(12) Patent Application Publication (10) Pub. No.: US 2004/0009477 A1 Fernandez Et Al
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Table 2. Significant
Table 2. Significant (Q < 0.05 and |d | > 0.5) transcripts from the meta-analysis Gene Chr Mb Gene Name Affy ProbeSet cDNA_IDs d HAP/LAP d HAP/LAP d d IS Average d Ztest P values Q-value Symbol ID (study #5) 1 2 STS B2m 2 122 beta-2 microglobulin 1452428_a_at AI848245 1.75334941 4 3.2 4 3.2316485 1.07398E-09 5.69E-08 Man2b1 8 84.4 mannosidase 2, alpha B1 1416340_a_at H4049B01 3.75722111 3.87309653 2.1 1.6 2.84852656 5.32443E-07 1.58E-05 1110032A03Rik 9 50.9 RIKEN cDNA 1110032A03 gene 1417211_a_at H4035E05 4 1.66015788 4 1.7 2.82772795 2.94266E-05 0.000527 NA 9 48.5 --- 1456111_at 3.43701477 1.85785922 4 2 2.8237185 9.97969E-08 3.48E-06 Scn4b 9 45.3 Sodium channel, type IV, beta 1434008_at AI844796 3.79536664 1.63774235 3.3 2.3 2.75319499 1.48057E-08 6.21E-07 polypeptide Gadd45gip1 8 84.1 RIKEN cDNA 2310040G17 gene 1417619_at 4 3.38875643 1.4 2 2.69163229 8.84279E-06 0.0001904 BC056474 15 12.1 Mus musculus cDNA clone 1424117_at H3030A06 3.95752801 2.42838452 1.9 2.2 2.62132809 1.3344E-08 5.66E-07 MGC:67360 IMAGE:6823629, complete cds NA 4 153 guanine nucleotide binding protein, 1454696_at -3.46081884 -4 -1.3 -1.6 -2.6026947 8.58458E-05 0.0012617 beta 1 Gnb1 4 153 guanine nucleotide binding protein, 1417432_a_at H3094D02 -3.13334396 -4 -1.6 -1.7 -2.5946297 1.04542E-05 0.0002202 beta 1 Gadd45gip1 8 84.1 RAD23a homolog (S. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Uncovering Ubiquitin and Ubiquitin-Like Signaling Networks Alfred C
REVIEW pubs.acs.org/CR Uncovering Ubiquitin and Ubiquitin-like Signaling Networks Alfred C. O. Vertegaal* Department of Molecular Cell Biology, Leiden University Medical Center, Albinusdreef 2, 2333 ZA Leiden, The Netherlands CONTENTS 8. Crosstalk between Post-Translational Modifications 7934 1. Introduction 7923 8.1. Crosstalk between Phosphorylation and 1.1. Ubiquitin and Ubiquitin-like Proteins 7924 Ubiquitylation 7934 1.2. Quantitative Proteomics 7924 8.2. Phosphorylation-Dependent SUMOylation 7935 8.3. Competition between Different Lysine 1.3. Setting the Scenery: Mass Spectrometry Modifications 7935 Based Investigation of Phosphorylation 8.4. Crosstalk between SUMOylation and the and Acetylation 7925 UbiquitinÀProteasome System 7935 2. Ubiquitin and Ubiquitin-like Protein Purification 9. Conclusions and Future Perspectives 7935 Approaches 7925 Author Information 7935 2.1. Epitope-Tagged Ubiquitin and Ubiquitin-like Biography 7935 Proteins 7925 Acknowledgment 7936 2.2. Traps Based on Ubiquitin- and Ubiquitin-like References 7936 Binding Domains 7926 2.3. Antibody-Based Purification of Ubiquitin and Ubiquitin-like Proteins 7926 1. INTRODUCTION 2.4. Challenges and Pitfalls 7926 Proteomes are significantly more complex than genomes 2.5. Summary 7926 and transcriptomes due to protein processing and extensive 3. Ubiquitin Proteomics 7927 post-translational modification (PTM) of proteins. Hundreds ff fi 3.1. Proteomic Studies Employing Tagged of di erent modi cations exist. Release 66 of the RESID database1 (http://www.ebi.ac.uk/RESID/) contains 559 dif- Ubiquitin 7927 ferent modifications, including small chemical modifications 3.2. Ubiquitin Binding Domains 7927 such as phosphorylation, acetylation, and methylation and mod- 3.3. Anti-Ubiquitin Antibodies 7927 ification by small proteins, including ubiquitin and ubiquitin- 3.4. -

Synthetic Lethal Screen Demonstrates That a JAK2 Inhibitor Suppresses a BCL6 Dependent IL10RA/JAK2/STAT3 Pathway in High Grade B-Cell Lymphoma
BCL6 suppresses an IL10RA/JAK2/STAT3 pathway Synthetic lethal screen demonstrates that a JAK2 inhibitor suppresses a BCL6 dependent IL10RA/JAK2/STAT3 pathway in high grade B-cell lymphoma. Daniel Beck1,6, Jenny Zobel3,6, Ruth Barber1,2,6, Sian Evans1, Larissa Lezina1, Rebecca L. Allchin1, Matthew Blades4, Richard Elliott5, Christopher J. Lord5, Alan Ashworth5, Andrew C.G. Porter3, Simon D. Wagner1 1Department of Cancer Studies, Ernest and Helen Scott Haematology Research Institute and, 2 Leicester Diagnostic and Drug Development (LD3) Centre, University of Leicester, Lancaster Road, Leicester LE1 7HB, UK, 3Department of Haematology, Imperial College London, Hammersmith Campus, Du Cane Road, London W12 0NN, UK. 4Bioinformatics and Biostatistics Analysis Support Hub (B/BASH), University of Leicester, Lancaster Road, Leicester LE1 9HN and 5The Breakthrough Breast Cancer Research Centre, The Institute of Cancer Research, 237 Fulham Road, London, SW3 6JB, UK. 6The first three authors contributed equally to this work Running title: BCL6 suppresses an IL10RA/JAK2/STAT3 pathway. To whom correspondence should be addressed: Simon D. Wagner, Department of Cancer Studies, Room 104, Hodgkin Building, University of Leicester, Lancaster Road, Leicester LE1 7HB, UK. Tel: 0441162525584, Fax: 0441162525616, Email: [email protected] Keywords: cancer therapy, Janus kinase (JAK), lymphocyte, lymphoma, transcription factor, B-cell lymphoma 6 (BCL-6), synthetic lethal screen. ABSTRACT which shows higher levels of IL10RA, JAK2 and We demonstrate the usefulness of synthetic lethal STAT3 but lower levels of BCL6 than GC- screening of a conditionally BCL6 deficient DLBCL and might be usefully combined with Burkitt lymphoma cell line, DG75-AB7, with a novel approaches such as inhibition of IL10RA. -
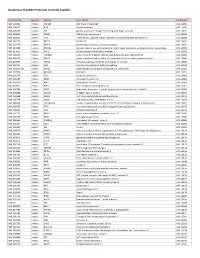
Quantigene Flowrna Probe Sets Currently Available
QuantiGene FlowRNA Probe Sets Currently Available Accession No. Species Symbol Gene Name Catalog No. NM_003452 Human ZNF189 zinc finger protein 189 VA1-10009 NM_000057 Human BLM Bloom syndrome VA1-10010 NM_005269 Human GLI glioma-associated oncogene homolog (zinc finger protein) VA1-10011 NM_002614 Human PDZK1 PDZ domain containing 1 VA1-10015 NM_003225 Human TFF1 Trefoil factor 1 (breast cancer, estrogen-inducible sequence expressed in) VA1-10016 NM_002276 Human KRT19 keratin 19 VA1-10022 NM_002659 Human PLAUR plasminogen activator, urokinase receptor VA1-10025 NM_017669 Human ERCC6L excision repair cross-complementing rodent repair deficiency, complementation group 6-like VA1-10029 NM_017699 Human SIDT1 SID1 transmembrane family, member 1 VA1-10032 NM_000077 Human CDKN2A cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) VA1-10040 NM_003150 Human STAT3 signal transducer and activator of transcripton 3 (acute-phase response factor) VA1-10046 NM_004707 Human ATG12 ATG12 autophagy related 12 homolog (S. cerevisiae) VA1-10047 NM_000737 Human CGB chorionic gonadotropin, beta polypeptide VA1-10048 NM_001017420 Human ESCO2 establishment of cohesion 1 homolog 2 (S. cerevisiae) VA1-10050 NM_197978 Human HEMGN hemogen VA1-10051 NM_001738 Human CA1 Carbonic anhydrase I VA1-10052 NM_000184 Human HBG2 Hemoglobin, gamma G VA1-10053 NM_005330 Human HBE1 Hemoglobin, epsilon 1 VA1-10054 NR_003367 Human PVT1 Pvt1 oncogene homolog (mouse) VA1-10061 NM_000454 Human SOD1 Superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Heat Shock Factors: Integrators of Cell Stress, Development and Lifespan
REVIEWS Heat shock factors: integrators of cell stress, development and lifespan Malin Åkerfelt*‡, Richard I. Morimoto§ and Lea Sistonen*‡ Abstract | Heat shock factors (HSFs) are essential for all organisms to survive exposures to acute stress. They are best known as inducible transcriptional regulators of genes encoding molecular chaperones and other stress proteins. Four members of the HSF family are also important for normal development and lifespan-enhancing pathways, and the repertoire of HSF targets has thus expanded well beyond the heat shock genes. These unexpected observations have uncovered complex layers of post-translational regulation of HSFs that integrate the metabolic state of the cell with stress biology, and in doing so control fundamental aspects of the health of the proteome and ageing. Polytene chromosome In the early 1960s, Ritossa made the seminal discovery of shock response. Here, we present the recent discover- A chromosome that undergoes temperature-induced puffs in polytene chromosomes ies of novel target genes and physiological functions multiple rounds of DNA of Drosophila melanogaster larvae salivary glands1. of HSFs, which have changed the view that HSFs act replication, without cell A decade later, it was shown that the puffing pattern solely in the heat shock response. Based on the current division, and produces many sister chromatids that remain corresponded to a robust activation of genes encoding knowledge of small-molecule activators and inhibitors of synapsed together; for the heat shock proteins (HSPs), which function as HSFs, we also highlight the potential for pharmacologic example, in larval salivary molecular chaperones2. The heat shock response is a modulation of HSF-mediated gene regulation. -

Supplementary Material
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Neurol Neurosurg Psychiatry Page 1 / 45 SUPPLEMENTARY MATERIAL Appendix A1: Neuropsychological protocol. Appendix A2: Description of the four cases at the transitional stage. Table A1: Clinical status and center proportion in each batch. Table A2: Complete output from EdgeR. Table A3: List of the putative target genes. Table A4: Complete output from DIANA-miRPath v.3. Table A5: Comparison of studies investigating miRNAs from brain samples. Figure A1: Stratified nested cross-validation. Figure A2: Expression heatmap of miRNA signature. Figure A3: Bootstrapped ROC AUC scores. Figure A4: ROC AUC scores with 100 different fold splits. Figure A5: Presymptomatic subjects probability scores. Figure A6: Heatmap of the level of enrichment in KEGG pathways. Kmetzsch V, et al. J Neurol Neurosurg Psychiatry 2021; 92:485–493. doi: 10.1136/jnnp-2020-324647 BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Neurol Neurosurg Psychiatry Appendix A1. Neuropsychological protocol The PREV-DEMALS cognitive evaluation included standardized neuropsychological tests to investigate all cognitive domains, and in particular frontal lobe functions. The scores were provided previously (Bertrand et al., 2018). Briefly, global cognitive efficiency was evaluated by means of Mini-Mental State Examination (MMSE) and Mattis Dementia Rating Scale (MDRS). Frontal executive functions were assessed with Frontal Assessment Battery (FAB), forward and backward digit spans, Trail Making Test part A and B (TMT-A and TMT-B), Wisconsin Card Sorting Test (WCST), and Symbol-Digit Modalities test. -

The Influence of Cell Cycle Regulation on Chemotherapy
International Journal of Molecular Sciences Review The Influence of Cell Cycle Regulation on Chemotherapy Ying Sun 1, Yang Liu 1, Xiaoli Ma 2 and Hao Hu 1,* 1 Institute of Biomedical Materials and Engineering, College of Materials Science and Engineering, Qingdao University, Qingdao 266071, China; [email protected] (Y.S.); [email protected] (Y.L.) 2 Qingdao Institute of Measurement Technology, Qingdao 266000, China; [email protected] * Correspondence: [email protected] Abstract: Cell cycle regulation is orchestrated by a complex network of interactions between proteins, enzymes, cytokines, and cell cycle signaling pathways, and is vital for cell proliferation, growth, and repair. The occurrence, development, and metastasis of tumors are closely related to the cell cycle. Cell cycle regulation can be synergistic with chemotherapy in two aspects: inhibition or promotion. The sensitivity of tumor cells to chemotherapeutic drugs can be improved with the cooperation of cell cycle regulation strategies. This review presented the mechanism of the commonly used chemotherapeutic drugs and the effect of the cell cycle on tumorigenesis and development, and the interaction between chemotherapy and cell cycle regulation in cancer treatment was briefly introduced. The current collaborative strategies of chemotherapy and cell cycle regulation are discussed in detail. Finally, we outline the challenges and perspectives about the improvement of combination strategies for cancer therapy. Keywords: chemotherapy; cell cycle regulation; drug delivery systems; combination chemotherapy; cancer therapy Citation: Sun, Y.; Liu, Y.; Ma, X.; Hu, H. The Influence of Cell Cycle Regulation on Chemotherapy. Int. J. 1. Introduction Mol. Sci. 2021, 22, 6923. https:// Chemotherapy is currently one of the main methods of tumor treatment [1]. -

Investigation Into the Effect of LRRK2-Rab10 Protein Interactions on the Proboscis Extension Response of the Fruit Fly Drosophila Melanogaster
Investigation into the effect of LRRK2-Rab10 protein interactions on the Proboscis Extension Response of the fruit fly Drosophila melanogaster Laura Covill Masters by Research University of York Biology December 2018 Abstract Parkinson’s Disease (PD) is a debilitating disease which affects 1% of the population worldwide and is characterised by stiffness, tremor and bradykinesia. PD is a complex disease with many suspected genetic and environmental causes, and it is critical to understand all the pathways involved in disease progression to develop effective therapies for PD, which currently has no cure. A kinase- coding gene, LRRK2 has emerged as a focal point for much PD research, particularly PD-associated SNP LRRK2-G2019S, which leads to LRRK2 overactivity. Rab proteins, a series of small GTPases, have been identified among the proteins phosphorylated by LRRK2. These interactions may be modelled in the fruit fly Drosophila melanogaster. Using optogenetics in the fly, this project investigates the relationship between the LRRK2-G2019S and Rab10 interaction, and the speed and degree of tremor of Proboscis Extension Response (PER) by triggering a PER in fly lines of different genotypes. Significant bradykinesia in Rab10 null flies which was not recreated in flies with dopaminergic neuron Rab10RNAi suggests that the bradykinesia PER phenotype is caused by off-target effect of Rab10-KO in another tissue of the fly than the dopaminergic neurons. Over-expression of Rab10 in dopaminergic neurons of flies also expressing LRRK2-G2019S produced -

CHML Promotes Liver Cancer Metastasis by Facilitating Rab14 Recycle
ARTICLE https://doi.org/10.1038/s41467-019-10364-0 OPEN CHML promotes liver cancer metastasis by facilitating Rab14 recycle Tian-Wei Chen1,2,3, Fen-Fen Yin1,2, Yan-Mei Yuan1,2, Dong-Xian Guan1, Erbin Zhang1,2, Feng-Kun Zhang1,2, Hao Jiang1,2, Ning Ma1,2, Jing-Jing Wang1, Qian-Zhi Ni1,2, Lin Qiu1, Jing Feng3, Xue-Li Zhang3, Ying Bao4, Kang Wang5, Shu-Qun Cheng5, Xiao-Fan Wang6, Xiang Wang7, Jing-Jing Li1,2 & Dong Xie1,2,8,9 Metastasis-associated recurrence is the major cause of poor prognosis in hepatocellular 1234567890():,; carcinoma (HCC), however, the underlying mechanisms remain largely elusive. In this study, we report that expression of choroideremia-like (CHML) is increased in HCC, associated with poor survival, early recurrence and more satellite nodules in HCC patients. CHML promotes migration, invasion and metastasis of HCC cells, in a Rab14-dependent manner. Mechanism study reveals that CHML facilitates constant recycling of Rab14 by escorting Rab14 to the membrane. Furthermore, we identify several metastasis regulators as cargoes carried by Rab14-positive vesicles, including Mucin13 and CD44, which may contribute to metastasis- promoting effects of CHML. Altogether, our data establish CHML as a potential promoter of HCC metastasis, and the CHML-Rab14 axis may be a promising therapeutic target for HCC. 1 CAS Key Laboratory of Nutrition, Metabolism and Food Safety, Shanghai Institute of Nutrition and Health, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Xuhui district 200031, China. 2 University of Chinese Academy of Sciences, Chinese Academy of Sciences, Xuhui district 200031, China. 3 Department of General Surgery, Fengxian Hospital Affiliated to Southern Medical University, 6600 Nanfeng Road, Shanghai 201499, China. -

DNA Topoisomerase I and DNA Gyrase As Targets for TB Therapy, Drug Discov Today (2016), J.Drudis.2016.11.006
Drug Discovery Today Volume 00, Number 00 November 2016 REVIEWS GENE TO SCREEN DNA topoisomerase I and DNA gyrase as targets for TB therapy Reviews 1,2 1 3 Valakunja Nagaraja , Adwait A. Godbole , Sara R. Henderson and 3 Anthony Maxwell 1 Department of Microbiology and Cell Biology, Indian Institute of Science, Bangalore 560 012, India 2 Jawaharlal Nehru Centre for Advanced Scientific Research, Bangalore 560064, India 3 Department of Biological Chemistry, John Innes Centre, Norwich Research Park, Norwich NR4 7UH, UK Tuberculosis (TB) is the deadliest bacterial disease in the world. New therapeutic agents are urgently needed to replace existing drugs for which resistance is a significant problem. DNA topoisomerases are well-validated targets for antimicrobial and anticancer chemotherapies. Although bacterial topoisomerase I has yet to be exploited as a target for clinical antibiotics, DNA gyrase has been extensively targeted, including the highly clinically successful fluoroquinolones, which have been utilized in TB therapy. Here, we review the exploitation of topoisomerases as antibacterial targets and summarize progress in developing new agents to target DNA topoisomerase I and DNA gyrase from Mycobacterium tuberculosis. Introduction provides for a certain degree of overlap in their functions. For Although the information content of DNA is essentially indepen- instance, in Escherichia coli, there are four topoisomerases: two dent of how the DNA is knotted or twisted, the access to this type I [topoisomerase (topo) I and topo III] and two type II (DNA information depends on the topology of the DNA. DNA topoi- gyrase and topo IV). In vitro, all four enzymes are capable of DNA somerases are ubiquitous enzymes that maintain the topological relaxation, whereas, in vivo, their roles tend to be more special- homeostasis within the cell during these DNA transaction pro- ized, for example, gyrase introduces negative supercoils, whereas cesses [1–3].