RBX1/ROC1 Disruption Results in Early Embryonic Lethality Due to Proliferation Failure, Partially Rescued by Simultaneous Loss of P27
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genome-Wide Analysis and Characterization of F-Box Gene Family in Gossypium Hirsutum L Shulin Zhang1,2, Zailong Tian1, Haipeng Li1, Yutao Guo1, Yanqi Zhang1, Jeremy A
Zhang et al. BMC Genomics (2019) 20:993 https://doi.org/10.1186/s12864-019-6280-2 RESEARCH ARTICLE Open Access Genome-wide analysis and characterization of F-box gene family in Gossypium hirsutum L Shulin Zhang1,2, Zailong Tian1, Haipeng Li1, Yutao Guo1, Yanqi Zhang1, Jeremy A. Roberts3, Xuebin Zhang1* and Yuchen Miao1* Abstract Background: F-box proteins are substrate-recognition components of the Skp1-Rbx1-Cul1-F-box protein (SCF) ubiquitin ligases. By selectively targeting the key regulatory proteins or enzymes for ubiquitination and 26S proteasome mediated degradation, F-box proteins play diverse roles in plant growth/development and in the responses of plants to both environmental and endogenous signals. Studies of F-box proteins from the model plant Arabidopsis and from many additional plant species have demonstrated that they belong to a super gene family, and function across almost all aspects of the plant life cycle. However, systematic exploration of F-box family genes in the important fiber crop cotton (Gossypium hirsutum) has not been previously performed. The genome- wide analysis of the cotton F-box gene family is now possible thanks to the completion of several cotton genome sequencing projects. Results: In current study, we first conducted a genome-wide investigation of cotton F-box family genes by reference to the published F-box protein sequences from other plant species. 592 F-box protein encoding genes were identified in the Gossypium hirsutume acc.TM-1 genome and, subsequently, we were able to present their gene structures, chromosomal locations, syntenic relationships with their parent species. In addition, duplication modes analysis showed that cotton F-box genes were distributed to 26 chromosomes, with the maximum number of genes being detected on chromosome 5. -

The Cardiomyocyte Cell Cycle," Novartis Found Symposium 274 (2006): 196-276
DePauw University Scholarly and Creative Work from DePauw University Biology Faculty publications Biology 2006 The ac rdiomyocyte cell cycle Pascal J. Lafontant DePauw University, [email protected] Follow this and additional works at: http://scholarship.depauw.edu/bio_facpubs Part of the Biology Commons Recommended Citation Lafontant, Pascal J. E. and Loren J. Field. "The cardiomyocyte cell cycle," Novartis Found Symposium 274 (2006): 196-276. This Article is brought to you for free and open access by the Biology at Scholarly and Creative Work from DePauw University. It has been accepted for inclusion in Biology Faculty publications by an authorized administrator of Scholarly and Creative Work from DePauw University. For more information, please contact [email protected]. NIH Public Access Author Manuscript Novartis Found Symp. Author manuscript; available in PMC 2009 January 20. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Novartis Found Symp. 2006 ; 274: 196±276. The cardiomyocyte cell cycle Pascal J. E. Lafontant and Loren J. Field From the Wells Center for Pediatric Research and Krannert Institute of Cardiology, Indiana University School of Medicine, Indianapolis, IN 46202-5225. Abstract Many forms of cardiac disease are characterized by cardiomyocyte death due to necrosis, apoptosis and/or oncosis. Recently, the notion of promoting cardiac regeneration as a means to replace damaged heart tissue has engendered considerable interest. One approach to accomplish heart muscle regeneration entails promoting cardiomyocyte cell cycle activity in the surviving myocardium. Genetically modified mice have provided useful model systems to test the efficacy of specific pathways to promote cardiomyocyte proliferation in normal and diseased hearts. -

Expression Profiling of KLF4
Expression Profiling of KLF4 AJCR0000006 Supplemental Data Figure S1. Snapshot of enriched gene sets identified by GSEA in Klf4-null MEFs. Figure S2. Snapshot of enriched gene sets identified by GSEA in wild type MEFs. 98 Am J Cancer Res 2011;1(1):85-97 Table S1: Functional Annotation Clustering of Genes Up-Regulated in Klf4 -Null MEFs ILLUMINA_ID Gene Symbol Gene Name (Description) P -value Fold-Change Cell Cycle 8.00E-03 ILMN_1217331 Mcm6 MINICHROMOSOME MAINTENANCE DEFICIENT 6 40.36 ILMN_2723931 E2f6 E2F TRANSCRIPTION FACTOR 6 26.8 ILMN_2724570 Mapk12 MITOGEN-ACTIVATED PROTEIN KINASE 12 22.19 ILMN_1218470 Cdk2 CYCLIN-DEPENDENT KINASE 2 9.32 ILMN_1234909 Tipin TIMELESS INTERACTING PROTEIN 5.3 ILMN_1212692 Mapk13 SAPK/ERK/KINASE 4 4.96 ILMN_2666690 Cul7 CULLIN 7 2.23 ILMN_2681776 Mapk6 MITOGEN ACTIVATED PROTEIN KINASE 4 2.11 ILMN_2652909 Ddit3 DNA-DAMAGE INDUCIBLE TRANSCRIPT 3 2.07 ILMN_2742152 Gadd45a GROWTH ARREST AND DNA-DAMAGE-INDUCIBLE 45 ALPHA 1.92 ILMN_1212787 Pttg1 PITUITARY TUMOR-TRANSFORMING 1 1.8 ILMN_1216721 Cdk5 CYCLIN-DEPENDENT KINASE 5 1.78 ILMN_1227009 Gas2l1 GROWTH ARREST-SPECIFIC 2 LIKE 1 1.74 ILMN_2663009 Rassf5 RAS ASSOCIATION (RALGDS/AF-6) DOMAIN FAMILY 5 1.64 ILMN_1220454 Anapc13 ANAPHASE PROMOTING COMPLEX SUBUNIT 13 1.61 ILMN_1216213 Incenp INNER CENTROMERE PROTEIN 1.56 ILMN_1256301 Rcc2 REGULATOR OF CHROMOSOME CONDENSATION 2 1.53 Extracellular Matrix 5.80E-06 ILMN_2735184 Col18a1 PROCOLLAGEN, TYPE XVIII, ALPHA 1 51.5 ILMN_1223997 Crtap CARTILAGE ASSOCIATED PROTEIN 32.74 ILMN_2753809 Mmp3 MATRIX METALLOPEPTIDASE -

Neddylation: a Novel Modulator of the Tumor Microenvironment Lisha Zhou1,2*†, Yanyu Jiang3†, Qin Luo1, Lihui Li1 and Lijun Jia1*
Zhou et al. Molecular Cancer (2019) 18:77 https://doi.org/10.1186/s12943-019-0979-1 REVIEW Open Access Neddylation: a novel modulator of the tumor microenvironment Lisha Zhou1,2*†, Yanyu Jiang3†, Qin Luo1, Lihui Li1 and Lijun Jia1* Abstract Neddylation, a post-translational modification that adds an ubiquitin-like protein NEDD8 to substrate proteins, modulates many important biological processes, including tumorigenesis. The process of protein neddylation is overactivated in multiple human cancers, providing a sound rationale for its targeting as an attractive anticancer therapeutic strategy, as evidence by the development of NEDD8-activating enzyme (NAE) inhibitor MLN4924 (also known as pevonedistat). Neddylation inhibition by MLN4924 exerts significantly anticancer effects mainly by triggering cell apoptosis, senescence and autophagy. Recently, intensive evidences reveal that inhibition of neddylation pathway, in addition to acting on tumor cells, also influences the functions of multiple important components of the tumor microenvironment (TME), including immune cells, cancer-associated fibroblasts (CAFs), cancer-associated endothelial cells (CAEs) and some factors, all of which are crucial for tumorigenesis. Here, we briefly summarize the latest progresses in this field to clarify the roles of neddylation in the TME, thus highlighting the overall anticancer efficacy of neddylaton inhibition. Keywords: Neddylation, Tumor microenvironment, Tumor-derived factors, Cancer-associated fibroblasts, Cancer- associated endothelial cells, Immune cells Introduction Overall, binding of NEDD8 molecules to target proteins Neddylation is a reversible covalent conjugation of an can affect their stability, subcellular localization, conform- ubiquitin-like molecule NEDD8 (neuronal precursor ation and function [4]. The best-characterized substrates cell-expressed developmentally down-regulated protein of neddylation are the cullin subunits of Cullin-RING li- 8) to a lysine residue of the substrate protein [1, 2]. -
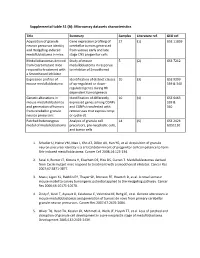
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Report 99517
Comprehensive Skeletal Dysplasias and Disorders Panel Plus REFERRING HEALTHCARE PROFESSIONAL NAME HOSPITAL PATIENT NAME DOB AGE GENDER ORDER ID 6 PRIMARY SAMPLE TYPE SAMPLE COLLECTION DATE CUSTOMER SAMPLE ID SUMMARY OF RESULTS PRIMARY FINDINGS The patient is heterozygous for TRPV4 c.2396C>T, p.(Pro799Leu), which is classified as pathogenic. Del/Dup (CNV) analysis Negative for explaining the patient’s phenotype. PRIMARY FINDINGS: SEQUENCE ALTERATIONS GENE TRANSCRIPT NOMENCLATURE GENOTYPE CONSEQUENCE INHERITANCE CLASSIFICATION TRPV4 NM_021625.4 c.2396C>T, p.(Pro799Leu) HET missense_variant AD Pathogenic ID ASSEMBLY POS REF/ALT GRCh37/hg19 12:110222183 G/A PHENOTYPE Brachyolmia (autosomal dominant type), Charcot-Marie-Tooth disease, Familial Digital arthropathy with brachydactyly, gnomAD AC/AN POLYPHEN SIFT MUTTASTER Hereditary motor and sensory neuropathy, 0/0 possibly damaging deleterious disease causing Metatropic dysplasia, Parastremmatic dwarfism, Spinal muscular atrophy, Spondyloepiphyseal dysplasia Maroteaux type, Spondylometaphyseal dysplasia Kozlowski type SEQUENCING PERFORMANCE METRICS PANEL GENES EXONS / REGIONS BASES BASES > 20X MEDIAN PERCENT COVERAGE > 20X Comprehensive Skeletal Dysplasias and 252 4053 816901 812241 199 99.43 Disorders Panel Blueprint Genetics Oy, Keilaranta 16 A-B, 02150 Espoo, Finland VAT number: FI22307900, CLIA ID Number: 99D2092375, CAP Number: 9257331 TARGET REGION AND GENE LIST The Blueprint Genetics Comprehensive Skeletal Dysplasias and Disorders Panel (version 4, Oct 19, 2019) Plus Analysis includes sequence -

Expression and Purification of Functional Recombinant CUL2
www.nature.com/scientificreports OPEN Expression and purifcation of functional recombinant CUL2•RBX1 from E. coli Stephanie Diaz1, Lihong Li1,2, Kankan Wang1 & Xing Liu1,2* Cullin-2 (CUL2) based cullin-RING ligases (CRL2s) comprise a family of ubiquitin E3 ligases that exist only in multi-cellular organisms and are crucial for cellular processes such as embryogenesis and viral pathogenesis. CUL2 is the scafold protein that binds one of the interchangeable substrate receptor modules, which consists of adaptor proteins and the substrate receptor protein. The VHL protein is a substrate receptor known to target hypoxia-inducible factor α (HIF1α) for ubiquitination and degradation. Because of its critical role in the ubiquitination of important cellular factors such as HIF1α, CRL2s have been investigated for their biological functions and the development of novel therapeutics against diseases. Given the importance of CRL2s in biological and biomedical research, methods that efciently produce functional CUL2 proteins will greatly facilitate studies on the mechanism and regulation of CRL2s. Here, we report two cost-efective systems for the expression and purifcation of recombinant human CUL2 from E. coli cells. The purifed CUL2 proteins were ~ 95% pure, could bind their substrate receptor modules, and were enzymatically active in transferring ubiquitin or ubiquitin-like protein to the corresponding substrate in in vitro assays. The presented methodological advancements will help advance research in CRL2 function and regulation. Protein turnover is a cellular regulatory system defned by the continuous synthesis and decomposition of specifc proteins to maintain the integrity of optimally functioning proteins 1,2. Abnormalities during protein turnover, specifcally during protein degradation, ofen result in human diseases such as cystic fbrosis and liposarcoma. -

Disruption of the Coordination Between Host Circadian Rhythms and Malaria
bioRxiv preprint doi: https://doi.org/10.1101/791046; this version posted October 2, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Disruption of the coordination between host circadian rhythms and malaria parasite development alters the duration of the intraerythrocytic cycle Authors Amit K. Subudhi1, Aidan J. O’Donnell2†, Abhinay Ramaprasad1†, Hussein M. Abkallo3, Abhinav Kaushik1, Hifzur R. Ansari1, Alyaa M Abdel-Haleem1, Fathia Ben Rached1, Osamu Kaneko4, Richard Culleton3,*, Sarah E. Reece2,*, Arnab Pain1,5,6* 1Pathogen Genomics Group, BESE Division, King Abdullah University of Science and Technology (KAUST), Thuwal, 23955-6900, Kingdom of Saudi Arabia 2Institute of Evolutionary Biology, and Institute of Immunology and Infection Research, University of Edinburgh, Edinburgh EH9 3FL, UK 3Malaria Unit, Department of Pathology, Institute of Tropical Medicine (NEKKEN), Nagasaki University, 1-12-4 Sakamoto, Nagasaki 852-8523, Japan 4Department of Protozoology, Institute of Tropical Medicine (NEKKEN), Nagasaki University, 1-12-4 Sakamoto, Nagasaki, 852-8523, Japan 5Center for Zoonosis Control, Global Institution for Collaborative Research and Education (GI-CoRE); Hokkaido University, N20 W10 Kita-ku, Sapporo, 001-0020 Japan 6Lead Contact Current address: Computational Bioscience Research Center, King Abdullah University of Science and Technology (KAUST), Thuwal, 23955-6900, Kingdom of Saudi Arabia. †Contributed equally 1 bioRxiv preprint doi: https://doi.org/10.1101/791046; this version posted October 2, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Anti-DCUN1D1 Antibody (ARG65153)
Product datasheet [email protected] ARG65153 Package: 100 μg anti-DCUN1D1 antibody Store at: -20°C Summary Product Description Goat Polyclonal antibody recognizes DCUN1D1 Tested Reactivity Hu Predict Reactivity Ms, Cow Tested Application IHC-P, WB Host Goat Clonality Polyclonal Isotype IgG Target Name DCUN1D1 Antigen Species Human Immunogen C-RPQIAGTKSTT Conjugation Un-conjugated Alternate Names SCRO; RP42; Defective in cullin neddylation protein 1-like protein 1; Squamous cell carcinoma-related oncogene; DCUN1 domain-containing protein 1; SCCRO; DCN1-like protein 1; DCUN1L1; DCNL1; Tes3 Application Instructions Application table Application Dilution IHC-P 5 - 10 µg/ml WB 0.3 - 1 µg/ml Application Note IHC-P: Antigen Retrieval: Steam tissue section in Citrate buffer (pH 6.0). WB: Recommend incubate at RT for 1h. * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Calculated Mw 30 kDa Properties Form Liquid Purification Purified from goat serum by ammonium sulphate precipitation followed by antigen affinity chromatography using the immunizing peptide. Buffer Tris saline (pH 7.3), 0.02% Sodium azide and 0.5% BSA Preservative 0.02% Sodium azide Stabilizer 0.5% BSA Concentration 0.5 mg/ml www.arigobio.com 1/2 Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C or below. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. Note For laboratory research only, not for drug, diagnostic or other use. -

A Graph-Theoretic Approach to Model Genomic Data and Identify Biological Modules Asscociated with Cancer Outcomes
A Graph-Theoretic Approach to Model Genomic Data and Identify Biological Modules Asscociated with Cancer Outcomes Deanna Petrochilos A dissertation presented in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Washington 2013 Reading Committee: Neil Abernethy, Chair John Gennari, Ali Shojaie Program Authorized to Offer Degree: Biomedical Informatics and Health Education UMI Number: 3588836 All rights reserved INFORMATION TO ALL USERS The quality of this reproduction is dependent upon the quality of the copy submitted. In the unlikely event that the author did not send a complete manuscript and there are missing pages, these will be noted. Also, if material had to be removed, a note will indicate the deletion. UMI 3588836 Published by ProQuest LLC (2013). Copyright in the Dissertation held by the Author. Microform Edition © ProQuest LLC. All rights reserved. This work is protected against unauthorized copying under Title 17, United States Code ProQuest LLC. 789 East Eisenhower Parkway P.O. Box 1346 Ann Arbor, MI 48106 - 1346 ©Copyright 2013 Deanna Petrochilos University of Washington Abstract Using Graph-Based Methods to Integrate and Analyze Cancer Genomic Data Deanna Petrochilos Chair of the Supervisory Committee: Assistant Professor Neil Abernethy Biomedical Informatics and Health Education Studies of the genetic basis of complex disease present statistical and methodological challenges in the discovery of reliable and high-confidence genes that reveal biological phenomena underlying the etiology of disease or gene signatures prognostic of disease outcomes. This dissertation examines the capacity of graph-theoretical methods to model and analyze genomic information and thus facilitate using prior knowledge to create a more discrete and functionally relevant feature space. -

Cytoplasmic Localized Ubiquitin Ligase Cullin 7 Binds to P53 and Promotes Cell Growth by Antagonizing P53 Function
Oncogene (2006) 25, 4534–4548 & 2006 Nature Publishing Group All rights reserved 0950-9232/06 $30.00 www.nature.com/onc ORIGINAL ARTICLE Cytoplasmic localized ubiquitin ligase cullin 7 binds to p53 and promotes cell growth by antagonizing p53 function P Andrews1,2,YJHe1 and Y Xiong1,2,3 1Lineberger Comprehensive Cancer Center, University of North Carolina, Chapel Hill, NC, USA; 2Department of Biochemistry and Biophysics, University of North Carolina, Chapel Hill, NC, USA and 3Program in Molecular Biology and Biotechnology, University of North Carolina, Chapel Hill, NC, USA Cullins are a family of evolutionarily conserved proteins monomeric manner (monoubiquitination) to cause that bind to the small RING fingerprotein,ROC1, to conformational change to the substrate or in a constitute potentially a large number of distinct E3 polyubiquitin chain which signals the substrate to be ubiquitin ligases. CUL7 mediates an essential function degraded by the 26S proteasome. Both ubiquitin formouse embryodevelopment and has been linked with activating and conjugating enzymes are well character- cell transformation by its physical association with the ized and contain highly conserved functional domains. SV40 large T antigen. We report here that, like its closely The identity and mechanism of E3 ubiquitin ligases, on related homolog PARC, CUL7 is localized predominantly the other hand, has been elusive and long postulated as in the cytoplasm and binds directly to p53. In contrast to an activity responsible for both recognizing substrates PARC, however, CUL7, even when overexpressed, did not and for catalysing polyubiquitin chain formation sequesterp53 in the cytoplasm. We have identified a (Hershko and Ciechanover, 1998). sequence in the N-terminal region of CUL7 that is highly Currently, two major families of E3 ligases have been conserved in PARC and a sequence spanning the described. -

Cullin 7 in Tumor Development: a Novel Potential Anti-Cancer Target
572 Neoplasma 2021; 68(3): 572–579 doi:10.4149/neo_2021_201207N1324 Cullin 7 in tumor development: a novel potential anti-cancer target Yumiao LI1,#, Aiming ZANG1,#, Jiejun FU2, Youchao JIA1,* 1Department of Medical Oncology, Affiliated Hospital of Hebei University, Hebei Key Laboratory of Cancer Radiotherapy and Chemotherapy, Baoding, Hebei, China; 2Department of Cell Biology and Genetics, Guangxi Medical University, Key Laboratory of Longevity and Aging-related Diseases of Chinese Ministry of Education, Center for Translational Medicine and School of Preclinical Medicine, Nanning, Guangxi, China *Correspondence: [email protected] #Contributed equally to this work. Received December 7, 2020 / Accepted January 25, 2021 As a core scaffold protein, Cullin 7 (Cul7) forms Skp1-Cullin-F-box (SCF) E3 ubiquitin ligase complexes with the regulator of cullins-1 (ROC1), S-phase kinase associated protein 1 (Skp1) and F-Box, and WD repeat domain containing 8 (Fbxw8). Alternatively, Cul7 can form a CRL7SMU1 complex with suppressor of Mec-8 and Unc-52 protein homolog (SMU1), damage-specific DNA binding protein 1 (DDB1), and ring finger protein 40 (RNF40), to promote cell growth. The mutations of Cul7 cause the 3-M dwarf syndrome, indicating Cul7 plays an important role in growth and development in humans and mice. Moreover, Cul7 regulates cell transformation, tumor protein p53 activity, cell senescence, and apoptosis, mutations in Cul7 are also involved in the development of tumors, indicating the characteristics of an oncogene. Cul7 is highly expressed in breast cancer, lung cancer, hepatocellular carcinoma, pancreatic cancer, ovarian cancer, and other malignant tumors where Cul7 promotes tumor development, cell transformation, and cell survival by regulating complex signaling pathways associated with protein degradation.