(12) Patent Application Publication (10) Pub. No.: US 2009/0163376 A1 Li Et Al
Total Page:16
File Type:pdf, Size:1020Kb
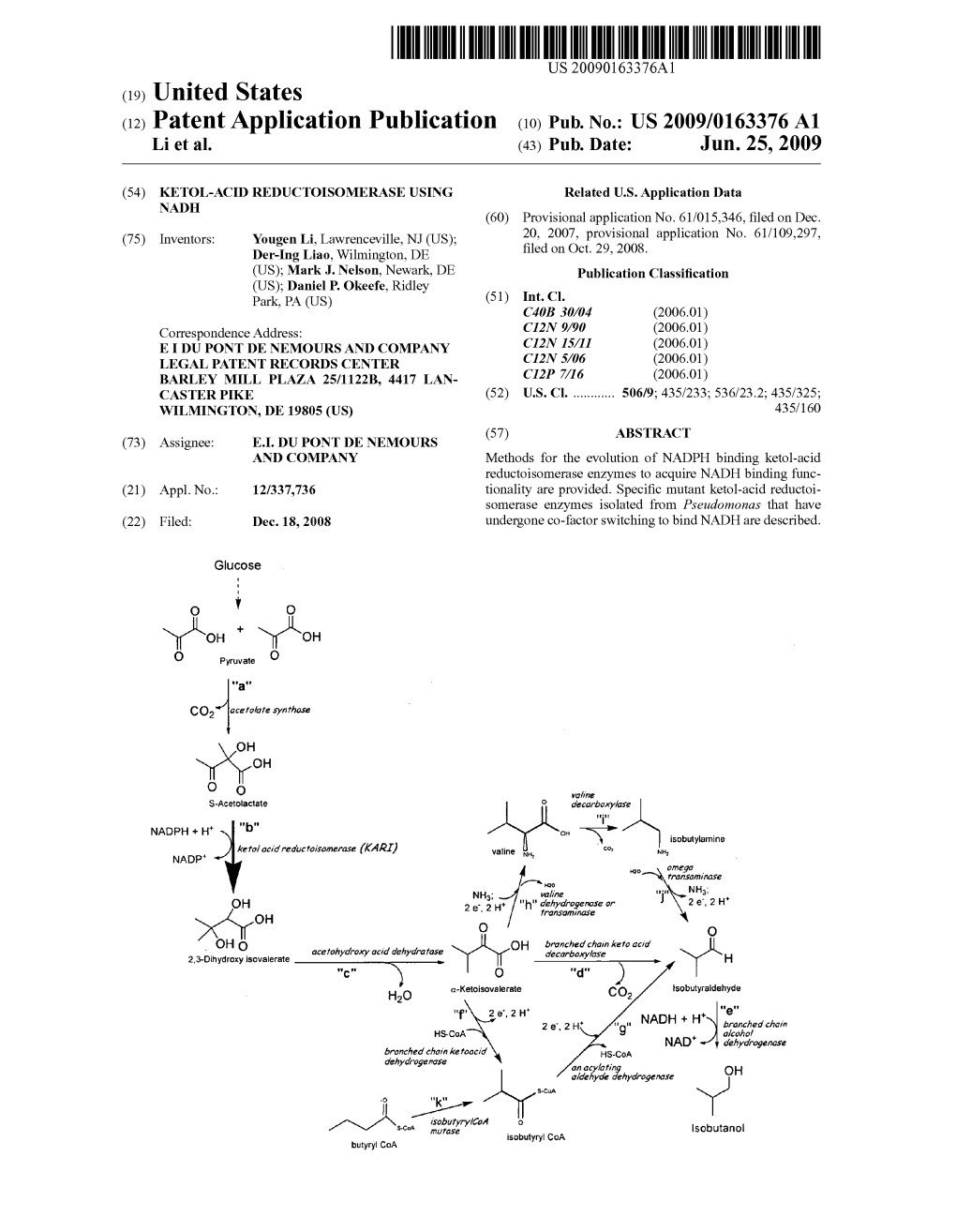
Load more
Recommended publications
-

Aliphatic 1-Amino Acid Decarboxylase from Ferns (Filicopsida) Thomas Hartmann
Aliphatic 1-Amino Acid Decarboxylase from Ferns (Filicopsida) Thomas Hartmann. Klaus Bax. and Renate Scholz Institut für Pharmazeutische Biologie der Technischen Universität Braunschweig, Mendels- sohnstr. 1, D-3300 Braunschweig. Bundesrepublik Deutschland Z. Naturforsch. 39c, 2 4 -3 0 (1984); received Septem ber 26, 1983 Ferns, Filicopsida, Polypodium vulgare. Aliphatic 1-Amino Acid Decarboxylase, Occurrence and Distribution A screening of 27 fern species (Filicopsida) out of 9 families revealed that 25 species were able to decarboxylate 1-leucine to 3-methylbutylamine (isoamylamine). The enzyme of Polvpodium vulgäre has partially been purified and characterisized. All attempts to solubilize it from acetone preparations failed; however, approx. 50% of total activity could be extracted from dry material in the presence of detergents at high concentration. The soluble enzyme was purified 132-fold. 1-Methionine was found the best substrate followed by norvaline, leucine, norleucine, isoleucine, homocysteine, valine. It has been confirmed that these substrates are decarboxylated by a single enzyme. The pH-optimum was at pH 5.0 (particulate preparation) and pH 4.5 (soluble enzyme). Decarboxylation is dependent on pyridoxal-5'-phosphate (PLP). A strictly substrate dependent coenzyme dissociation was observed which could largely be prevented by addition of 2 -oxo-acids, such as glyoxylate or pyruvate. Apodecarboxylase prepared by prolonged substrate incubation was found to be extremely labile at pH 4.5 but stable at pH 6.5. A comparison of the fern enzyme with bacterial valine decarboxylase (EC 4.1.1.14) and leucine decarboxylase of red algae revealed great similarities especially in substrate specificity. It is suggested to unify these activities as “aliphatic 1-amino acid decarboxylase”. -

(12) Patent Application Publication (10) Pub. No.: US 2009/0292100 A1 Fiene Et Al
US 20090292100A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2009/0292100 A1 Fiene et al. (43) Pub. Date: Nov. 26, 2009 (54) PROCESS FOR PREPARING (86). PCT No.: PCT/EP07/57646 PENTAMETHYLENE 1.5-DIISOCYANATE S371 (c)(1), (75) Inventors: Martin Fiene, Niederkirchen (DE): (2), (4) Date: Jan. 9, 2009 (DE);Eckhard Wolfgang Stroefer, Siegel, Mannheim (30) Foreign ApplicationO O Priority Data Limburgerhof (DE); Stephan Aug. 1, 2006 (EP) .................................. O61182.56.4 Freyer, Neustadt (DE); Oskar Zelder, Speyer (DE); Gerhard Publication Classification Schulz, Bad Duerkheim (DE) (51) Int. Cl. Correspondence Address: CSG 18/00 (2006.01) OBLON, SPIVAK, MCCLELLAND MAIER & CD7C 263/2 (2006.01) NEUSTADT, L.L.P. CI2P I3/00 (2006.01) 194O DUKE STREET CD7C 263/10 (2006.01) ALEXANDRIA, VA 22314 (US) (52) U.S. Cl. ........... 528/85; 560/348; 435/128; 560/347; 560/355 (73) Assignee: BASFSE, LUDWIGSHAFEN (DE) (57) ABSTRACT (21) Appl. No.: 12/373,088 The present invention relates to a process for preparing pen tamethylene 1,5-diisocyanate, to pentamethylene 1,5-diiso (22) PCT Filed: Jul. 25, 2007 cyanate prepared in this way and to the use thereof. US 2009/0292100 A1 Nov. 26, 2009 PROCESS FOR PREPARING ene diisocyanates, especially pentamethylene 1,4-diisocyan PENTAMETHYLENE 1.5-DIISOCYANATE ate. Depending on its preparation, this proportion may be up to several % by weight. 0014. The pentamethylene 1,5-diisocyanate prepared in 0001. The present invention relates to a process for pre accordance with the invention has, in contrast, a proportion of paring pentamethylene 1,5-diisocyanate, to pentamethylene the branched pentamethylene diisocyanate isomers of in each 1.5-diisocyanate prepared in this way and to the use thereof. -

12) United States Patent (10
US007635572B2 (12) UnitedO States Patent (10) Patent No.: US 7,635,572 B2 Zhou et al. (45) Date of Patent: Dec. 22, 2009 (54) METHODS FOR CONDUCTING ASSAYS FOR 5,506,121 A 4/1996 Skerra et al. ENZYME ACTIVITY ON PROTEIN 5,510,270 A 4/1996 Fodor et al. MICROARRAYS 5,512,492 A 4/1996 Herron et al. 5,516,635 A 5/1996 Ekins et al. (75) Inventors: Fang X. Zhou, New Haven, CT (US); 5,532,128 A 7/1996 Eggers Barry Schweitzer, Cheshire, CT (US) 5,538,897 A 7/1996 Yates, III et al. s s 5,541,070 A 7/1996 Kauvar (73) Assignee: Life Technologies Corporation, .. S.E. al Carlsbad, CA (US) 5,585,069 A 12/1996 Zanzucchi et al. 5,585,639 A 12/1996 Dorsel et al. (*) Notice: Subject to any disclaimer, the term of this 5,593,838 A 1/1997 Zanzucchi et al. patent is extended or adjusted under 35 5,605,662 A 2f1997 Heller et al. U.S.C. 154(b) by 0 days. 5,620,850 A 4/1997 Bamdad et al. 5,624,711 A 4/1997 Sundberg et al. (21) Appl. No.: 10/865,431 5,627,369 A 5/1997 Vestal et al. 5,629,213 A 5/1997 Kornguth et al. (22) Filed: Jun. 9, 2004 (Continued) (65) Prior Publication Data FOREIGN PATENT DOCUMENTS US 2005/O118665 A1 Jun. 2, 2005 EP 596421 10, 1993 EP 0619321 12/1994 (51) Int. Cl. EP O664452 7, 1995 CI2O 1/50 (2006.01) EP O818467 1, 1998 (52) U.S. -

(12) United States Patent (10) Patent No.: US 9.273,330 B2 Bramucci Et Al
US00927.333 OB2 (12) United States Patent (10) Patent No.: US 9.273,330 B2 Bramucci et al. (45) Date of Patent: Mar. 1, 2016 (54) BUTANOL TOLERANCE IN 8,206,970 B2 6/2012 Eliot et al. MCROORGANISMS 8,222,017 B2 7/2012 Li et al. 8,241,878 B2 8/2012 Anthony et al. (71) Applicant: Butamax Advanced Biofuels LLC, 3. E: 858: Ration s al Wilmington, DE (US) 8,372,612 B2 2/2013 Larossa et al. 8,389,252 B2 3/2013 Larossa (72) Inventors: Michael G. Bramucci, Boothwyn, PA 8,455,224 B2 6/2013 Paul (US); Vasantha Nagarajan, 8,455.225 B2 6/2013 Bramucci et al. Wilmington,s DE (US) - E.Kw E. E. E.a 8,557.562 B2 10/2013 Bramucci et al. (73) Assignee: Butamax Advanced Biofuels LLC, 8,614,085 B2 12/2013 Van Dyk Wilmington, DE (US) 8,637,281 B2 1/2014 Paul et al. 8,637,289 B2 1/2014 Anthony et al. (*)c Notice:- r Sibi tO E. site th still 8,669,0948,652,823 B2 3/20142/2014 AnthonyFlint et al. et al. patent 1s extended or adjusted under 8,691,540 B2 4/2014 Bramucci et al. U.S.C. 154(b) by 2 days. 8,735,114 B2 5/2014 Donaldson et al. 8,765,433 B2 7/2014 Satagopan et al. (21) Appl. No.: 14/045,506 8,785,166 B2 7/2014 Anthony 8,795,992 B2 8/2014 Bramucci et al. (22) Filed: Oct. 3, 2013 5%. -

(12) Patent Application Publication (10) Pub. No.: US 2012/0266329 A1 Mathur Et Al
US 2012026.6329A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2012/0266329 A1 Mathur et al. (43) Pub. Date: Oct. 18, 2012 (54) NUCLEICACIDS AND PROTEINS AND CI2N 9/10 (2006.01) METHODS FOR MAKING AND USING THEMI CI2N 9/24 (2006.01) CI2N 9/02 (2006.01) (75) Inventors: Eric J. Mathur, Carlsbad, CA CI2N 9/06 (2006.01) (US); Cathy Chang, San Marcos, CI2P 2L/02 (2006.01) CA (US) CI2O I/04 (2006.01) CI2N 9/96 (2006.01) (73) Assignee: BP Corporation North America CI2N 5/82 (2006.01) Inc., Houston, TX (US) CI2N 15/53 (2006.01) CI2N IS/54 (2006.01) CI2N 15/57 2006.O1 (22) Filed: Feb. 20, 2012 CI2N IS/60 308: Related U.S. Application Data EN f :08: (62) Division of application No. 1 1/817,403, filed on May AOIH 5/00 (2006.01) 7, 2008, now Pat. No. 8,119,385, filed as application AOIH 5/10 (2006.01) No. PCT/US2006/007642 on Mar. 3, 2006. C07K I4/00 (2006.01) CI2N IS/II (2006.01) (60) Provisional application No. 60/658,984, filed on Mar. AOIH I/06 (2006.01) 4, 2005. CI2N 15/63 (2006.01) Publication Classification (52) U.S. Cl. ................... 800/293; 435/320.1; 435/252.3: 435/325; 435/254.11: 435/254.2:435/348; (51) Int. Cl. 435/419; 435/195; 435/196; 435/198: 435/233; CI2N 15/52 (2006.01) 435/201:435/232; 435/208; 435/227; 435/193; CI2N 15/85 (2006.01) 435/200; 435/189: 435/191: 435/69.1; 435/34; CI2N 5/86 (2006.01) 435/188:536/23.2; 435/468; 800/298; 800/320; CI2N 15/867 (2006.01) 800/317.2: 800/317.4: 800/320.3: 800/306; CI2N 5/864 (2006.01) 800/312 800/320.2: 800/317.3; 800/322; CI2N 5/8 (2006.01) 800/320.1; 530/350, 536/23.1: 800/278; 800/294 CI2N I/2 (2006.01) CI2N 5/10 (2006.01) (57) ABSTRACT CI2N L/15 (2006.01) CI2N I/19 (2006.01) The invention provides polypeptides, including enzymes, CI2N 9/14 (2006.01) structural proteins and binding proteins, polynucleotides CI2N 9/16 (2006.01) encoding these polypeptides, and methods of making and CI2N 9/20 (2006.01) using these polynucleotides and polypeptides. -

All Enzymes in BRENDA™ the Comprehensive Enzyme Information System
All enzymes in BRENDA™ The Comprehensive Enzyme Information System http://www.brenda-enzymes.org/index.php4?page=information/all_enzymes.php4 1.1.1.1 alcohol dehydrogenase 1.1.1.B1 D-arabitol-phosphate dehydrogenase 1.1.1.2 alcohol dehydrogenase (NADP+) 1.1.1.B3 (S)-specific secondary alcohol dehydrogenase 1.1.1.3 homoserine dehydrogenase 1.1.1.B4 (R)-specific secondary alcohol dehydrogenase 1.1.1.4 (R,R)-butanediol dehydrogenase 1.1.1.5 acetoin dehydrogenase 1.1.1.B5 NADP-retinol dehydrogenase 1.1.1.6 glycerol dehydrogenase 1.1.1.7 propanediol-phosphate dehydrogenase 1.1.1.8 glycerol-3-phosphate dehydrogenase (NAD+) 1.1.1.9 D-xylulose reductase 1.1.1.10 L-xylulose reductase 1.1.1.11 D-arabinitol 4-dehydrogenase 1.1.1.12 L-arabinitol 4-dehydrogenase 1.1.1.13 L-arabinitol 2-dehydrogenase 1.1.1.14 L-iditol 2-dehydrogenase 1.1.1.15 D-iditol 2-dehydrogenase 1.1.1.16 galactitol 2-dehydrogenase 1.1.1.17 mannitol-1-phosphate 5-dehydrogenase 1.1.1.18 inositol 2-dehydrogenase 1.1.1.19 glucuronate reductase 1.1.1.20 glucuronolactone reductase 1.1.1.21 aldehyde reductase 1.1.1.22 UDP-glucose 6-dehydrogenase 1.1.1.23 histidinol dehydrogenase 1.1.1.24 quinate dehydrogenase 1.1.1.25 shikimate dehydrogenase 1.1.1.26 glyoxylate reductase 1.1.1.27 L-lactate dehydrogenase 1.1.1.28 D-lactate dehydrogenase 1.1.1.29 glycerate dehydrogenase 1.1.1.30 3-hydroxybutyrate dehydrogenase 1.1.1.31 3-hydroxyisobutyrate dehydrogenase 1.1.1.32 mevaldate reductase 1.1.1.33 mevaldate reductase (NADPH) 1.1.1.34 hydroxymethylglutaryl-CoA reductase (NADPH) 1.1.1.35 3-hydroxyacyl-CoA -

(12) Patent Application Publication (10) Pub. No.: US 2015/0240226A1 Mathur Et Al
US 20150240226A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2015/0240226A1 Mathur et al. (43) Pub. Date: Aug. 27, 2015 (54) NUCLEICACIDS AND PROTEINS AND CI2N 9/16 (2006.01) METHODS FOR MAKING AND USING THEMI CI2N 9/02 (2006.01) CI2N 9/78 (2006.01) (71) Applicant: BP Corporation North America Inc., CI2N 9/12 (2006.01) Naperville, IL (US) CI2N 9/24 (2006.01) CI2O 1/02 (2006.01) (72) Inventors: Eric J. Mathur, San Diego, CA (US); CI2N 9/42 (2006.01) Cathy Chang, San Marcos, CA (US) (52) U.S. Cl. CPC. CI2N 9/88 (2013.01); C12O 1/02 (2013.01); (21) Appl. No.: 14/630,006 CI2O I/04 (2013.01): CI2N 9/80 (2013.01); CI2N 9/241.1 (2013.01); C12N 9/0065 (22) Filed: Feb. 24, 2015 (2013.01); C12N 9/2437 (2013.01); C12N 9/14 Related U.S. Application Data (2013.01); C12N 9/16 (2013.01); C12N 9/0061 (2013.01); C12N 9/78 (2013.01); C12N 9/0071 (62) Division of application No. 13/400,365, filed on Feb. (2013.01); C12N 9/1241 (2013.01): CI2N 20, 2012, now Pat. No. 8,962,800, which is a division 9/2482 (2013.01); C07K 2/00 (2013.01); C12Y of application No. 1 1/817,403, filed on May 7, 2008, 305/01004 (2013.01); C12Y 1 1 1/01016 now Pat. No. 8,119,385, filed as application No. PCT/ (2013.01); C12Y302/01004 (2013.01); C12Y US2006/007642 on Mar. 3, 2006. -

Comparative Modelling of a Novel Enzyme: Mus Musculus Leucine Decarboxylase
Turkish Journal of Chemistry Turk J Chem (2020) 44: 817 – 832 http://journals.tubitak.gov.tr/chem/ © TÜBİTAK Research Article doi:10.3906/kim-2003-63 Comparative modelling of a novel enzyme: Mus musculus leucine decarboxylase Arif Sercan ŞAHUTOĞLU∗ 1Department of Chemistry, Faculty of Science and Arts, Canakkale Onsekiz Mart University, Canakkale, Turkey Received: 30.03.2020 • Accepted/Published Online: 10.05.2020 • Final Version: 01.06.2020 Abstract: Leucine decarboxylase (LDC) is a recently proposed enzyme with no official enzyme commission number yet. It is encoded by the Mus musculus gene Gm853 which is expressed at kidneys, generating isopentylamine, an alkylmonoamine that has not been described to be formed by any metazoan enzyme yet. Although the relevance of LDC in mammalian physiology has not been fully determined, isopentylamine is a potential modulator which may have effects on insulin secretion and healthy gut microbiota formation. The LDC is a stable enzyme that specifically decarboxylates L-leucine but does not decarboxylate ornithine or lysine as its paralogues ornithine decarboxylase (ODC; EC: 4.1.1.17) and lysine decarboxylase (KDC; EC: 4.1.1.18) do. It does not act as an antizyme inhibitor and does not decarboxylate branched amino acids such as valine and isoleucine as it is another paralogue valine decarboxylase (VDC; EC: 4.1.1.14). The crystal structure of the enzyme has not been determined yet but there are homologous structures with complete coverage in Protein Data Bank (PDB) which makes LDC a good candidate for comparative modelling.In this study, homology models of LDC were generated and used in cofactor and substrate docking to understand the structure/function relationship underlying the unique selectivity of LDC enzyme. -

Supplementary Material (ESI) for Natural Product Reports
Electronic Supplementary Material (ESI) for Natural Product Reports. This journal is © The Royal Society of Chemistry 2014 Supplement to the paper of Alexey A. Lagunin, Rajesh K. Goel, Dinesh Y. Gawande, Priynka Pahwa, Tatyana A. Gloriozova, Alexander V. Dmitriev, Sergey M. Ivanov, Anastassia V. Rudik, Varvara I. Konova, Pavel V. Pogodin, Dmitry S. Druzhilovsky and Vladimir V. Poroikov “Chemo- and bioinformatics resources for in silico drug discovery from medicinal plants beyond their traditional use: a critical review” Contents PASS (Prediction of Activity Spectra for Substances) Approach S-1 Table S1. The lists of 122 known therapeutic effects for 50 analyzed medicinal plants with accuracy of PASS prediction calculated by a leave-one-out cross-validation procedure during the training and number of active compounds in PASS training set S-6 Table S2. The lists of 3,345 mechanisms of action that were predicted by PASS and were used in this study with accuracy of PASS prediction calculated by a leave-one-out cross-validation procedure during the training and number of active compounds in PASS training set S-9 Table S3. Comparison of direct PASS prediction results of known effects for phytoconstituents of 50 TIM plants with prediction of known effects through “mechanism-effect” and “target-pathway- effect” relationships from PharmaExpert S-79 S-1 PASS (Prediction of Activity Spectra for Substances) Approach PASS provides simultaneous predictions of many types of biological activity (activity spectrum) based on the structure of drug-like compounds. The approach used in PASS is based on the suggestion that biological activity of any drug-like compound is a function of its structure. -

Springer Handbook of Enzymes
Dietmar Schomburg Ida Schomburg (Eds.) Springer Handbook of Enzymes Alphabetical Name Index 1 23 © Springer-Verlag Berlin Heidelberg New York 2010 This work is subject to copyright. All rights reserved, whether in whole or part of the material con- cerned, specifically the right of translation, printing and reprinting, reproduction and storage in data- bases. The publisher cannot assume any legal responsibility for given data. Commercial distribution is only permitted with the publishers written consent. Springer Handbook of Enzymes, Vols. 1–39 + Supplements 1–7, Name Index 2.4.1.60 abequosyltransferase, Vol. 31, p. 468 2.7.1.157 N-acetylgalactosamine kinase, Vol. S2, p. 268 4.2.3.18 abietadiene synthase, Vol. S7,p.276 3.1.6.12 N-acetylgalactosamine-4-sulfatase, Vol. 11, p. 300 1.14.13.93 (+)-abscisic acid 8’-hydroxylase, Vol. S1, p. 602 3.1.6.4 N-acetylgalactosamine-6-sulfatase, Vol. 11, p. 267 1.2.3.14 abscisic-aldehyde oxidase, Vol. S1, p. 176 3.2.1.49 a-N-acetylgalactosaminidase, Vol. 13,p.10 1.2.1.10 acetaldehyde dehydrogenase (acetylating), Vol. 20, 3.2.1.53 b-N-acetylgalactosaminidase, Vol. 13,p.91 p. 115 2.4.99.3 a-N-acetylgalactosaminide a-2,6-sialyltransferase, 3.5.1.63 4-acetamidobutyrate deacetylase, Vol. 14,p.528 Vol. 33,p.335 3.5.1.51 4-acetamidobutyryl-CoA deacetylase, Vol. 14, 2.4.1.147 acetylgalactosaminyl-O-glycosyl-glycoprotein b- p. 482 1,3-N-acetylglucosaminyltransferase, Vol. 32, 3.5.1.29 2-(acetamidomethylene)succinate hydrolase, p. 287 Vol. -

(12) United States Patent (10) Patent No.: US 8,044,166 B2 Fiene Et Al
USOO80441 66B2 (12) United States Patent (10) Patent No.: US 8,044,166 B2 Fiene et al. (45) Date of Patent: Oct. 25, 2011 (54) PROCESS FOR PREPARING (56) References Cited PENTAMETHYLENE 1.5-DIISOCYANATE U.S. PATENT DOCUMENTS (75) Inventors: Martin Fiene, Niederkirchen (DE); 5,103,045 A * 4/1992 Robin et al. .................. 560,335 Eckhard Stroefer, Mannheim (DE); 2005, OOO3497 A1* 1/2005 Nishi et al. .................... 435/128 Wolfgang Siegel, Limburgerhof (DE); Stephan Freyer, Neustadt (DE); Oskar FOREIGN PATENT DOCUMENTS DE 1900 514 8, 1970 Zelder, Speyer (DE); Gerhard Schulz, DE 26 25 O75 A1 12, 1977 Bad Duerkheim (DE) DE 2625 O75 * 12, 1977 EP O 259 233 A2 3, 1988 (73) Assignee: BASF Aktiengesellschaft, EP O 161419 B1 8, 1989 Ludwigshafen (DE) EP 1482 055 A1 12, 2004 GB 1 225 450 3, 1971 (*) Notice: Subject to any disclaimer, the term of this WO WOO3,O99768 A1 12/2003 patent is extended or adjusted under 35 WO WO 2006.005603 A1 1, 2006 U.S.C. 154(b) by 336 days. OTHER PUBLICATIONS (21) Appl. No.: 12/373,088 Database Caplus Chemical Abstracts Service, Columbus, Ohio, US 1979, T. Lesiak etal "Preparation of Aliphatic diisocyanates without (22) PCT Fled: Jul. 25, 2007 using phosgene” XP002456997.* Enzyme Handbook, Dec 1, 1982, by Asakura Shoten.* (86) PCT NO.: PCT/EP2007/057646 Preparation of Aliphatic Diisocyanates without using Phosgene, Jun. 1969, by Sea Lesiak.* S371 (c)(1), T. Lesiak, et al., “Preparation of aliphatic diisocyanates without using (2), (4) Date: Jan. 9, 2009 phosgene'. Journal f. prakt. Chemie. Band 321, XP002456997. -

UCSF UC San Francisco Electronic Theses and Dissertations
UCSF UC San Francisco Electronic Theses and Dissertations Title Relating protein pharmacology by ligand chemistry Permalink https://escholarship.org/uc/item/5vp5h8g4 Author Keiser, Michael James Publication Date 2009 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California Copyright 2009 by Michael James Keiser i i To my family iii Acknowledgements I thank my advisor Brian Shoichet, for combining concrete foundations of support with the strong beams of frank advice that give it structure. I thank Brian for knowing when to actively guide and when to lead by example. From Brian I learned the power of falsifiable hypotheses defined such that either result, expected or not, will advance the field—and that the unexpected is often the most intriguing. I thank also John Irwin, who was my rotation advisor and fellow traveler down the many roads of this thesis, and who no matter how busy has always made time for me, even out of thin air. I was excited and apprehensive the morning of my first interviews at UCSF, but I ended that day exhilarated. The research atmosphere of Mission Bay is like no other. Professors Patricia Babbitt, Andrej Sali, and Jim Wells have provided years of essential advice, enthusiasm, and direction both in their roles on my committee and off of it. Patsy mapped the lay of the land, Andrej lit the way with lights statistical, and Jim knew where we were going. Critical to the stories that fill the following pages has been the ready support, energy, and expertise of our collaborators.