Thalassemia, a Human Blood Disorder
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Developmental Genetics of Hemoglobin Synthesis in Chironomus Darrel Starr English Iowa State University
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 1968 The developmental genetics of hemoglobin synthesis in Chironomus Darrel Starr English Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Genetics Commons Recommended Citation English, Darrel Starr, "The developmental genetics of hemoglobin synthesis in Chironomus " (1968). Retrospective Theses and Dissertations. 3660. https://lib.dr.iastate.edu/rtd/3660 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. This dissertation has been microfilmed exactly as received 6 8-14,785 ENGLISH, Barrel Starr, 1936- THE DEVELOPMENTAL GENETICS OF HEMOGLOBIN SYNTHESIS IN CHIRONOMUS. Iowa State University, Ph.D., 1968 Biology- Genetics University Microfilms, Inc., Ann Arbor, Michigan THE DEVELOPMENTAL GENETICS OF HEMOGLOBIN SYNTHESIS IN CHIRONOMUS by Darrel Starr English A Dissertation Submitted to the Graduate Faculty in Partial Fulfillment of The Requirements for the Degree of DOCTOR OF PHILOSOPHY Major Subject: Genetics Approved: Signature was redacted for privacy. In Charge of MajdA Work Signature was redacted for privacy. Head ^ Major Department Signature was redacted for privacy. -

Hypoxia-Induced Alpha-Globin Expression in Syncytiotrophoblasts Mimics the Pattern Observed in Preeclamptic Placentas
International Journal of Molecular Sciences Article Hypoxia-Induced Alpha-Globin Expression in Syncytiotrophoblasts Mimics the Pattern Observed in Preeclamptic Placentas Zahra Masoumi 1,* , Lena Erlandsson 1, Eva Hansson 1, Mattias Magnusson 2, Eva Mezey 3 and Stefan R. Hansson 1,4 1 Department of Clinical Sciences Lund, Division of Obstetrics and Gynecology, Lund University, SE-22184 Lund, Sweden; [email protected] (L.E.); [email protected] (E.H.); [email protected] (S.R.H.) 2 Department of Molecular Medicine and Gene Therapy, Lund University, SE-22184 Lund, Sweden; [email protected] 3 Adult Stem Cell Section, National Institute of Dental and Craniofacial Research, National Institutes of Health, 9000 Rockville Pike, Bethesda, MD 20892, USA; [email protected] 4 Skåne University Hospital, SE-22184 Lund, Sweden * Correspondence: [email protected] Abstract: Preeclampsia (PE) is a pregnancy disorder associated with placental dysfunction and elevated fetal hemoglobin (HbF). Early in pregnancy the placenta harbors hematopoietic stem and progenitor cells (HSPCs) and is an extramedullary source of erythropoiesis. However, globin ex- pression is not unique to erythroid cells and can be triggered by hypoxia. To investigate the role of the placenta in increasing globin levels previously reported in PE, flow cytometry, histological and Citation: Masoumi, Z.; Erlandsson, immunostaining and in situ analyses were used on placenta samples and ex vivo explant cultures. L.; Hansson, E.; Magnusson, M.; Our results indicated that in PE pregnancies, placental HSPC homing and erythropoiesis were not Mezey, E.; Hansson, S.R. affected. Non-erythroid alpha-globin mRNA and protein, but not gamma-globin, were detected in Hypoxia-Induced Alpha-Globin Expression in Syncytiotrophoblasts syncytiotrophoblasts and stroma of PE placenta samples. -

8.5 X12.5 Doublelines.P65
Cambridge University Press 978-0-521-87519-6 - Disorders of Hemoglobin: Genetics, Pathophysiology, and Clinical Management, Second Edition Edited by Martin H. Steinberg, Bernard G. Forget, Douglas R. Higgs and David J. Weatherall Index More information anti-inflammatory therapy, 762–763 thalassemia-related complications, 779 sulfasalazine, nuclear factor (NF)-kB, 762 transplant-related complications, 778–779 targeting ET-1, 762–763 S-linked haplotypes, African/Indo-European, anti-oxidant therapy targeting erythrocyte, 638–640 765–766 burst forming unit-erythroid (BFU-E), 10, 29 deferiprone, 765 oral glutamine, 765 calcium-activated potassium channel (Gardos oral N-acetyl-cysteine, 765–766 pathway), 167–168 anti-oxidant therapy targeting vascular, 763–765 capillary electrophoresis, 660 Index Apo A-I mimetics, 764 capillary IEF, 660 NO, 763–764 carbon monoxide poisoning, 613–616 statins, 764 clinical features, 615 xanthine oxidase inhibitors, 764–765 diagnosis, treatment, 615–616 anti-thrombotic therapy epidemiology, 613–614 -thalassemia, 761–762 cardiac, arterial abnormalities, 151 sickle cell disease, 761–762 cardiac abnormalities, ATRX syndrome, 305 aortagonad-mesonephros (AGM), 6 cardiovascular disease, 652 Apo A-I mimetics, 764 cation content, cellular hydration, 164–172 apoptosis, vasculature permeability, 193–194 calcium-activated potassium channel, 167–168 assays, assay systems, 7, 142 cation permeability alterations, 166–167 ATMDS. See ␣ thalassemia-myelodysplastic cell calcium, calcium pump activity, 167 syndromes cell sodium, -

Serum Albumin OS=Homo Sapiens
Protein Name Cluster of Glial fibrillary acidic protein OS=Homo sapiens GN=GFAP PE=1 SV=1 (P14136) Serum albumin OS=Homo sapiens GN=ALB PE=1 SV=2 Cluster of Isoform 3 of Plectin OS=Homo sapiens GN=PLEC (Q15149-3) Cluster of Hemoglobin subunit beta OS=Homo sapiens GN=HBB PE=1 SV=2 (P68871) Vimentin OS=Homo sapiens GN=VIM PE=1 SV=4 Cluster of Tubulin beta-3 chain OS=Homo sapiens GN=TUBB3 PE=1 SV=2 (Q13509) Cluster of Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 (P60709) Cluster of Tubulin alpha-1B chain OS=Homo sapiens GN=TUBA1B PE=1 SV=1 (P68363) Cluster of Isoform 2 of Spectrin alpha chain, non-erythrocytic 1 OS=Homo sapiens GN=SPTAN1 (Q13813-2) Hemoglobin subunit alpha OS=Homo sapiens GN=HBA1 PE=1 SV=2 Cluster of Spectrin beta chain, non-erythrocytic 1 OS=Homo sapiens GN=SPTBN1 PE=1 SV=2 (Q01082) Cluster of Pyruvate kinase isozymes M1/M2 OS=Homo sapiens GN=PKM PE=1 SV=4 (P14618) Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 Clathrin heavy chain 1 OS=Homo sapiens GN=CLTC PE=1 SV=5 Filamin-A OS=Homo sapiens GN=FLNA PE=1 SV=4 Cytoplasmic dynein 1 heavy chain 1 OS=Homo sapiens GN=DYNC1H1 PE=1 SV=5 Cluster of ATPase, Na+/K+ transporting, alpha 2 (+) polypeptide OS=Homo sapiens GN=ATP1A2 PE=3 SV=1 (B1AKY9) Fibrinogen beta chain OS=Homo sapiens GN=FGB PE=1 SV=2 Fibrinogen alpha chain OS=Homo sapiens GN=FGA PE=1 SV=2 Dihydropyrimidinase-related protein 2 OS=Homo sapiens GN=DPYSL2 PE=1 SV=1 Cluster of Alpha-actinin-1 OS=Homo sapiens GN=ACTN1 PE=1 SV=2 (P12814) 60 kDa heat shock protein, mitochondrial OS=Homo -

Newborn Screening Result for Bart's Hemoglobin
NEWBORN SCREENING RESULT FOR BART’S HEMOGLOBIN Physician’s information sheet developed by the Nebraska Newborn Screening Program with review by James Harper, MD, Pediatric Hematologist with UNMC Follow-up program, and member of the Nebraska Newborn Screening Advisory Committee. BACKGROUND The alpha thalassemias result from the loss of alpha globin genes. There are normally four genes for alpha globin production so that the loss of one to four genes is possible. The lack of one gene causes alpha thalassemia 2 (silent carrier) with no clinically detectable problems but may cause small amounts of hemoglobin Barts to be present in newborn blood samples. Alpha thalassemia trait (Alpha thalassemia 1) results from loss of two genes and causes a mild microcytic anemia which may resemble iron deficiency anemia. The loss of three genes causes hemoglobin H diseases which is a moderately severe form of thalassemia. The lack of all four genes causes hydrops fetalis and is usually fatal in utero. In general, only the loss of one or two genes is seen in African Americans. Individuals from Southeast Asia and the Mediterranean may have all four types of alpha thalassemia. The percentage of hemoglobin Barts in the blood sample may indicate the number of alpha genes that have been lost. However, the percentage of hemoglobin Barts is not directly measurable with the current methodology used by the newborn screening laboratory. Only the presence of Barts hemoglobin in relation to fetal and adult hemoglobin, and variants S, C, D and E can be detected. RECOMMENDED WORK UP In addition to the standard newborn hemoglobinopathy confirmation (hemoglobin electrophoresis), to separate those patients with alpha thalassemia silent carrier from the patients with alpha thalassemia trait, we recommend that these babies have the following labs drawn at their 6 month well baby check: CBC with retic count, ferritin, and a hemoglobin electropheresis. -

Adult, Embryonic and Fetal Hemoglobin Are Expressed in Human Glioblastoma Cells
514 INTERNATIONAL JOURNAL OF ONCOLOGY 44: 514-520, 2014 Adult, embryonic and fetal hemoglobin are expressed in human glioblastoma cells MARWAN EMARA1,2, A. ROBERT TURNER1 and JOAN ALLALUNIS-TURNER1 1Department of Oncology, University of Alberta and Alberta Health Services, Cross Cancer Institute, Edmonton, AB T6G 1Z2, Canada; 2Center for Aging and Associated Diseases, Zewail City of Science and Technology, Cairo, Egypt Received September 7, 2013; Accepted October 7, 2013 DOI: 10.3892/ijo.2013.2186 Abstract. Hemoglobin is a hemoprotein, produced mainly in Introduction erythrocytes circulating in the blood. However, non-erythroid hemoglobins have been previously reported in other cell Globins are hemo-containing proteins, have the ability to types including human and rodent neurons of embryonic bind gaseous ligands [oxygen (O2), nitric oxide (NO) and and adult brain, but not astrocytes and oligodendrocytes. carbon monoxide (CO)] reversibly. They have been described Human glioblastoma multiforme (GBM) is the most aggres- in prokaryotes, fungi, plants and animals with an enormous sive tumor among gliomas. However, despite extensive basic diversity of structure and function (1). To date, hemoglobin, and clinical research studies on GBM cells, little is known myoglobin, neuroglobin (Ngb) and cytoglobin (Cygb) repre- about glial defence mechanisms that allow these cells to sent the vertebrate globin family with distinct function and survive and resist various types of treatment. We have tissue distributions (2). During ontogeny, developing erythro- shown previously that the newest members of vertebrate blasts sequentially express embryonic {[Gower 1 (ζ2ε2), globin family, neuroglobin (Ngb) and cytoglobin (Cygb), are Gower 2 (α2ε2), and Portland 1 (ζ2γ2)] to fetal [Hb F(α2γ2)] expressed in human GBM cells. -
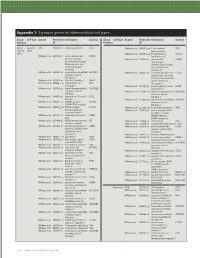
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

LOINC Top 2000++ Lab Observations V1.6 (PDF)
LOINC Mapper's Guide to Top 2000++ US Lab Tests v1.6 June 2017 Page 1 of 112 B C EFGH I P LOINC # Long Common Name Class Rank Example Example Comments System Override UCUM UCUM Adjusted 1 Display 2 General Guidance 1) Ask your test kit and instrument manufacturer(s) and referral labs about which LOINC codes are relevant for their products. Increasingly, test kit and instrument manufacturers are requesting LOINC codes for their new test. Some of the larger manufacturers have mapped their routine tests done on to LOINC codes. Check with these in vitro diagnostic companies for the LOINC codes relevant for their tests. In addition, the largest referral laboratories in the US have mapped their high- to medium-volume tests to LOINC. Getting the LOINC mappings from either of these sources will save you time. 2) When mapping, search against the LOINC common test list. In RELMA and on search.loinc.org you can set the search parameters to only look at the common tests. Work through the mapping by lab section. Realize that LOINC does not encompass terms that may be used in your lab system for internal accounting or “diagnostic” variables that are provided as indicators that might be used to trigger a follow up test, but are not supposed to be reported to the ordering provider because the results are not reliable enough. Blood cell counters usually report such indicators. 3) Obtain a master list of tests for mapping. RELMA has a function that will convert a large set of HL7 result (ORU) messages into a database that carries the name of the order, the units of measure, and sample data that can be the source of frequency statistics for deciding which terms to tackle first. -

Molecular Epidemiology and Hematologic Characterization of Δβ
Jiang et al. BMC Medical Genetics (2020) 21:43 https://doi.org/10.1186/s12881-020-0981-x RESEARCH ARTICLE Open Access Molecular epidemiology and hematologic characterization of δβ-thalassemia and hereditary persistence of fetal hemoglobin in 125,661 families of greater Guangzhou area, the metropolis of southern China Fan Jiang1,2, Liandong Zuo2, Dongzhi Li2, Jian Li2, Xuewei Tang2, Guilan Chen2, Jianying Zhou2, Hang Lu2 and Can Liao1,2* Abstract Background: Individuals with δβ-thalassemia/HPFH and β-thalassemia usually present with intermedia or thalassemia major. No large-scale survey on HPFH/δβ-thalassemia in southern China has been reported to date. The purpose of this study was to examine the molecular epidemiology and hematologic characteristics of these disorders in Guangzhou, the largest city in Southern China, to offer advice for thalassemia screening programs and genetic counseling. Methods: A total of 125,661 couples participated in pregestational thalassemia screening. 654 subjects with fetal hemoglobin (HbF) level ≥ 5% were selected for further investigation. Gap-PCR combined with Multiplex ligation dependent probe amplification (MLPA) was used to screen for β-globin gene cluster deletions. Gene sequencing for the promoter region of HBG1 /HBG2 gene was performed for all those subjects. Results: A total of 654 individuals had hemoglobin (HbF) levels≥5, and 0.12% of the couples were found to be heterozygous for HPFH/δβ-thalassemia, including Chinese Gγ (Aγδβ)0-thal, Southeast Asia HPFH (SEA-HPFH), Taiwanese deletion and Hb Lepore–Boston–Washington. The highest prevalence was observed in the Huadu district and the lowest in the Nansha district. Three cases were identified as carrying β-globin gene cluster deletions, which had not been previously reported. -

Identification of Novel HPFH-Like Mutations by CRISPR Base Editing That Elevates the Expression of Fetal Hemoglobin
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.30.178715; this version posted July 1, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Identification of novel HPFH-like mutations by CRISPR base editing that elevates the expression of fetal hemoglobin Nithin Sam Ravi1,3, Beeke Wienert4,8,9, Stacia K. Wyman4, Jonathan Vu4, Aswin Anand Pai2, Poonkuzhali Balasubramanian2, Yukio Nakamura7, Ryo Kurita10, Srujan Marepally1, Saravanabhavan Thangavel1, Shaji R. Velayudhan1,2, Alok Srivastava1,2, Mark A. DeWitt 4,5, Jacob E. Corn4,6, Kumarasamypet M. Mohankumar1* 1. Centre for Stem Cell Research (a unit of inStem, Bengaluru), Christian Medical College Campus, Bagayam, Vellore-632002, Tamil Nadu, India. 2. Department of Haematology, Christian Medical College Hospital, IDA Scudder Rd, Vellore-632002, Tamil Nadu, India. 3. Sree Chitra Tirunal Institute for Medical Sciences and Technology, Thiruvananthapuram - 695 011, Kerala, India. 4. Innovative Genomics Institute, University of California Berkeley, Berkeley, California- 94704, USA. 5. Department of Microbiology, Immunology, and Molecular Genetics, University of California - Los Angeles, Los Angeles, California - 90095, USA. 6. Institute of Molecular Health Sciences, Department of Biology, ETH Zurich, Otto-Stern- Weg, Zurich - 8092, Switzerland. 7. Cell Engineering Division, RIKEN BioResource Center, Tsukuba, Ibaraki 305-0074, Japan. 8. Institute of Data Science and Biotechnology, Gladstone Institutes, San Francisco, California -94158, USA 9. School of Biotechnology and Biomolecular Sciences, University of New South Wales - 2052, Sydney, Australia. 10. Research and Development Department Central Blood Institute Blood Service Headquarters Japanese Red Cross Society, 2-1-67 Tatsumi, Tokyo, 135-8521, Japan *Corresponding Author: Dr. -

Code Lists Page 1
Code lists Page 1 Code lists AESEV Page 2 AESEV Codelist Name: Severity/Intensity Scale for Adverse Events Description: A scale that defines the degree or state of disease existing in a patient as a result of the occurrence of an adverse event. (NCI) C41338,1; Grade 1 C41339,2; Grade 2 C41340,3; Grade 3 AGEU Page 3 AGEU Codelist Name: Age Unit Description: Those units of time that are routinely used to express the age of a subject. C25301,Days C25529,Hours; h; hr C29846,Month C29844,Week C29848,Year CMDOSFRM Page 4 CMDOSFRM Codelist Name: Concomitant Medication Dose Form Description: A terminology subset of the CDISC SDTM Pharmaceutical Dosage Form codelist created for CDASH Concomitant Medication Dose Form codelist. (NCI) C42887,AEROSOL; aer C25158,CAPSULE; cap C28944,CREAM; C42933,GAS; C42934,GEL; C42966,OINTMENT; oint C42968,PATCH; C42972,POWDER; C42989,SPRAY; C42993,SUPPOSITORY; supp C42994,SUSPENSION; susp C42998,TABLET; tab CMDOSFRQ Page 5 CMDOSFRQ Codelist Name: Concomitant Medication Dosing Frequency per Interval Description: A terminology subset of the CDISC SDTM Frequency codelist created for CDASH Concomitant Medication Dosing Frequency per Interval codelist. (NCI) C64496,BID; BD; Twice per day C64499,PRN; As needed C25473,QD; Daily C64530,QID; 4 times per day C64498,QM; Every Month; Per Month C64525,QOD; Every other day C64527,TID; 3 times per day C17998,UNKNOWN; U; UNK; Unknown CMDOSU Page 6 CMDOSU Codelist Name: Concomitant Medication Dose Units Description: A terminology subset of the CDISC SDTM Unit codelist created for CDASH Concomitant Medication Dose Units codelist. (NCI) C48480,CAPSULE; Capsule Dosing Unit; cap C48155,g; Gram C48579,IU; IE; International Unit C28253,mg; Milligram C28254,mL; Milliliter; cm3 C65060,PUFF; Puff Dosing Unit C48542,TABLET; Tablet Dosing Unit; tab C48152,ug; Microgram; mcg CMROUTE Page 7 CMROUTE Codelist Name: Concomitant Medication Route of Administration Description: A terminology subset of the CDISC SDTM Route codelist created for CDASH Concomitant Medication Route of Administration codelist. -

Normal, C=Critical
Patient Report |FINAL Client: Example Client ABC123 Patient: Patient, Example 123 Test Drive Salt Lake City, UT 84108 DOB 7/22/1970 UNITED STATES Gender: Female Patient Identifiers: 01234567890ABCD, 012345 Physician: Doctor, Example Visit Number (FIN): 01234567890ABCD Collection Date: 00/00/0000 00:00 Hemoglobin Evaluation Reflexive Cascade ARUP test code 2005792 Hemoglobin A 97.1 % (Ref Interval: 95.0-97.9) Hemoglobin A2 2.6 % (Ref Interval: 2.0-3.5) Hemoglobin F 0.3 % (Ref Interval: 0.0-2.1) REFERENCE INTERVAL: Hemoglobin F Access complete set of age- and/or gender-specific reference intervals for this test in the ARUP Laboratory Test Directory (aruplab.com). Hemoglobin S 0.0 % (Ref Interval: 0.0-0.0) Hemoglobin C 0.0 % (Ref Interval: 0.0-0.0) Hemoglobin E 0.0 % (Ref Interval: 0.0-0.0) Hemoglobin - Other 0.0 % (Ref Interval: 0.0-0.0) Sickle Cell Solubility Not Performed Hemoglobin, Capillary Electrophoresis Performed Hemoglobin Evaluation See Note Beta Globin Full Gene Sequencing Not Applicable H=High, L=Low, *=Abnormal, C=Critical Patient: Patient, Example ARUP Accession: 21-048-400496 Patient Identifiers: 01234567890ABCD, 012345 Visit Number (FIN): 01234567890ABCD Page 1 of 4 | Printed: 3/11/2021 11:41:09 AM 4848 Patient Report |FINAL Beta Globin (HBB) Del/Dup Result Not Applicable Alpha Thalassemia HBA1 and HBA2 Seq Not Applicable Gamma Globin (HBG1 and HBG2) Sequencing Not Applicable Hemoglobin Cascade Interpretation See Note H=High, L=Low, *=Abnormal, C=Critical Patient: Patient, Example ARUP Accession: 21-048-400496 Patient Identifiers: 01234567890ABCD, 012345 Visit Number (FIN): 01234567890ABCD Page 2 of 4 | Printed: 3/11/2021 11:41:09 AM 4848 Patient Report |FINAL RESULT Normal hemoglobin evaluation.