Signal Transducer and Activator of Transcription 3-Mediated Regulation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Table 2. Significant
Table 2. Significant (Q < 0.05 and |d | > 0.5) transcripts from the meta-analysis Gene Chr Mb Gene Name Affy ProbeSet cDNA_IDs d HAP/LAP d HAP/LAP d d IS Average d Ztest P values Q-value Symbol ID (study #5) 1 2 STS B2m 2 122 beta-2 microglobulin 1452428_a_at AI848245 1.75334941 4 3.2 4 3.2316485 1.07398E-09 5.69E-08 Man2b1 8 84.4 mannosidase 2, alpha B1 1416340_a_at H4049B01 3.75722111 3.87309653 2.1 1.6 2.84852656 5.32443E-07 1.58E-05 1110032A03Rik 9 50.9 RIKEN cDNA 1110032A03 gene 1417211_a_at H4035E05 4 1.66015788 4 1.7 2.82772795 2.94266E-05 0.000527 NA 9 48.5 --- 1456111_at 3.43701477 1.85785922 4 2 2.8237185 9.97969E-08 3.48E-06 Scn4b 9 45.3 Sodium channel, type IV, beta 1434008_at AI844796 3.79536664 1.63774235 3.3 2.3 2.75319499 1.48057E-08 6.21E-07 polypeptide Gadd45gip1 8 84.1 RIKEN cDNA 2310040G17 gene 1417619_at 4 3.38875643 1.4 2 2.69163229 8.84279E-06 0.0001904 BC056474 15 12.1 Mus musculus cDNA clone 1424117_at H3030A06 3.95752801 2.42838452 1.9 2.2 2.62132809 1.3344E-08 5.66E-07 MGC:67360 IMAGE:6823629, complete cds NA 4 153 guanine nucleotide binding protein, 1454696_at -3.46081884 -4 -1.3 -1.6 -2.6026947 8.58458E-05 0.0012617 beta 1 Gnb1 4 153 guanine nucleotide binding protein, 1417432_a_at H3094D02 -3.13334396 -4 -1.6 -1.7 -2.5946297 1.04542E-05 0.0002202 beta 1 Gadd45gip1 8 84.1 RAD23a homolog (S. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Uncovering Ubiquitin and Ubiquitin-Like Signaling Networks Alfred C
REVIEW pubs.acs.org/CR Uncovering Ubiquitin and Ubiquitin-like Signaling Networks Alfred C. O. Vertegaal* Department of Molecular Cell Biology, Leiden University Medical Center, Albinusdreef 2, 2333 ZA Leiden, The Netherlands CONTENTS 8. Crosstalk between Post-Translational Modifications 7934 1. Introduction 7923 8.1. Crosstalk between Phosphorylation and 1.1. Ubiquitin and Ubiquitin-like Proteins 7924 Ubiquitylation 7934 1.2. Quantitative Proteomics 7924 8.2. Phosphorylation-Dependent SUMOylation 7935 8.3. Competition between Different Lysine 1.3. Setting the Scenery: Mass Spectrometry Modifications 7935 Based Investigation of Phosphorylation 8.4. Crosstalk between SUMOylation and the and Acetylation 7925 UbiquitinÀProteasome System 7935 2. Ubiquitin and Ubiquitin-like Protein Purification 9. Conclusions and Future Perspectives 7935 Approaches 7925 Author Information 7935 2.1. Epitope-Tagged Ubiquitin and Ubiquitin-like Biography 7935 Proteins 7925 Acknowledgment 7936 2.2. Traps Based on Ubiquitin- and Ubiquitin-like References 7936 Binding Domains 7926 2.3. Antibody-Based Purification of Ubiquitin and Ubiquitin-like Proteins 7926 1. INTRODUCTION 2.4. Challenges and Pitfalls 7926 Proteomes are significantly more complex than genomes 2.5. Summary 7926 and transcriptomes due to protein processing and extensive 3. Ubiquitin Proteomics 7927 post-translational modification (PTM) of proteins. Hundreds ff fi 3.1. Proteomic Studies Employing Tagged of di erent modi cations exist. Release 66 of the RESID database1 (http://www.ebi.ac.uk/RESID/) contains 559 dif- Ubiquitin 7927 ferent modifications, including small chemical modifications 3.2. Ubiquitin Binding Domains 7927 such as phosphorylation, acetylation, and methylation and mod- 3.3. Anti-Ubiquitin Antibodies 7927 ification by small proteins, including ubiquitin and ubiquitin- 3.4. -
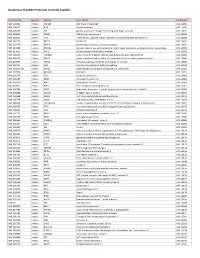
Quantigene Flowrna Probe Sets Currently Available
QuantiGene FlowRNA Probe Sets Currently Available Accession No. Species Symbol Gene Name Catalog No. NM_003452 Human ZNF189 zinc finger protein 189 VA1-10009 NM_000057 Human BLM Bloom syndrome VA1-10010 NM_005269 Human GLI glioma-associated oncogene homolog (zinc finger protein) VA1-10011 NM_002614 Human PDZK1 PDZ domain containing 1 VA1-10015 NM_003225 Human TFF1 Trefoil factor 1 (breast cancer, estrogen-inducible sequence expressed in) VA1-10016 NM_002276 Human KRT19 keratin 19 VA1-10022 NM_002659 Human PLAUR plasminogen activator, urokinase receptor VA1-10025 NM_017669 Human ERCC6L excision repair cross-complementing rodent repair deficiency, complementation group 6-like VA1-10029 NM_017699 Human SIDT1 SID1 transmembrane family, member 1 VA1-10032 NM_000077 Human CDKN2A cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) VA1-10040 NM_003150 Human STAT3 signal transducer and activator of transcripton 3 (acute-phase response factor) VA1-10046 NM_004707 Human ATG12 ATG12 autophagy related 12 homolog (S. cerevisiae) VA1-10047 NM_000737 Human CGB chorionic gonadotropin, beta polypeptide VA1-10048 NM_001017420 Human ESCO2 establishment of cohesion 1 homolog 2 (S. cerevisiae) VA1-10050 NM_197978 Human HEMGN hemogen VA1-10051 NM_001738 Human CA1 Carbonic anhydrase I VA1-10052 NM_000184 Human HBG2 Hemoglobin, gamma G VA1-10053 NM_005330 Human HBE1 Hemoglobin, epsilon 1 VA1-10054 NR_003367 Human PVT1 Pvt1 oncogene homolog (mouse) VA1-10061 NM_000454 Human SOD1 Superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) -

Heat Shock Factors: Integrators of Cell Stress, Development and Lifespan
REVIEWS Heat shock factors: integrators of cell stress, development and lifespan Malin Åkerfelt*‡, Richard I. Morimoto§ and Lea Sistonen*‡ Abstract | Heat shock factors (HSFs) are essential for all organisms to survive exposures to acute stress. They are best known as inducible transcriptional regulators of genes encoding molecular chaperones and other stress proteins. Four members of the HSF family are also important for normal development and lifespan-enhancing pathways, and the repertoire of HSF targets has thus expanded well beyond the heat shock genes. These unexpected observations have uncovered complex layers of post-translational regulation of HSFs that integrate the metabolic state of the cell with stress biology, and in doing so control fundamental aspects of the health of the proteome and ageing. Polytene chromosome In the early 1960s, Ritossa made the seminal discovery of shock response. Here, we present the recent discover- A chromosome that undergoes temperature-induced puffs in polytene chromosomes ies of novel target genes and physiological functions multiple rounds of DNA of Drosophila melanogaster larvae salivary glands1. of HSFs, which have changed the view that HSFs act replication, without cell A decade later, it was shown that the puffing pattern solely in the heat shock response. Based on the current division, and produces many sister chromatids that remain corresponded to a robust activation of genes encoding knowledge of small-molecule activators and inhibitors of synapsed together; for the heat shock proteins (HSPs), which function as HSFs, we also highlight the potential for pharmacologic example, in larval salivary molecular chaperones2. The heat shock response is a modulation of HSF-mediated gene regulation. -

UBE2I Sirna Set I UBE2I Sirna Set I
Catalog # Aliquot Size U224-911-05 3 x 5 nmol U224-911-20 3 x 20 nmol U224-911-50 3 x 50 nmol UBE2I siRNA Set I siRNA duplexes targeted against three exon regions Catalog # U224-911 Lot # Z2109-25 Specificity Formulation UBE2I siRNAs are designed to specifically knock-down The siRNAs are supplied as a lyophilized powder and human UBE2I expression. shipped at room temperature. Product Description Reconstitution Protocol UBE2I siRNA is a pool of three individual synthetic siRNA Briefly centrifuge the tubes (maximum RCF 4,000g) to duplexes designed to knock-down human UBE2I mRNA collect lyophilized siRNA at the bottom of the tube. expression. Each siRNA is 19-25 bases in length. The gene Resuspend the siRNA in 50 µl of DEPC-treated water accession number is NM_003345. (supplied by researcher), which results in a 1x stock solution (10 µM). Gently pipet the solution 3-5 times to mix Gene Aliases and avoid the introduction of bubbles. Optional: aliquot C358B7.1; P18; UBC9 1x stock solutions for storage. Storage and Stability Related Products The lyophilized powder is stable for at least 4 weeks at room temperature. It is recommended that the Product Name Catalog Number lyophilized and resuspended siRNAs are stored at or UBE2E3 (UBCH9) Protein U219-30H below -20oC. After resuspension, siRNA stock solutions ≥2 UBE2F Protein U220-30H µM can undergo up to 50 freeze-thaw cycles without UBE2G1 (UBC7) Protein U221-30H significant degradation. For long-term storage, it is UBE2H Protein U223-30H recommended that the siRNA is stored at -70oC. For most UBE2I Protein U224-30H favorable performance, avoid repeated handling and UBE2J1 Protein U225-30G multiple freeze/thaw cycles. -

Host Cell Factors Necessary for Influenza a Infection: Meta-Analysis of Genome Wide Studies
Host Cell Factors Necessary for Influenza A Infection: Meta-Analysis of Genome Wide Studies Juliana S. Capitanio and Richard W. Wozniak Department of Cell Biology, Faculty of Medicine and Dentistry, University of Alberta Abstract: The Influenza A virus belongs to the Orthomyxoviridae family. Influenza virus infection occurs yearly in all countries of the world. It usually kills between 250,000 and 500,000 people and causes severe illness in millions more. Over the last century alone we have seen 3 global influenza pandemics. The great human and financial cost of this disease has made it the second most studied virus today, behind HIV. Recently, several genome-wide RNA interference studies have focused on identifying host molecules that participate in Influen- za infection. We used nine of these studies for this meta-analysis. Even though the overlap among genes identified in multiple screens was small, network analysis indicates that similar protein complexes and biological functions of the host were present. As a result, several host gene complexes important for the Influenza virus life cycle were identified. The biological function and the relevance of each identified protein complex in the Influenza virus life cycle is further detailed in this paper. Background and PA bound to the viral genome via nucleoprotein (NP). The viral core is enveloped by a lipid membrane derived from Influenza virus the host cell. The viral protein M1 underlies the membrane and anchors NEP/NS2. Hemagglutinin (HA), neuraminidase Viruses are the simplest life form on earth. They parasite host (NA), and M2 proteins are inserted into the envelope, facing organisms and subvert the host cellular machinery for differ- the viral exterior. -

The MALDI TOF E2/E3 Ligase Assay As an Universal Tool for Drug Discovery in the Ubiquitin Pathway
bioRxiv preprint doi: https://doi.org/10.1101/224600; this version posted November 29, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The MALDI TOF E2/E3 ligase assay as an universal tool for drug discovery in the ubiquitin pathway Virginia De Cesare*1, Clare Johnson2, Victoria Barlow2, James Hastie2 Axel Knebel1 and 5 Matthias Trost*1,3 1MRC Protein Phosphorylation and Ubiquitylation Unit, University of Dundee, Dow St, Dundee, DD1 5EH, Scotland, UK; 2MRC Protein Phosphorylation and Ubiquitylation Unit Reagents and Services, University of Dundee, Dow St, Dundee, DD1 5EH, Scotland, UK; . 3Institute for Cell and Molecular Biosciences, Newcastle University, Framlington Place, Newcastle-upon-Tyne, 10 NE2 1HH, UK *To whom correspondence should be addressed: Virginia De Cesare ([email protected]) Matthias Trost ([email protected]) 15 Contact information: V.D.C.: MRC PPU, University of Dundee, Dow St, Dundee, DD1 5EH, Phone: +44 1382 20 85822 M.T.: Newcastle University, Institute for Cell and Molecular Biosciences, Framlington Place, Newcastle-upon-Tyne, NE2 4HH, Phone: +44 191 2087009 Key words: Ubiquitin, E3 ligase, E2 enzyme, MALDI TOF, mass spectrometry, drug 25 discovery, high-throughput, assay, MDM2, HOIP, ITCH 1 bioRxiv preprint doi: https://doi.org/10.1101/224600; this version posted November 29, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

S41467-019-12388-Y.Pdf
ARTICLE https://doi.org/10.1038/s41467-019-12388-y OPEN A tri-ionic anchor mechanism drives Ube2N-specific recruitment and K63-chain ubiquitination in TRIM ligases Leo Kiss1,4, Jingwei Zeng 1,4, Claire F. Dickson1,2, Donna L. Mallery1, Ji-Chun Yang 1, Stephen H. McLaughlin 1, Andreas Boland1,3, David Neuhaus1,5 & Leo C. James1,5* 1234567890():,; The cytosolic antibody receptor TRIM21 possesses unique ubiquitination activity that drives broad-spectrum anti-pathogen targeting and underpins the protein depletion technology Trim-Away. This activity is dependent on formation of self-anchored, K63-linked ubiquitin chains by the heterodimeric E2 enzyme Ube2N/Ube2V2. Here we reveal how TRIM21 facilitates ubiquitin transfer and differentiates this E2 from other closely related enzymes. A tri-ionic motif provides optimally distributed anchor points that allow TRIM21 to wrap an Ube2N~Ub complex around its RING domain, locking the closed conformation and promoting ubiquitin discharge. Mutation of these anchor points inhibits ubiquitination with Ube2N/ Ube2V2, viral neutralization and immune signalling. We show that the same mechanism is employed by the anti-HIV restriction factor TRIM5 and identify spatially conserved ionic anchor points in other Ube2N-recruiting RING E3s. The tri-ionic motif is exclusively required for Ube2N but not Ube2D1 activity and provides a generic E2-specific catalysis mechanism for RING E3s. 1 Medical Research Council Laboratory of Molecular Biology, Cambridge, UK. 2Present address: University of New South Wales, Sydney, NSW, Australia. 3Present address: Department of Molecular Biology, Science III, University of Geneva, Geneva, Switzerland. 4These authors contributed equally: Leo Kiss, Jingwei Zeng. 5These authors jointly supervised: David Neuhaus, Leo C. -

HRD1 and UBE2J1 Target Misfolded MHC Class I Heavy Chains for Endoplasmic Reticulum- Associated Degradation
HRD1 and UBE2J1 target misfolded MHC class I heavy chains for endoplasmic reticulum- associated degradation Marian L. Burr, Florencia Cano, Stanislava Svobodova, Louise H. Boyle, Jessica M. Boname, and Paul J. Lehner1 Cambridge Institute for Medical Research, University of Cambridge, Cambridge CB2 0XY, United Kingdom Edited* by Peter Cresswell, Yale University School of Medicine, New Haven, CT, and approved December 21, 2010 (received for review October 29, 2010) The assembly of MHC class I molecules is governed by stringent orthologs of yeast Hrd1p, HRD1 (Synoviolin) and gp78 (AMFR), endoplasmic reticulum (ER) quality control mechanisms. MHC class I have distinct substrate specificities, exemplifying the increased heavy chains that fail to achieve their native conformation in complexity of mammalian ERAD (6, 7). HRD1 is implicated in the complex with β2-microglobulin (β2m) and peptide are targeted for pathogenesis of rheumatoid arthritis (10), and reported substrates ER-associated degradation. This requires ubiquitination of the MHC include α1-antitrypsin variants (11) and amyloid precursor protein class I heavy chain and its dislocation from the ER to the cytosol for (12). Three E2 ubiquitin-conjugating enzymes function in mam- proteasome-mediated degradation, although the cellular machin- malian ERAD: UBE2J1 and UBE2J2 (yeast Ubc6p orthologs) and ery involved in this process is unknown. Using an siRNA functional UBE2G2 (yeast Ubc7p ortholog) (7, 8, 13). screen in β2m-depleted cells, we identify an essential role for the E3 Many viruses down-regulate cell-surface MHC I to evade im- ligase HRD1 (Synoviolin) together with the E2 ubiquitin-conjugating mune detection (2). The human cytomegalovirus US2 and US11 gene products bind newly synthesized MHC I HCs and initiate enzyme UBE2J1 in the ubiquitination and dislocation of misfolded their rapid dislocation to the cytosol for proteasomal degradation MHC class I heavy chains. -

Ubiquitinating Enzyme Systems Eric Yao, Msc
TECHNICAL BULLETIN Ubiquitinating Enzyme Systems Eric Yao, MSc. Scientist, Product Management - SignalChem Biotech Inc. Target Ub Recognizing Domain Target A E1 E2 Ub Target P Ub P Ub P E3 Ub Ub Poly-Ub A P P E1 E2 E2 - Interacting Ub P Domain Ub Ub Ub Monomeric 26 S Proteosome Ubiquitin Polymeric Ubiquitin Ub Ub Ub Ub Ub Ub Ub Ub Ub Ub Deconjugation/ Ub Ub Ub Recycling Target Degradation Protein modifications by ubiquitin (Ub) or ubiquitin-like proteins (UBLs) participate in many critical cellular processes such as cell-cycle regulation, DNA repair, oncogenesis, antiviral pathways and most notably, proteasomal degradation of target proteins. Ubiquitination and modification by UBLs share a similar catalytic cascade which requires the sequential action of three classes of enzymes: E1 activating enzymes, E2 conjugating enzymes and E3 ligases. Recent research has linked dysregulation of the Ub/UBLs modification system to numerous diseases including cancer, immunological disorders and neurodegeneration. Thus, the high substrate specificity provided by combinations of over 30 E2’s and over 600 E3’s makes these enzymes emerging drug targets. In response to a growing market demand, SignalChem has developed an extensive array of products encompassing enzymes, Ub/UBL modifiers and substrates in the ubiquitination, SUMOylation, ISGylation and NEDDylation processes. Using Promega’s AMP-GloTM technology and an optimized assay protocol, we have identified and validated a variety of functional combinations of the enzyme components. With the established protocol, each enzyme in the catalytic cascade has been assessed for their activity towards generation of free AMP. In addition, inhibition profiles of the ubiquitinating enzymes have been obtained using the assay system, further demonstrating their potential to be used in high-throughput screening to identify lead compounds for drug discovery and development programs. -

Comparative Analysis of the Ubiquitin-Proteasome System in Homo Sapiens and Saccharomyces Cerevisiae
Comparative Analysis of the Ubiquitin-proteasome system in Homo sapiens and Saccharomyces cerevisiae Inaugural-Dissertation zur Erlangung des Doktorgrades der Mathematisch-Naturwissenschaftlichen Fakultät der Universität zu Köln vorgelegt von Hartmut Scheel aus Rheinbach Köln, 2005 Berichterstatter: Prof. Dr. R. Jürgen Dohmen Prof. Dr. Thomas Langer Dr. Kay Hofmann Tag der mündlichen Prüfung: 18.07.2005 Zusammenfassung I Zusammenfassung Das Ubiquitin-Proteasom System (UPS) stellt den wichtigsten Abbauweg für intrazelluläre Proteine in eukaryotischen Zellen dar. Das abzubauende Protein wird zunächst über eine Enzym-Kaskade mit einer kovalent gebundenen Ubiquitinkette markiert. Anschließend wird das konjugierte Substrat vom Proteasom erkannt und proteolytisch gespalten. Ubiquitin besitzt eine Reihe von Homologen, die ebenfalls posttranslational an Proteine gekoppelt werden können, wie z.B. SUMO und NEDD8. Die hierbei verwendeten Aktivierungs- und Konjugations-Kaskaden sind vollständig analog zu der des Ubiquitin- Systems. Es ist charakteristisch für das UPS, daß sich die Vielzahl der daran beteiligten Proteine aus nur wenigen Proteinfamilien rekrutiert, die durch gemeinsame, funktionale Homologiedomänen gekennzeichnet sind. Einige dieser funktionalen Domänen sind auch in den Modifikations-Systemen der Ubiquitin-Homologen zu finden, jedoch verfügen diese Systeme zusätzlich über spezifische Domänentypen. Homologiedomänen lassen sich als mathematische Modelle in Form von Domänen- deskriptoren (Profile) beschreiben. Diese Deskriptoren können wiederum dazu verwendet werden, mit Hilfe geeigneter Verfahren eine gegebene Proteinsequenz auf das Vorliegen von entsprechenden Homologiedomänen zu untersuchen. Da die im UPS involvierten Homologie- domänen fast ausschließlich auf dieses System und seine Analoga beschränkt sind, können domänen-spezifische Profile zur Katalogisierung der UPS-relevanten Proteine einer Spezies verwendet werden. Auf dieser Basis können dann die entsprechenden UPS-Repertoires verschiedener Spezies miteinander verglichen werden.