Mrna Encoding Sec61β, a Tail-Anchored Protein, Is Localized on the Endoplasmic Reticulum Xianying A
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Patterns of Green Fluorescent Protein Expression in Transgenic Plants
Color profile: Generic CMYK printer profile Composite Default screen Plant Molecular Biology Reporter 18: 141a–141i, 2000 © 2000 International Society for Plant Molecular Biology. Printed in Canada. Publish by Abstract Patterns of Green Fluorescent Protein Expression in Transgenic Plants BRIAN K. HARPER1 and C. NEAL STEWART JR.2,* 1Novartis Agricultural Biotechnology Research, Inc., 3054 Cornwallis Rd., Research Triangle Park, NC 27709; 2Dept of Biology, University of North Carolina, Greensboro, NC 27402-6174 Abstract. Modified forms of genes encoding green fluorescent protein (GFP) can be mac- roscopically detected when expressed in whole plants. This technology has opened up new uses for GFP such as monitoring transgene presence and expression in the environment once it is linked or fused to a gene of interest. When whole-plant or whole-organ GFP vi- sualization is required, GFP should be predictably expressed and reliably fluorescent. In this study the whole plant expression and fluorescence patterns of a mGFP5er gene driven by the cauliflower mosaic virus 35S promoter was studied in intact GFP-expressing trans- genic tobacco (Nicotiana tabacum cv. Xanthi). It was shown that GFP synthesis levels in single plant organs were similar to GUS activity levels from published data when driven by the same promoter. Under the control of the 35S promoter, high expression of GFP can be used to visualize stems, young leaves, flowers, and organs where the 35S promoter is most active. Modified forms of GFP could replace GUS as the visual marker gene of choice. Key words: expression patterns, green fluorescent protein, marker genes Patterns of IntroductionGFP expression Harper and Stewart Since the discovery of green fluorescent protein (GFP) from the jellyfish Aequorea victoria it has become a frequently used tool in biology. -

Supplementary Materials and Method Immunostaining and Western Blot
Supplementary Materials and Method Immunostaining and Western Blot Analysis For immunofluorescence staining, mouse and human cells were fixed with 4% paraformaldehyde- PBS for 15 min. Following Triton-X100 permeabilization and blocking, cells were incubated with primary antibodies overnight at 4°C following with Alexa 594-conjugated secondary antibodies at 4°C for 1 hour (Thermo Fisher Scientific, 1:1000). Samples were mounted using VECTASHIELD Antifade Mounting Medium with DAPI (Vector Laboratories) and immunofluorescence was detected using Olympus confocal microscopy. For western blot analysis, cells were lysed on ice using RIPA buffer supplemented with protease and phosphatase inhibitors (Sigma). Primary Antibodies for Immunostaining and Western Blot Analysis: Yap (14074, Cell Signaling), pYAP (4911, Cell Signaling), Lats1 (3477, Cell Signaling), pLats1( 8654, Cell Signaling), Wnt5a (2530, Cell Signaling), cleaved Caspase-3 (9661, Cell Signaling), Ki-67 (VP-K451, Vector Laboratories), Cyr61 (sc-13100, Santa Cruz Biotechnology), CTGF (sc-14939, Santa Cruz Biotechnology), AXL (8661, Cell Signaling), pErk (4376, Cell Signaling), pMEK (4376, Cell Signaling), Ck-19 (16858-1-AP, Proteintech), Actin (A2228, Sigma Aldrich), Vinculin (V4139, Sigma Aldrich), Kras (sc-30, Santa Cruz Biotechnology). Ectopic expression of YAP1 and WNT5A in mouse and human cells To generate YAP1S127A-expressing stable Pa04C cells, Pa04C cells were transfected with a linearized pcDNA3.1 plasmid with or without YAP1 cDNA containing S127A substitution. Two days post-transfection using Lipofectamine1000, cultures were selected in G418 (Sigma) and single clones were picked and expanded for further analysis. Overexpression of YAPS127A or WNT5A in human or mouse cells other than Pa04C were acheieved with lentivral infection. Briefly, lentivirus infection was performed by transfecting 293T cells with either GFP control, YAP1S127A, or WNT5A cloned in pHAGE lentivirus vector {EF1α promoter-GW-IRES-eGFP (GW: Gateway modified)}. -
Download Validation Data
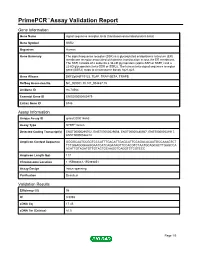
PrimePCR™Assay Validation Report Gene Information Gene Name signal sequence receptor, beta (translocon-associated protein beta) Gene Symbol SSR2 Organism Human Gene Summary The signal sequence receptor (SSR) is a glycosylated endoplasmic reticulum (ER) membrane receptor associated with protein translocation across the ER membrane. The SSR consists of 2 subunits a 34-kD glycoprotein (alpha-SSR or SSR1) and a 22-kD glycoprotein (beta-SSR or SSR2). The human beta-signal sequence receptor gene (SSR2) maps to chromosome bands 1q21-q23. Gene Aliases DKFZp686F19123, TLAP, TRAP-BETA, TRAPB RefSeq Accession No. NC_000001.10, NT_004487.19 UniGene ID Hs.74564 Ensembl Gene ID ENSG00000163479 Entrez Gene ID 6746 Assay Information Unique Assay ID qHsaCID0014663 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000295702, ENST00000529008, ENST00000480567, ENST00000531917, ENST00000526212 Amplicon Context Sequence GGGGCAATCCGGTCCCATTTGACATTGAGCATTCCAGACACAATGCCAAAGTCT TCTGGAGGGAAGGAATCATCAGATAGTTCCACGTCTAATGCAGCACTTGAGCCA ACATTGTAGATGTTGTACTGCAAGGTCAGGTCTCGTCCC Amplicon Length (bp) 117 Chromosome Location 1:155988061-155989851 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 98 R2 0.9998 cDNA Cq 17.45 cDNA Tm (Celsius) 81.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report SSR2, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference -

A Clearer Picture of the ER Translocon Complex Max Gemmer and Friedrich Förster*
© 2020. Published by The Company of Biologists Ltd | Journal of Cell Science (2020) 133, jcs231340. doi:10.1242/jcs.231340 REVIEW A clearer picture of the ER translocon complex Max Gemmer and Friedrich Förster* ABSTRACT et al., 1986). SP-equivalent N-terminal transmembrane helices that The endoplasmic reticulum (ER) translocon complex is the main gate are not cleaved off can also target proteins to the ER through the into the secretory pathway, facilitating the translocation of nascent same mechanism. In this SRP-dependent co-translational ER- peptides into the ER lumen or their integration into the lipid membrane. targeting mode, ribosomes associate with the ER membrane via ER Protein biogenesis in the ER involves additional processes, many of translocon complexes. These membrane protein complexes them occurring co-translationally while the nascent protein resides at translocate nascent soluble proteins into the ER, integrate nascent the translocon complex, including recruitment of ER-targeted membrane proteins into the ER membrane, mediate protein folding ribosome–nascent-chain complexes, glycosylation, signal peptide and membrane protein topogenesis, and modify them chemically. In cleavage, membrane protein topogenesis and folding. To perform addition to co-translational protein import and translocation, distinct such varied functions on a broad range of substrates, the ER ER translocon complexes enable post-translational translocation and translocon complex has different accessory components that membrane integration. This post-translational pathway is widespread associate with it either stably or transiently. Here, we review recent in yeast (Panzner et al., 1995), whereas higher eukaryotes primarily structural and functional insights into this dynamically constituted use it for relatively short peptides (Schlenstedt and Zimmermann, central hub in the ER and its components. -

Scholarworks@UNO
University of New Orleans ScholarWorks@UNO University of New Orleans Theses and Dissertations Dissertations and Theses Summer 8-4-2011 Identification and characterization of enzymes involved in the biosynthesis of different phycobiliproteins in cyanobacteria Avijit Biswas University of New Orleans, [email protected] Follow this and additional works at: https://scholarworks.uno.edu/td Part of the Biochemistry, Biophysics, and Structural Biology Commons Recommended Citation Biswas, Avijit, "Identification and characterization of enzymes involved in the biosynthesis of different phycobiliproteins in cyanobacteria" (2011). University of New Orleans Theses and Dissertations. 446. https://scholarworks.uno.edu/td/446 This Dissertation-Restricted is protected by copyright and/or related rights. It has been brought to you by ScholarWorks@UNO with permission from the rights-holder(s). You are free to use this Dissertation-Restricted in any way that is permitted by the copyright and related rights legislation that applies to your use. For other uses you need to obtain permission from the rights-holder(s) directly, unless additional rights are indicated by a Creative Commons license in the record and/or on the work itself. This Dissertation-Restricted has been accepted for inclusion in University of New Orleans Theses and Dissertations by an authorized administrator of ScholarWorks@UNO. For more information, please contact [email protected]. Identification and characterization of enzymes involved in biosynthesis of different phycobiliproteins in cyanobacteria A Thesis Submitted to the Graduate Faculty of the University of New Orleans in partial fulfillment of the requirements for the degree of Doctor of Philosophy In Chemistry (Biochemistry) By Avijit Biswas B.S. -

Development and Applications of Superfolder and Split Fluorescent Protein Detection Systems in Biology
International Journal of Molecular Sciences Review Development and Applications of Superfolder and Split Fluorescent Protein Detection Systems in Biology Jean-Denis Pedelacq 1,* and Stéphanie Cabantous 2,* 1 Institut de Pharmacologie et de Biologie Structurale, IPBS, Université de Toulouse, CNRS, UPS, 31077 Toulouse, France 2 Centre de Recherche en Cancérologie de Toulouse (CRCT), Inserm, Université Paul Sabatier-Toulouse III, CNRS, 31037 Toulouse, France * Correspondence: [email protected] (J.-D.P.); [email protected] (S.C.) Received: 15 June 2019; Accepted: 8 July 2019; Published: 15 July 2019 Abstract: Molecular engineering of the green fluorescent protein (GFP) into a robust and stable variant named Superfolder GFP (sfGFP) has revolutionized the field of biosensor development and the use of fluorescent markers in diverse area of biology. sfGFP-based self-associating bipartite split-FP systems have been widely exploited to monitor soluble expression in vitro, localization, and trafficking of proteins in cellulo. A more recent class of split-FP variants, named « tripartite » split-FP,that rely on the self-assembly of three GFP fragments, is particularly well suited for the detection of protein–protein interactions. In this review, we describe the different steps and evolutions that have led to the diversification of superfolder and split-FP reporter systems, and we report an update of their applications in various areas of biology, from structural biology to cell biology. Keywords: fluorescent protein; superfolder; split-GFP; bipartite; tripartite; folding; PPI 1. Superfolder Fluorescent Proteins: Progenitor of Split Fluorescent Protein (FP) Systems Previously described mutations that improve the physical properties and expression of green fluorescent protein (GFP) color variants in the host organism have already been the subject of several reviews [1–4] and will not be described here. -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

Intrinsic Indicators for Specimen Degradation
Laboratory Investigation (2013) 93, 242–253 & 2013 USCAP, Inc All rights reserved 0023-6837/13 $32.00 Intrinsic indicators for specimen degradation Jie Li1, Catherine Kil1, Kelly Considine1, Bartosz Smarkucki1, Michael C Stankewich1, Brian Balgley2 and Alexander O Vortmeyer1 Variable degrees of molecular degradation occur in human surgical specimens before clinical examination and severely affect analytical results. We therefore initiated an investigation to identify protein markers for tissue degradation assessment. We exposed 4 cell lines and 64 surgical/autopsy specimens to defined periods of time at room temperature before procurement (experimental cold ischemic time (CIT)-dependent tissue degradation model). Using two-dimen- sional fluorescence difference gel electrophoresis in conjunction with mass spectrometry, we performed comparative proteomic analyses on cells at different CIT exposures and identified proteins with CIT-dependent changes. The results were validated by testing clinical specimens with western blot analysis. We identified 26 proteins that underwent dynamic changes (characterized by continuous quantitative changes, isoelectric changes, and/or proteolytic cleavages) in our degradation model. These changes are strongly associated with the length of CIT. We demonstrate these proteins to represent universal tissue degradation indicators (TDIs) in clinical specimens. We also devised and implemented a unique degradation measure by calculating the quantitative ratio between TDIs’ intact forms and their respective degradation- -

A Crosstalk Between the RNA Binding Protein Smaug and the Hedgehog Pathway Links Cell Signaling to Mrna Regulation in Drosophila Lucía Bruzzone
A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila Lucía Bruzzone To cite this version: Lucía Bruzzone. A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila. Cellular Biology. Université Sorbonne Paris Cité, 2018. English. NNT : 2018USPCC234. tel-02899776 HAL Id: tel-02899776 https://tel.archives-ouvertes.fr/tel-02899776 Submitted on 15 Jul 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Thèse de doctorat de l’Université Sorbonne Paris Cité Préparée à l’Université Paris Diderot Ecole doctorale HOB n° 561 Institut Jacques Monod / Equipe Développement, Signalisation et Trafic A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in Drosophila Lucía Bruzzone Thèse de doctorat de Biologie Dirigée par Anne Plessis Présentée et soutenue publiquement à Paris le 19 mars 2018 Président du jury: Alain Zider / Professeur Université Paris Diderot -

Investigations on the Impact of Toxic Cyanobacteria on Fish : As
INVESTIGATIONS ON THE IMPACT OF TOXIC CYANOBACTERIA ON FISH - AS EXEMPLIFIED BY THE COREGONIDS IN LAKE AMMERSEE - DISSERTATION Zur Erlangung des akademischen Grades des Doktors der Naturwissenschaften an der Universität Konstanz Fachbereich Biologie Vorgelegt von BERNHARD ERNST Tag der mündlichen Prüfung: 05. Nov. 2008 Referent: Prof. Dr. Daniel Dietrich Referent: Prof. Dr. Karl-Otto Rothhaupt Referent: Prof. Dr. Alexander Bürkle 2 »Erst seit gestern und nur für einen Tag auf diesem Planeten weilend, können wir nur hoffen, einen Blick auf das Wissen zu erhaschen, das wir vermutlich nie erlangen werden« Horace-Bénédict de Saussure (1740-1799) Pionier der modernen Alpenforschung & Wegbereiter des Alpinismus 3 ZUSAMMENFASSUNG Giftige Cyanobakterien beeinträchtigen Organismen verschiedenster Entwicklungsstufen und trophischer Ebenen. Besonders bedroht sind aquatische Organismen, weil sie von Cyanobakterien sehr vielfältig beeinflussbar sind und ihnen zudem oft nur sehr begrenzt ausweichen können. Zu den toxinreichsten Cyanobakterien gehören Arten der Gattung Planktothrix. Hierzu zählt auch die Burgunderblutalge Planktothrix rubescens, eine Cyanobakterienart die über die letzten Jahrzehnte im Besonderen in den Seen der Voralpenregionen zunehmend an Bedeutung gewonnen hat. An einigen dieser Voralpenseen treten seit dem Erstarken von P. rubescens existenzielle, fischereiwirtschaftliche Probleme auf, die wesentlich auf markante Wachstumseinbrüche bei den Coregonenbeständen (Coregonus sp.; i.e. Renken, Felchen, etc.) zurückzuführen sind. So auch -

Immunohistochemistry Stain Offerings
immunohistochemistry stain offerings TRUSTED PATHOLOGISTS. INVALUABLE ANSWERS.™ MARCHMAY 20172021 www.aruplab.com/ap-ihcaruplab.com/ap-ihc InformationInformation in this brochurein this brochure is current is current as of as May of March 2021. 2017. All content All content is subject is subject to tochange. change. Please contactPlease ARUPcontact ClientARUP Services Client Services at 800-522-2787 at (800) 522-2787 with any with questions any questions or concerns.or concerns. ARUP LABORATORIES As a nonprofit, academic institution of the University of Utah and its Department We believe in of Pathology, ARUP believes in collaborating, sharing and contributing to laboratory science in ways that benefit our clients and their patients. collaborating, Our test menu is one of the broadest in the industry, encompassing more sharing and than 3,000 tests, including highly specialized and esoteric assays. We offer comprehensive testing in the areas of genetics, molecular oncology, pediatrics, contributing pain management, and more. to laboratory ARUP’s clients include many of the nation’s university teaching hospitals and children’s hospitals, as well as multihospital groups, major commercial science in ways laboratories, and group purchasing organizations. We believe that healthcare should be delivered as close to the patient as possible, which is why we support that provide our clients’ efforts to be the principal healthcare provider in the communities they serve by offering highly complex assays and accompanying consultative support. the best value Offering analytics, consulting, and decision support services, ARUP provides for the patient. clients with the utilization management tools necessary to prosper in this time of value-based care. -

X-Ray Structure of a Protein-Conducting Channel
articles X-ray structure of a protein-conducting channel Bert van den Berg1*, William M. Clemons Jr1*, Ian Collinson2, Yorgo Modis3, Enno Hartmann4, Stephen C. Harrison3 & Tom A. Rapoport1 1Howard Hughes Medical Institute and Department of Cell Biology, Harvard Medical School, 240 Longwood Avenue, Boston, Massachusetts 02115, USA 2Max Planck Institute of Biophysics, Marie-Curie-Strasse 13-15, D-60439 Frankfurt am Main, Germany 3Howard Hughes Medical Institute, Children’s Hospital and Harvard Medical School, 320 Longwood Avenue, Boston, Massachusetts 02115, USA 4University Luebeck, Institute for Biology, Ratzeburger Allee 160, Luebeck, D-23538, Germany * These authors contributed equally to this work ........................................................................................................................................................................................................................... A conserved heterotrimeric membrane protein complex, the Sec61 or SecY complex, forms a protein-conducting channel, allowing polypeptides to be transferred across or integrated into membranes. We report the crystal structure of the complex from Methanococcus jannaschii at a resolution of 3.2 A˚ . The structure suggests that one copy of the heterotrimer serves as a functional translocation channel. The a-subunit has two linked halves, transmembrane segments 1–5 and 6–10, clamped together by the g-subunit. A cytoplasmic funnel leading into the channel is plugged by a short helix. Plug displacement can open the channel into an ‘hourglass’ with a ring of hydrophobic residues at its constriction. This ring may form a seal around the translocating polypeptide, hindering the permeation of other molecules. The structure also suggests mechanisms for signal-sequence recognition and for the lateral exit of transmembrane segments of nascent membrane proteins into lipid, and indicates binding sites for partners that provide the driving force for translocation.