Report 99517
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Clinical and Genetic Characterisation of Hereditary Motor Neuropathies
Clinical and genetic characterisation of hereditary motor neuropathies Boglarka Bansagi Institute of Genetic Medicine January 2017 A Thesis submitted for the degree of Doctor of Philosophy at Newcastle University 'I dare do all that may become a man; who dares do more is none.' (Shakespeare) „Szívet vagy mundért cserélhet az is, ki házat, hazát holtig nem cserél, de hű maradhat idegenben is, kiben népe mostoha sorsa él.” (Tollas Tibor) Author’s declaration This Thesis is submitted to Newcastle University for the degree of Doctor of Philosophy at Newcastle University. I, Boglarka Bansagi, confirm that the work presented in this Thesis is my own. Where information has been derived from other sources, it has been indicated in the Thesis. I can confirm that none of the material offered in this Thesis has been previously submitted by me for a degree or qualification in this or any other university. Abstract Inherited peripheral neuropathies or Charcot-Marie-Tooth disease (CMT) are common neuromuscular conditions, characterised by distal motor atrophy and weakness with variable range of sensory impairment and classified according to demyelinating (CMT1) or axonal (CMT2) pathology. The number of genes causing CMT has rapidly increased due to improved genetic testing technology, even though gene identification has remained challenging in some subgroups of CMT. Hereditary motor neuropathies (HMN) encompass heterogeneous groups of disorders caused by motor axon and neuron pathology. The distal hereditary motor neuropathies (dHMN) are rare length-dependent conditions, which show significant clinical and genetic overlap with motor neuron diseases. Several (>30) causative genes have been identified for ~20% of dHMN patients, which predicts extreme genetic heterogeneity in this group. -

The Cardiomyocyte Cell Cycle," Novartis Found Symposium 274 (2006): 196-276
DePauw University Scholarly and Creative Work from DePauw University Biology Faculty publications Biology 2006 The ac rdiomyocyte cell cycle Pascal J. Lafontant DePauw University, [email protected] Follow this and additional works at: http://scholarship.depauw.edu/bio_facpubs Part of the Biology Commons Recommended Citation Lafontant, Pascal J. E. and Loren J. Field. "The cardiomyocyte cell cycle," Novartis Found Symposium 274 (2006): 196-276. This Article is brought to you for free and open access by the Biology at Scholarly and Creative Work from DePauw University. It has been accepted for inclusion in Biology Faculty publications by an authorized administrator of Scholarly and Creative Work from DePauw University. For more information, please contact [email protected]. NIH Public Access Author Manuscript Novartis Found Symp. Author manuscript; available in PMC 2009 January 20. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Novartis Found Symp. 2006 ; 274: 196±276. The cardiomyocyte cell cycle Pascal J. E. Lafontant and Loren J. Field From the Wells Center for Pediatric Research and Krannert Institute of Cardiology, Indiana University School of Medicine, Indianapolis, IN 46202-5225. Abstract Many forms of cardiac disease are characterized by cardiomyocyte death due to necrosis, apoptosis and/or oncosis. Recently, the notion of promoting cardiac regeneration as a means to replace damaged heart tissue has engendered considerable interest. One approach to accomplish heart muscle regeneration entails promoting cardiomyocyte cell cycle activity in the surviving myocardium. Genetically modified mice have provided useful model systems to test the efficacy of specific pathways to promote cardiomyocyte proliferation in normal and diseased hearts. -

Expression Profiling of KLF4
Expression Profiling of KLF4 AJCR0000006 Supplemental Data Figure S1. Snapshot of enriched gene sets identified by GSEA in Klf4-null MEFs. Figure S2. Snapshot of enriched gene sets identified by GSEA in wild type MEFs. 98 Am J Cancer Res 2011;1(1):85-97 Table S1: Functional Annotation Clustering of Genes Up-Regulated in Klf4 -Null MEFs ILLUMINA_ID Gene Symbol Gene Name (Description) P -value Fold-Change Cell Cycle 8.00E-03 ILMN_1217331 Mcm6 MINICHROMOSOME MAINTENANCE DEFICIENT 6 40.36 ILMN_2723931 E2f6 E2F TRANSCRIPTION FACTOR 6 26.8 ILMN_2724570 Mapk12 MITOGEN-ACTIVATED PROTEIN KINASE 12 22.19 ILMN_1218470 Cdk2 CYCLIN-DEPENDENT KINASE 2 9.32 ILMN_1234909 Tipin TIMELESS INTERACTING PROTEIN 5.3 ILMN_1212692 Mapk13 SAPK/ERK/KINASE 4 4.96 ILMN_2666690 Cul7 CULLIN 7 2.23 ILMN_2681776 Mapk6 MITOGEN ACTIVATED PROTEIN KINASE 4 2.11 ILMN_2652909 Ddit3 DNA-DAMAGE INDUCIBLE TRANSCRIPT 3 2.07 ILMN_2742152 Gadd45a GROWTH ARREST AND DNA-DAMAGE-INDUCIBLE 45 ALPHA 1.92 ILMN_1212787 Pttg1 PITUITARY TUMOR-TRANSFORMING 1 1.8 ILMN_1216721 Cdk5 CYCLIN-DEPENDENT KINASE 5 1.78 ILMN_1227009 Gas2l1 GROWTH ARREST-SPECIFIC 2 LIKE 1 1.74 ILMN_2663009 Rassf5 RAS ASSOCIATION (RALGDS/AF-6) DOMAIN FAMILY 5 1.64 ILMN_1220454 Anapc13 ANAPHASE PROMOTING COMPLEX SUBUNIT 13 1.61 ILMN_1216213 Incenp INNER CENTROMERE PROTEIN 1.56 ILMN_1256301 Rcc2 REGULATOR OF CHROMOSOME CONDENSATION 2 1.53 Extracellular Matrix 5.80E-06 ILMN_2735184 Col18a1 PROCOLLAGEN, TYPE XVIII, ALPHA 1 51.5 ILMN_1223997 Crtap CARTILAGE ASSOCIATED PROTEIN 32.74 ILMN_2753809 Mmp3 MATRIX METALLOPEPTIDASE -
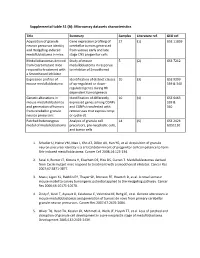
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Sleeping Beauty Transposon Mutagenesis Identifies Genes That
Sleeping Beauty transposon mutagenesis identifies PNAS PLUS genes that cooperate with mutant Smad4 in gastric cancer development Haruna Takedaa,b, Alistair G. Rustc,d, Jerrold M. Warda, Christopher Chin Kuan Yewa, Nancy A. Jenkinsa,e, and Neal G. Copelanda,e,1 aDivision of Genomics and Genetics, Institute of Molecular and Cell Biology, Agency for Science, Technology and Research, Singapore 138673; bDepartment of Pathology, School of Medicine, Kanazawa Medical University, Ishikawa 920-0293, Japan; cExperimental Cancer Genetics, Wellcome Trust Sanger Institute, Cambridge CB10 1HH, United Kingdom; dTumour Profiling Unit, The Institute of Cancer Research, Chester Beatty Laboratories, London SW3 6JB, United Kingdom; and eCancer Research Program, Houston Methodist Research Institute, Houston, TX 77030 Contributed by Neal G. Copeland, February 27, 2016 (sent for review October 15, 2015; reviewed by Yoshiaki Ito and David A. Largaespada) Mutations in SMAD4 predispose to the development of gastroin- animal models that mimic human GC, researchers have infected testinal cancer, which is the third leading cause of cancer-related mice with H. pylori and then, treated them with carcinogens. They deaths. To identify genes driving gastric cancer (GC) development, have also used genetic engineering to develop a variety of trans- we performed a Sleeping Beauty (SB) transposon mutagenesis genic and KO mouse models of GC (10). Smad4 KO mice are one + − screen in the stomach of Smad4 / mutant mice. This screen iden- GC model that has been of particular interest to us (11, 12). tified 59 candidate GC trunk drivers and a much larger number of Heterozygous Smad4 KO mice develop polyps in the pyloric re- candidate GC progression genes. -

Orphanet Report Series Rare Diseases Collection
Marche des Maladies Rares – Alliance Maladies Rares Orphanet Report Series Rare Diseases collection DecemberOctober 2013 2009 List of rare diseases and synonyms Listed in alphabetical order www.orpha.net 20102206 Rare diseases listed in alphabetical order ORPHA ORPHA ORPHA Disease name Disease name Disease name Number Number Number 289157 1-alpha-hydroxylase deficiency 309127 3-hydroxyacyl-CoA dehydrogenase 228384 5q14.3 microdeletion syndrome deficiency 293948 1p21.3 microdeletion syndrome 314655 5q31.3 microdeletion syndrome 939 3-hydroxyisobutyric aciduria 1606 1p36 deletion syndrome 228415 5q35 microduplication syndrome 2616 3M syndrome 250989 1q21.1 microdeletion syndrome 96125 6p subtelomeric deletion syndrome 2616 3-M syndrome 250994 1q21.1 microduplication syndrome 251046 6p22 microdeletion syndrome 293843 3MC syndrome 250999 1q41q42 microdeletion syndrome 96125 6p25 microdeletion syndrome 6 3-methylcrotonylglycinuria 250999 1q41-q42 microdeletion syndrome 99135 6-phosphogluconate dehydrogenase 67046 3-methylglutaconic aciduria type 1 deficiency 238769 1q44 microdeletion syndrome 111 3-methylglutaconic aciduria type 2 13 6-pyruvoyl-tetrahydropterin synthase 976 2,8 dihydroxyadenine urolithiasis deficiency 67047 3-methylglutaconic aciduria type 3 869 2A syndrome 75857 6q terminal deletion 67048 3-methylglutaconic aciduria type 4 79154 2-aminoadipic 2-oxoadipic aciduria 171829 6q16 deletion syndrome 66634 3-methylglutaconic aciduria type 5 19 2-hydroxyglutaric acidemia 251056 6q25 microdeletion syndrome 352328 3-methylglutaconic -

A Yeast Phenomic Model for the Influence of Warburg Metabolism on Genetic Buffering of Doxorubicin Sean M
Santos and Hartman Cancer & Metabolism (2019) 7:9 https://doi.org/10.1186/s40170-019-0201-3 RESEARCH Open Access A yeast phenomic model for the influence of Warburg metabolism on genetic buffering of doxorubicin Sean M. Santos and John L. Hartman IV* Abstract Background: The influence of the Warburg phenomenon on chemotherapy response is unknown. Saccharomyces cerevisiae mimics the Warburg effect, repressing respiration in the presence of adequate glucose. Yeast phenomic experiments were conducted to assess potential influences of Warburg metabolism on gene-drug interaction underlying the cellular response to doxorubicin. Homologous genes from yeast phenomic and cancer pharmacogenomics data were analyzed to infer evolutionary conservation of gene-drug interaction and predict therapeutic relevance. Methods: Cell proliferation phenotypes (CPPs) of the yeast gene knockout/knockdown library were measured by quantitative high-throughput cell array phenotyping (Q-HTCP), treating with escalating doxorubicin concentrations under conditions of respiratory or glycolytic metabolism. Doxorubicin-gene interaction was quantified by departure of CPPs observed for the doxorubicin-treated mutant strain from that expected based on an interaction model. Recursive expectation-maximization clustering (REMc) and Gene Ontology (GO)-based analyses of interactions identified functional biological modules that differentially buffer or promote doxorubicin cytotoxicity with respect to Warburg metabolism. Yeast phenomic and cancer pharmacogenomics data were integrated to predict differential gene expression causally influencing doxorubicin anti-tumor efficacy. Results: Yeast compromised for genes functioning in chromatin organization, and several other cellular processes are more resistant to doxorubicin under glycolytic conditions. Thus, the Warburg transition appears to alleviate requirements for cellular functions that buffer doxorubicin cytotoxicity in a respiratory context. -

A Set of Regulatory Genes Co-Expressed in Embryonic Human Brain Is Implicated in Disrupted Speech Development
Molecular Psychiatry https://doi.org/10.1038/s41380-018-0020-x ARTICLE A set of regulatory genes co-expressed in embryonic human brain is implicated in disrupted speech development 1 1 1 2 3 Else Eising ● Amaia Carrion-Castillo ● Arianna Vino ● Edythe A. Strand ● Kathy J. Jakielski ● 4,5 6 7 8 9 Thomas S. Scerri ● Michael S. Hildebrand ● Richard Webster ● Alan Ma ● Bernard Mazoyer ● 1,10 4,5 6,11 6,12 13 Clyde Francks ● Melanie Bahlo ● Ingrid E. Scheffer ● Angela T. Morgan ● Lawrence D. Shriberg ● Simon E. Fisher 1,10 Received: 22 September 2017 / Revised: 3 December 2017 / Accepted: 2 January 2018 © The Author(s) 2018. This article is published with open access Abstract Genetic investigations of people with impaired development of spoken language provide windows into key aspects of human biology. Over 15 years after FOXP2 was identified, most speech and language impairments remain unexplained at the molecular level. We sequenced whole genomes of nineteen unrelated individuals diagnosed with childhood apraxia of speech, a rare disorder enriched for causative mutations of large effect. Where DNA was available from unaffected parents, CHD3 SETD1A WDR5 fi 1234567890();,: we discovered de novo mutations, implicating genes, including , and . In other probands, we identi ed novel loss-of-function variants affecting KAT6A, SETBP1, ZFHX4, TNRC6B and MKL2, regulatory genes with links to neurodevelopment. Several of the new candidates interact with each other or with known speech-related genes. Moreover, they show significant clustering within a single co-expression module of genes highly expressed during early human brain development. This study highlights gene regulatory pathways in the developing brain that may contribute to acquisition of proficient speech. -
Blueprint Genetics Comprehensive Growth Disorders / Skeletal
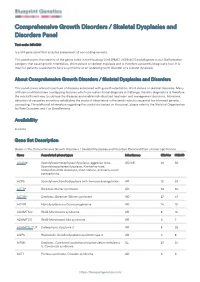
Comprehensive Growth Disorders / Skeletal Dysplasias and Disorders Panel Test code: MA4301 Is a 374 gene panel that includes assessment of non-coding variants. This panel covers the majority of the genes listed in the Nosology 2015 (PMID: 26394607) and all genes in our Malformation category that cause growth retardation, short stature or skeletal dysplasia and is therefore a powerful diagnostic tool. It is ideal for patients suspected to have a syndromic or an isolated growth disorder or a skeletal dysplasia. About Comprehensive Growth Disorders / Skeletal Dysplasias and Disorders This panel covers a broad spectrum of diseases associated with growth retardation, short stature or skeletal dysplasia. Many of these conditions have overlapping features which can make clinical diagnosis a challenge. Genetic diagnostics is therefore the most efficient way to subtype the diseases and enable individualized treatment and management decisions. Moreover, detection of causative mutations establishes the mode of inheritance in the family which is essential for informed genetic counseling. For additional information regarding the conditions tested on this panel, please refer to the National Organization for Rare Disorders and / or GeneReviews. Availability 4 weeks Gene Set Description Genes in the Comprehensive Growth Disorders / Skeletal Dysplasias and Disorders Panel and their clinical significance Gene Associated phenotypes Inheritance ClinVar HGMD ACAN# Spondyloepimetaphyseal dysplasia, aggrecan type, AD/AR 20 56 Spondyloepiphyseal dysplasia, Kimberley -

Blueprint Genetics Comprehensive Skeletal Dysplasias and Disorders
Comprehensive Skeletal Dysplasias and Disorders Panel Test code: MA3301 Is a 251 gene panel that includes assessment of non-coding variants. Is ideal for patients with a clinical suspicion of disorders involving the skeletal system. About Comprehensive Skeletal Dysplasias and Disorders This panel covers a broad spectrum of skeletal disorders including common and rare skeletal dysplasias (eg. achondroplasia, COL2A1 related dysplasias, diastrophic dysplasia, various types of spondylo-metaphyseal dysplasias), various ciliopathies with skeletal involvement (eg. short rib-polydactylies, asphyxiating thoracic dysplasia dysplasias and Ellis-van Creveld syndrome), various subtypes of osteogenesis imperfecta, campomelic dysplasia, slender bone dysplasias, dysplasias with multiple joint dislocations, chondrodysplasia punctata group of disorders, neonatal osteosclerotic dysplasias, osteopetrosis and related disorders, abnormal mineralization group of disorders (eg hypopohosphatasia), osteolysis group of disorders, disorders with disorganized development of skeletal components, overgrowth syndromes with skeletal involvement, craniosynostosis syndromes, dysostoses with predominant craniofacial involvement, dysostoses with predominant vertebral involvement, patellar dysostoses, brachydactylies, some disorders with limb hypoplasia-reduction defects, ectrodactyly with and without other manifestations, polydactyly-syndactyly-triphalangism group of disorders, and disorders with defects in joint formation and synostoses. Availability 4 weeks Gene Set Description -

Oncoscore: a Novel, Internet-Based Tool to Assess the Oncogenic Potential of Genes
www.nature.com/scientificreports OPEN OncoScore: a novel, Internet- based tool to assess the oncogenic potential of genes Received: 06 July 2016 Rocco Piazza1, Daniele Ramazzotti2, Roberta Spinelli1, Alessandra Pirola3, Luca De Sano4, Accepted: 15 March 2017 Pierangelo Ferrari3, Vera Magistroni1, Nicoletta Cordani1, Nitesh Sharma5 & Published: 07 April 2017 Carlo Gambacorti-Passerini1 The complicated, evolving landscape of cancer mutations poses a formidable challenge to identify cancer genes among the large lists of mutations typically generated in NGS experiments. The ability to prioritize these variants is therefore of paramount importance. To address this issue we developed OncoScore, a text-mining tool that ranks genes according to their association with cancer, based on available biomedical literature. Receiver operating characteristic curve and the area under the curve (AUC) metrics on manually curated datasets confirmed the excellent discriminating capability of OncoScore (OncoScore cut-off threshold = 21.09; AUC = 90.3%, 95% CI: 88.1–92.5%), indicating that OncoScore provides useful results in cases where an efficient prioritization of cancer-associated genes is needed. The huge amount of data emerging from NGS projects is bringing a revolution in molecular medicine, leading to the discovery of a large number of new somatic alterations that are associated with the onset and/or progression of cancer. However, researchers are facing a formidable challenge in prioritizing cancer genes among the variants generated by NGS experiments. Despite the development of a significant number of tools devoted to cancer driver prediction, limited effort has been dedicated to tools able to generate a gene-centered Oncogenic Score based on the evidence already available in the scientific literature. -

Title: a Yeast Phenomic Model for the Influence of Warburg Metabolism on Genetic
bioRxiv preprint doi: https://doi.org/10.1101/517490; this version posted January 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 1 Title Page: 2 3 Title: A yeast phenomic model for the influence of Warburg metabolism on genetic 4 buffering of doxorubicin 5 6 Authors: Sean M. Santos1 and John L. Hartman IV1 7 1. University of Alabama at Birmingham, Department of Genetics, Birmingham, AL 8 Email: [email protected], [email protected] 9 Corresponding author: [email protected] 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 1 bioRxiv preprint doi: https://doi.org/10.1101/517490; this version posted January 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 26 Abstract: 27 Background: 28 Saccharomyces cerevisiae represses respiration in the presence of adequate glucose, 29 mimicking the Warburg effect, termed aerobic glycolysis. We conducted yeast phenomic 30 experiments to characterize differential doxorubicin-gene interaction, in the context of 31 respiration vs. glycolysis. The resulting systems level biology about doxorubicin 32 cytotoxicity, including the influence of the Warburg effect, was integrated with cancer 33 pharmacogenomics data to identify potentially causal correlations between differential 34 gene expression and anti-cancer efficacy.