A Multi-Cancer Gene Signature Associated with Stromal Activation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic Variation Across the Human Olfactory Receptor Repertoire Alters Odor Perception
bioRxiv preprint doi: https://doi.org/10.1101/212431; this version posted November 1, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Genetic variation across the human olfactory receptor repertoire alters odor perception Casey Trimmer1,*, Andreas Keller2, Nicolle R. Murphy1, Lindsey L. Snyder1, Jason R. Willer3, Maira Nagai4,5, Nicholas Katsanis3, Leslie B. Vosshall2,6,7, Hiroaki Matsunami4,8, and Joel D. Mainland1,9 1Monell Chemical Senses Center, Philadelphia, Pennsylvania, USA 2Laboratory of Neurogenetics and Behavior, The Rockefeller University, New York, New York, USA 3Center for Human Disease Modeling, Duke University Medical Center, Durham, North Carolina, USA 4Department of Molecular Genetics and Microbiology, Duke University Medical Center, Durham, North Carolina, USA 5Department of Biochemistry, University of Sao Paulo, Sao Paulo, Brazil 6Howard Hughes Medical Institute, New York, New York, USA 7Kavli Neural Systems Institute, New York, New York, USA 8Department of Neurobiology and Duke Institute for Brain Sciences, Duke University Medical Center, Durham, North Carolina, USA 9Department of Neuroscience, University of Pennsylvania School of Medicine, Philadelphia, Pennsylvania, USA *[email protected] ABSTRACT The human olfactory receptor repertoire is characterized by an abundance of genetic variation that affects receptor response, but the perceptual effects of this variation are unclear. To address this issue, we sequenced the OR repertoire in 332 individuals and examined the relationship between genetic variation and 276 olfactory phenotypes, including the perceived intensity and pleasantness of 68 odorants at two concentrations, detection thresholds of three odorants, and general olfactory acuity. -

Olfactory Receptors Involved in the Perception
(19) TZZ¥ZZ__T (11) EP 3 004 157 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: C07K 14/705 (2006.01) C07K 14/72 (2006.01) 01.11.2017 Bulletin 2017/44 G01N 33/50 (2006.01) G01N 33/566 (2006.01) G01N 33/68 (2006.01) (21) Application number: 13730138.8 (86) International application number: (22) Date of filing: 31.05.2013 PCT/EP2013/061243 (87) International publication number: WO 2014/191047 (04.12.2014 Gazette 2014/49) (54) OLFACTORY RECEPTORS INVOLVED IN THE PERCEPTION OF SWEAT CARBOXYLIC ACIDS AND THE USE THEREOF AN DER WAHRNEHMUNG VON SCHWEISSCARBONSÄUREN BETEILIGTE GERUCHSREZEPTOREN UND VERWENDUNG DAVON RÉCEPTEURS OLFACTIFS IMPLIQUÉS DANS LA PERCEPTION D’ACIDES CARBOXYLIQUES DE SUEUR ET LEUR UTILISATION (84) Designated Contracting States: • VEITHEN, Alex AL AT BE BG CH CY CZ DE DK EE ES FI FR GB 1476 Genappe (BE) GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL PT RO RS SE SI SK SM TR (74) Representative: Vanhalst, Koen et al De Clercq & Partners cvba (43) Date of publication of application: E. Gevaertdreef 10a 13.04.2016 Bulletin 2016/15 9830 Sint-Martens-Latem (BE) (73) Proprietor: ChemCom S.A. (56) References cited: 1070 Brussels (BE) WO-A1-2012/029922 WO-A2-01/27158 WO-A2-2006/094704 US-A1- 2003 207 337 (72) Inventors: • CHATELAIN, Pierre 1150 Brussels (BE) Note: Within nine months of the publication of the mention of the grant of the European patent in the European Patent Bulletin, any person may give notice to the European Patent Office of opposition to that patent, in accordance with the Implementing Regulations. -

Whole-Exome Sequencing in a Japanese Pedigree Implicates A
Bone 122 (2019) 193–198 Contents lists available at ScienceDirect Bone journal homepage: www.elsevier.com/locate/bone Full Length Article Whole-exome sequencing in a Japanese pedigree implicates a rare non- T synonymous single-nucleotide variant in BEST3 as a candidate for mandibular prognathism ⁎ Takashi S. Kajiia, , Akira Okab, Fumio Saitoc, Jun Mitsuid, Junichiro Iidac a Section of Orthodontics, Department of Oral Growth and Development, Fukuoka Dental College, 2-15-1 Tamura, Sawara-ku, Fukuoka 814-0193, Japan b Institute of Medical Sciences, Tokai University, 143 Shimokasuya, Isehara, Kanagawa 259-1193, Japan c Department of Orthodontics, Division of Oral Functional Science, Graduate School of Dental Medicine, Hokkaido University, Kita 13 Nishi 7, Kita-ku, Sapporo 060-8586, Japan d Department of Neurology, Graduate School of Medicine, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-8655, Japan ARTICLE INFO ABSTRACT Keywords: Mandibular prognathism is a phenotype of facial deformity seen in populations around the world, but with Genetics higher incidence among East Asian populations. Five genome-wide nonparametric linkage analyses and a Whole-exome sequencing genome-wide association study to identify susceptibility loci of the phenotype have shown inconsistent results. Bestrophin 3 To explore variants related to mandibular prognathism, we undertook whole-exome sequencing in a Japanese Mandibular prognathism pedigree. The pedigree was ascertained as mandibular prognathism. The pedigree comprised 15 individuals from Endochondral growth 4 generations. Four affected individuals across 2 generations and 5 unaffected individuals were chosenfor Japanese whole-exome sequencing. Five non-synonymous single-nucleotide variants (SNVs) of UBASH3B, OR6M1, OR8D4, OR8B4, and BEST3 genes were detected in all 4 affected individuals, but in none of the 5 unaffected individuals. -
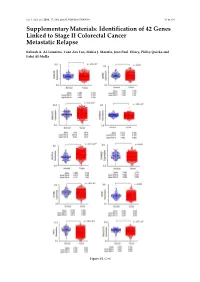
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

PDF Datasheet
Product Datasheet Recombinant Human OR5P2 GST (N-Term) Protein H00120065-P01-2ug Unit Size: 2 ug Store at -80C. Avoid freeze-thaw cycles. Protocols, Publications, Related Products, Reviews, Research Tools and Images at: www.novusbio.com/H00120065-P01 Updated 11/4/2020 v.20.1 Earn rewards for product reviews and publications. Submit a publication at www.novusbio.com/publications Submit a review at www.novusbio.com/reviews/destination/H00120065-P01 Page 1 of 3 v.20.1 Updated 11/4/2020 H00120065-P01-2ug Recombinant Human OR5P2 GST (N-Term) Protein Product Information Unit Size 2 ug Concentration Please see the vial label for concentration. If unlisted please contact technical services. Storage Store at -80C. Avoid freeze-thaw cycles. Preservative No Preservative Purity >80% by SDS-PAGE and Coomassie blue staining Buffer 50 mM Tris-HCl, 10 mM reduced Glutathione, pH 8.0 in the elution buffer. Target Molecular Weight 62.2 kDa Product Description Description A recombinant protein with GST-tag at N-terminal corresponding to the amino acids 1 - 322 of Human OR5P2 full-length ORF Source: Wheat Germ (in vitro) Amino Acid Sequence: MNSLKDGNHTALTGFILLGLTDDPILRVILFMIILSGNLSIIILIRISSQLHHPMYFFL SHLAFADMAYSSSVTPNMLVNFLVERNTVSYLGCAIQLGSAAFFATVECVLLAA MAYDRFVAICSPLLYSTKMSTQVSVQLLLVVYIAGFLIAVSYTTSFYFLLFCGPN QVNHFFCDFAPLLELSCSDISVSTVVLSFSSGSIIVVTVCVIAVCYIYILITILKMRS TEGHHKAFSTCTSHLTVVTLFYGTITFIYVMPNFSYSTDQNKVVSVLYTVVIPML NPLIYSLRNKEIKGALKRELVRKILSHDACYFSRTSNNDIT Gene ID 120065 Gene Symbol OR5P2 Species Human Preparation Method in vitro wheat germ expression system Details of Functionality This protein was produced in an in vitro wheat germ expression system that should preserve correct conformational folding that is necessary for biological function. While it is possible that this protein could display some level of activity, the functionality of this protein has not been explicitly measured or validated. -

The Mutational Landscape of Human Olfactory G Protein-Coupled Receptors
Jimenez et al. BMC Biology (2021) 19:21 https://doi.org/10.1186/s12915-021-00962-0 RESEARCH ARTICLE Open Access The mutational landscape of human olfactory G protein-coupled receptors Ramón Cierco Jimenez1,2, Nil Casajuana-Martin1, Adrián García-Recio1, Lidia Alcántara1, Leonardo Pardo1, Mercedes Campillo1 and Angel Gonzalez1* Abstract Background: Olfactory receptors (ORs) constitute a large family of sensory proteins that enable us to recognize a wide range of chemical volatiles in the environment. By contrast to the extensive information about human olfactory thresholds for thousands of odorants, studies of the genetic influence on olfaction are limited to a few examples. To annotate on a broad scale the impact of mutations at the structural level, here we analyzed a compendium of 119,069 natural variants in human ORs collected from the public domain. Results: OR mutations were categorized depending on their genomic and protein contexts, as well as their frequency of occurrence in several human populations. Functional interpretation of the natural changes was estimated from the increasing knowledge of the structure and function of the G protein-coupled receptor (GPCR) family, to which ORs belong. Our analysis reveals an extraordinary diversity of natural variations in the olfactory gene repertoire between individuals and populations, with a significant number of changes occurring at the structurally conserved regions. A particular attention is paid to mutations in positions linked to the conserved GPCR activation mechanism that could imply phenotypic variation in the olfactory perception. An interactive web application (hORMdb, Human Olfactory Receptor Mutation Database) was developed for the management and visualization of this mutational dataset. -

WO 2019/068007 Al Figure 2
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/068007 Al 04 April 2019 (04.04.2019) W 1P O PCT (51) International Patent Classification: (72) Inventors; and C12N 15/10 (2006.01) C07K 16/28 (2006.01) (71) Applicants: GROSS, Gideon [EVIL]; IE-1-5 Address C12N 5/10 (2006.0 1) C12Q 1/6809 (20 18.0 1) M.P. Korazim, 1292200 Moshav Almagor (IL). GIBSON, C07K 14/705 (2006.01) A61P 35/00 (2006.01) Will [US/US]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., C07K 14/725 (2006.01) P.O. Box 4044, 7403635 Ness Ziona (TL). DAHARY, Dvir [EilL]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., P.O. (21) International Application Number: Box 4044, 7403635 Ness Ziona (IL). BEIMAN, Merav PCT/US2018/053583 [EilL]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., P.O. (22) International Filing Date: Box 4044, 7403635 Ness Ziona (E.). 28 September 2018 (28.09.2018) (74) Agent: MACDOUGALL, Christina, A. et al; Morgan, (25) Filing Language: English Lewis & Bockius LLP, One Market, Spear Tower, SanFran- cisco, CA 94105 (US). (26) Publication Language: English (81) Designated States (unless otherwise indicated, for every (30) Priority Data: kind of national protection available): AE, AG, AL, AM, 62/564,454 28 September 2017 (28.09.2017) US AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, 62/649,429 28 March 2018 (28.03.2018) US CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, (71) Applicant: IMMP ACT-BIO LTD. -

Olfactory Receptors in Non-Chemosensory Tissues
BMB Reports Invited Mini Review Olfactory receptors in non-chemosensory tissues NaNa Kang & JaeHyung Koo* Department of Brain Science, Daegu Gyeongbuk Institute of Science and Technology (DGIST), Daegu 711-873, Korea Olfactory receptors (ORs) detect volatile chemicals that lead to freezing behavior (3-5). the initial perception of smell in the brain. The olfactory re- ORs are localized in the cilia of olfactory sensory neurons ceptor (OR) is the first protein that recognizes odorants in the (OSNs) in the olfactory epithelium (OE) and are activated by olfactory signal pathway and it is present in over 1,000 genes chemical cues, typically odorants at the molecular level, in mice. It is also the largest member of the G protein-coupled which lead to the perception of smell in the brain (6). receptors (GPCRs). Most ORs are extensively expressed in the Tremendous research was conducted since Buck and Axel iso- nasal olfactory epithelium where they perform the appropriate lated ORs as an OE-specific expression in 1991 (7). OR genes, physiological functions that fit their location. However, recent the largest family among the G protein-coupled receptors whole-genome sequencing shows that ORs have been found (GPCRs) (8), constitute more than 1,000 genes on the mouse outside of the olfactory system, suggesting that ORs may play chromosome (9, 10) and more than 450 genes in the human an important role in the ectopic expression of non-chemo- genome (11, 12). sensory tissues. The ectopic expressions of ORs and their phys- Odorant activation shows a distinct signal transduction iological functions have attracted more attention recently since pathway for odorant perception. -

Sean Raspet – Molecules
1. Commercial name: Fructaplex© IUPAC Name: 2-(3,3-dimethylcyclohexyl)-2,5,5-trimethyl-1,3-dioxane SMILES: CC1(C)CCCC(C1)C2(C)OCC(C)(C)CO2 Molecular weight: 240.39 g/mol Volume (cubic Angstroems): 258.88 Atoms number (non-hydrogen): 17 miLogP: 4.43 Structure: Biological Properties: Predicted Druglikenessi: GPCR ligand -0.23 Ion channel modulator -0.03 Kinase inhibitor -0.6 Nuclear receptor ligand 0.15 Protease inhibitor -0.28 Enzyme inhibitor 0.15 Commercial name: Fructaplex© IUPAC Name: 2-(3,3-dimethylcyclohexyl)-2,5,5-trimethyl-1,3-dioxane SMILES: CC1(C)CCCC(C1)C2(C)OCC(C)(C)CO2 Predicted Olfactory Receptor Activityii: OR2L13 83.715% OR1G1 82.761% OR10J5 80.569% OR2W1 78.180% OR7A2 77.696% 2. Commercial name: Sylvoxime© IUPAC Name: N-[4-(1-ethoxyethenyl)-3,3,5,5tetramethylcyclohexylidene]hydroxylamine SMILES: CCOC(=C)C1C(C)(C)CC(CC1(C)C)=NO Molecular weight: 239.36 Volume (cubic Angstroems): 252.83 Atoms number (non-hydrogen): 17 miLogP: 4.33 Structure: Biological Properties: Predicted Druglikeness: GPCR ligand -0.6 Ion channel modulator -0.41 Kinase inhibitor -0.93 Nuclear receptor ligand -0.17 Protease inhibitor -0.39 Enzyme inhibitor 0.01 Commercial name: Sylvoxime© IUPAC Name: N-[4-(1-ethoxyethenyl)-3,3,5,5tetramethylcyclohexylidene]hydroxylamine SMILES: CCOC(=C)C1C(C)(C)CC(CC1(C)C)=NO Predicted Olfactory Receptor Activity: OR52D1 71.900% OR1G1 70.394% 0R52I2 70.392% OR52I1 70.390% OR2Y1 70.378% 3. Commercial name: Hyperflor© IUPAC Name: 2-benzyl-1,3-dioxan-5-one SMILES: O=C1COC(CC2=CC=CC=C2)OC1 Molecular weight: 192.21 g/mol Volume -

Genetic Characterization of Greek Population Isolates Reveals Strong Genetic Drift at Missense and Trait-Associated Variants
ARTICLE Received 22 Apr 2014 | Accepted 22 Sep 2014 | Published 6 Nov 2014 DOI: 10.1038/ncomms6345 OPEN Genetic characterization of Greek population isolates reveals strong genetic drift at missense and trait-associated variants Kalliope Panoutsopoulou1,*, Konstantinos Hatzikotoulas1,*, Dionysia Kiara Xifara2,3, Vincenza Colonna4, Aliki-Eleni Farmaki5, Graham R.S. Ritchie1,6, Lorraine Southam1,2, Arthur Gilly1, Ioanna Tachmazidou1, Segun Fatumo1,7,8, Angela Matchan1, Nigel W. Rayner1,2,9, Ioanna Ntalla5,10, Massimo Mezzavilla1,11, Yuan Chen1, Chrysoula Kiagiadaki12, Eleni Zengini13,14, Vasiliki Mamakou13,15, Antonis Athanasiadis16, Margarita Giannakopoulou17, Vassiliki-Eirini Kariakli5, Rebecca N. Nsubuga18, Alex Karabarinde18, Manjinder Sandhu1,8, Gil McVean2, Chris Tyler-Smith1, Emmanouil Tsafantakis12, Maria Karaleftheri16, Yali Xue1, George Dedoussis5 & Eleftheria Zeggini1 Isolated populations are emerging as a powerful study design in the search for low-frequency and rare variant associations with complex phenotypes. Here we genotype 2,296 samples from two isolated Greek populations, the Pomak villages (HELIC-Pomak) in the North of Greece and the Mylopotamos villages (HELIC-MANOLIS) in Crete. We compare their genomic characteristics to the general Greek population and establish them as genetic isolates. In the MANOLIS cohort, we observe an enrichment of missense variants among the variants that have drifted up in frequency by more than fivefold. In the Pomak cohort, we find novel associations at variants on chr11p15.4 showing large allele frequency increases (from 0.2% in the general Greek population to 4.6% in the isolate) with haematological traits, for example, with mean corpuscular volume (rs7116019, P ¼ 2.3 Â 10 À 26). We replicate this association in a second set of Pomak samples (combined P ¼ 2.0 Â 10 À 36). -

Ncomms-18-33880B
ORE Open Research Exeter TITLE Genome-wide association study identifies genetic loci for self-reported habitual sleep duration supported by accelerometer-derived estimates. AUTHORS Dashti, HS; Jones, SE; Wood, AR; et al. JOURNAL Nature Communications DEPOSITED IN ORE 15 March 2019 This version available at http://hdl.handle.net/10871/36477 COPYRIGHT AND REUSE Open Research Exeter makes this work available in accordance with publisher policies. A NOTE ON VERSIONS The version presented here may differ from the published version. If citing, you are advised to consult the published version for pagination, volume/issue and date of publication NCOMMS-18-33880B Dashti et al. a. Sleep Duration Heritability = 9.8 (0.1)% ) P ( 10 log - Chromosome 1 2 3 4 5 6 7 10 11 14 17 19 C1orf94 PAX8 PCCB BANK1 PAM SCAND3 FOXP2 LOC100131719 HSD17B12 LOC100128215 LOC644157 PIN1 GJB5 LOC100130100 PPP2R3A LCORL PPIP5K2 LOC646160 MAD1L1 GPR26 METT5D1 PRKD1 LOC644172 OLFM2 PDE4B VRK2 STAG1 LOC645174 GIN1 FAM83B CHCHD3 MLLT10 OR2BH1P NOVA1 PER1 DAB1 GALNT3 MSL2 PRKG2 C5ORF30 MEA1 DNAJC1 C11orf63 ADCK1 PFAS DPYD SCN1A BBX, CCSER1 SLC6A3 PPP2R5D 8 SKIDA1 DRD2 FRDAP SMAD5-AS5 20 TTC21B LOC285205 LOC100132531 SMAD5-AS2 PPP1R3B EGR2 PTPRJ RTN1 VAMP2 YWHAB PABPCP2 LOC100128160 LOC285577 SRF PABPC1L LOC100129150 ADO OR4X2 12 FNTB AURKB LOC100133235 FOXP1 FGFR4 CUL7 FGF8 OR4B1 GPX2 ARHGEF15 SLC8A1 IL20RB V27 SLC34A1 DNPH1 KSR2 9 NFKB2 OR4S1 MVK MAX RANGRF MBOAT2 NPM1P17 RGS14 CUL9 ZCCHC7 PITX3 OR4X1 CHURC1 BORCS6 MAP2 ERC2 LMAN2 MRPL2 KCTD10 MRPL41 PSD TRPC6 UBE3B RAB15 CTC1, -

Clinical, Molecular, and Immune Analysis of Dabrafenib-Trametinib
Supplementary Online Content Chen G, McQuade JL, Panka DJ, et al. Clinical, molecular and immune analysis of dabrafenib-trametinib combination treatment for metastatic melanoma that progressed during BRAF inhibitor monotherapy: a phase 2 clinical trial. JAMA Oncology. Published online April 28, 2016. doi:10.1001/jamaoncol.2016.0509. eMethods. eReferences. eTable 1. Clinical efficacy eTable 2. Adverse events eTable 3. Correlation of baseline patient characteristics with treatment outcomes eTable 4. Patient responses and baseline IHC results eFigure 1. Kaplan-Meier analysis of overall survival eFigure 2. Correlation between IHC and RNAseq results eFigure 3. pPRAS40 expression and PFS eFigure 4. Baseline and treatment-induced changes in immune infiltrates eFigure 5. PD-L1 expression eTable 5. Nonsynonymous mutations detected by WES in baseline tumors This supplementary material has been provided by the authors to give readers additional information about their work. © 2016 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 09/30/2021 eMethods Whole exome sequencing Whole exome capture libraries for both tumor and normal samples were constructed using 100ng genomic DNA input and following the protocol as described by Fisher et al.,3 with the following adapter modification: Illumina paired end adapters were replaced with palindromic forked adapters with unique 8 base index sequences embedded within the adapter. In-solution hybrid selection was performed using the Illumina Rapid Capture Exome enrichment kit with 38Mb target territory (29Mb baited). The targeted region includes 98.3% of the intervals in the Refseq exome database. Dual-indexed libraries were pooled into groups of up to 96 samples prior to hybridization.