT placentas.