Lecture 20 Eukaryotic Genes and Genomes Ii
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Glucan Phosphorylase-Catalyzed Enzymatic Reactions Using Analog Substrates to Synthesize Non-Natural Oligo- and Polysaccharides
catalysts Review α-Glucan Phosphorylase-Catalyzed Enzymatic Reactions Using Analog Substrates to Synthesize Non-Natural Oligo- and Polysaccharides Jun-ichi Kadokawa Department of Chemistry, Biotechnology, and Chemical Engineering, Graduate School of Science and Engineering, Kagoshima University, 1-21-40 Korimoto, Kagoshima 860-0065, Japan; [email protected]; Tel.: +81-99-285-7743 Received: 9 October 2018; Accepted: 16 October 2018; Published: 19 October 2018 Abstract: As natural oligo- and polysaccharides are important biomass resources and exhibit vital biological functions, non-natural oligo- and polysaccharides with a well-defined structure can be expected to act as new functional materials with specific natures and properties. α-Glucan phosphorylase (GP) is one of the enzymes that have been used as catalysts for practical synthesis of oligo- and polysaccharides. By means of weak specificity for the recognition of substrates by GP, non-natural oligo- and polysaccharides has precisely been synthesized. GP-catalyzed enzymatic glycosylations using several analog substrates as glycosyl donors have been carried out to produce oligosaccharides having different monosaccharide residues at the non-reducing end. Glycogen, a highly branched natural polysaccharide, has been used as the polymeric glycosyl acceptor and primer for the GP-catalyzed glycosylation and polymerization to obtain glycogen-based non-natural polysaccharide materials. Under the conditions of removal of inorganic phosphate, thermostable GP-catalyzed enzymatic polymerization of analog monomers occurred to give amylose analog polysaccharides. Keywords: analog substrate; α-glucan phosphorylase; non-natural oligo- and polysaccharides 1. Introduction Oligo- and polysaccharides are widely distributed in nature and enact specific important biological functions in accordance with their chemical structures [1]. -

Defective Galactose Oxidation in a Patient with Glycogen Storage Disease and Fanconi Syndrome
Pediatr. Res. 17: 157-161 (1983) Defective Galactose Oxidation in a Patient with Glycogen Storage Disease and Fanconi Syndrome M. BRIVET,"" N. MOATTI, A. CORRIAT, A. LEMONNIER, AND M. ODIEVRE Laboratoire Central de Biochimie du Centre Hospitalier de Bichre, 94270 Kremlin-Bicetre, France [M. B., A. C.]; Faculte des Sciences Pharmaceutiques et Biologiques de I'Universite Paris-Sud, 92290 Chatenay-Malabry, France [N. M., A. L.]; and Faculte de Midecine de I'Universiti Paris-Sud et Unite de Recherches d'Hepatologie Infantile, INSERM U 56, 94270 Kremlin-Bicetre. France [M. 0.1 Summary The patient's diet was supplemented with 25-OH-cholecalci- ferol, phosphorus, calcium, and bicarbonate. With this treatment, Carbohydrate metabolism was studied in a child with atypical the serum phosphate concentration increased, but remained be- glycogen storage disease and Fanconi syndrome. Massive gluco- tween 0.8 and 1.0 mmole/liter, whereas the plasma carbon dioxide suria, partial resistance to glucagon and abnormal responses to level returned to normal (18-22 mmole/liter). Rickets was only carbohydrate loads, mainly in the form of major impairment of partially controlled. galactose utilization were found, as reported in previous cases. Increased blood lactate to pyruvate ratios, observed in a few cases of idiopathic Fanconi syndrome, were not present. [l-14ClGalac- METHODS tose oxidation was normal in erythrocytes, but reduced in fresh All studies of the patient and of the subjects who served as minced liver tissue, despite normal activities of hepatic galactoki- controls were undertaken after obtaining parental or personal nase, uridyltransferase, and UDP-glucose 4epirnerase in hornog- consent. enates of frozen liver. -

Chem331 Glycogen Metabolism
Glycogen metabolism Glycogen review - 1,4 and 1,6 α-glycosidic links ~ every 10 sugars are branched - open helix with many non-reducing ends. Effective storage of glucose Glucose storage Liver glycogen 4.0% 72 g Muscle glycogen 0.7% 245 g Blood Glucose 0.1% 10 g Large amount of water associated with glycogen - 0.5% of total weight Glycogen stored in granules in cytosol w/proteins for synthesis, degradation and control There are very different means of control of glycogen metabolism between liver and muscle Glycogen biosynthetic and degradative cycle Two different pathways - which do not share enzymes like glycolysis and gluconeogenesis glucose -> glycogen glycogenesis - biosynthetic glycogen -> glucose 1-P glycogenolysis - breakdown Evidence for two paths - Patients lacking phosphorylase can still synthesize glycogen - hormonal regulation of both directions Glycogenolysis (glycogen breakdown)- Glycogen Phosphorylase glycogen (n) + Pi -> glucose 1-p + glycogen (n-1) • Enzyme binds and cleaves glycogen into monomers at the end of the polymer (reducing ends of glycogen) • Dimmer interacting at the N-terminus. • rate limiting - controlled step in glycogen breakdown • glycogen phosphorylase - cleavage of 1,4 α glycosidic bond by Pi NOT H2O • Energy of phosphorolysis vs. hydrolysis -low standard state free energy change -transfer potential -driven by Pi concentration -Hydrolysis would require additional step s/ cost of ATP - Think of the difference between adding a phosphate group with hydrolysis • phosphorylation locks glucose in cell (imp. for muscle) • Phosphorylase binds glycogen at storage site and the catalytic site is 4 to 5 glucose residues away from the catalytic site. • Phosphorylase removes 1 residue at a time from glycogen until 4 glucose residues away on either side of 1,6 branch point – stericaly hindered by glycogen storage site • Cleaves without releasing at storage site • general acid/base catalysts • Inorganic phosphate attacks the terminal glucose residue passing through an oxonium ion intermediate. -

The Origins of Protein Phosphorylation
historical perspective The origins of protein phosphorylation Philip Cohen The reversible phosphorylation of proteins is central to the regulation of most aspects of cell func- tion but, even after the first protein kinase was identified, the general significance of this discovery was slow to be appreciated. Here I review the discovery of protein phosphorylation and give a per- sonal view of the key findings that have helped to shape the field as we know it today. he days when protein phosphorylation was an abstruse backwater, best talked Tabout between consenting adults in private, are over. My colleagues no longer cringe on hearing that “phosphorylase kinase phosphorylates phosphorylase” and their eyes no longer glaze over when a “”kinase kinase kinase” is mentioned. This is because protein phosphorylation has gradu- ally become an integral part of all the sys- tems they are studying themselves. Indeed it would be difficult to find anyone today who would disagree with the statement that “the reversible phosphorylation of proteins regu- lates nearly every aspect of cell life”. Phosphorylation and dephosphorylation, catalysed by protein kinases and protein phosphatases, can modify the function of a protein in almost every conceivable way; for Carl and Gerty Cori, the 1947 Nobel Laureates. Picture: Science Photo Library. example by increasing or decreasing its bio- logical activity, by stabilizing it or marking it for destruction, by facilitating or inhibiting movement between subcellular compart- so long before its general significance liver enzyme that catalysed the phosphory- ments, or by initiating or disrupting pro- was appreciated? lation of casein3. Soon after, Fischer and tein–protein interactions. -

Glycogenosis Due to Liver and Muscle Phosphorylase Kinase Deficiency
Pediat. Res. 15: 299-303 (198 1) genetics muscle glycogenosis phosphorylase kinase deficiency liver Glycogenosis Due to Liver and Muscle Phosphorylase Kinase Deficiency N. BASHAN. T. C. IANCU. A. LERNER. D. FRASER, R. POTASHNIK. AND S. W. MOSES'"' Pediatric Research Laborarorv. Soroka Medical Center. Iaculr~of Health Sciences. Ben-Gurion Universi!,' of Negev. Beer-Sheva, and Department of Pediatrics. Carmel Hospiral. Huifa. Israel Summary hepatomegaly. The family history disclosed that two sisters were similarly affected, whereas one older brother was apparently A four-year-old Israeli Arab boy was found to have glycogen healthy. accumulation in both liver and muscle without clinical symptoms. Past history was unremarkable. The patient's height was below Liver phosphorylase kinase (PK) activity was 20% of normal, the third percentile for his age in contrast to a normal weight. He resulting in undetectable activity of phosphorylase a. Muscle PK had a doll face and a protuberant abdomen. The liver was palpable activity was about 25% of normal, resulting in a marked decrease 9 cm below the costal margin. Slight muscular hypotonia and of phosphorylase a activity. weakness were noticeable with normal tendon reflexes. He had Two sisters showed a similar pattern, whereas one brother had slightly abnormal liver function tests. a fasting blood sugar of 72 normal PK activity. The patient's liver protein kinase activity was mg %, a normal glucagon test. and no lactic acidemia or uricemia normal. Addition of exogenous protein kinase did not affect PK but slight lipidemia. Electronmicroscopic studies of a liver biopsy activity, whereas exogenous PK restored phosphorylase activity revealed marked deposition of glycogen. -
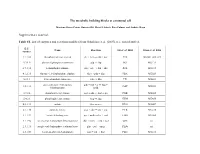
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

Histidine Kinases and the Missing Phosphoproteome from Prokaryotes to Eukaryotes Kevin Adam and Tony Hunter
Laboratory Investigation (2018) 98, 233–247 © 2018 USCAP, Inc All rights reserved 0023-6837/18 $32.00 PATHOBIOLOGY IN FOCUS Histidine kinases and the missing phosphoproteome from prokaryotes to eukaryotes Kevin Adam and Tony Hunter Protein phosphorylation is the most common type of post-translational modification in eukaryotes. The phosphoproteome is defined as the complete set of experimentally detectable phosphorylation sites present in a cell’s proteome under various conditions. However, we are still far from identifying all the phosphorylation sites in a cell mainly due to the lack of information about phosphorylation events involving residues other than Ser, Thr and Tyr. Four types of phosphate–protein linkage exist and these generate nine different phosphoresidues—pSer, pThr, pTyr, pHis, pLys, pArg, pAsp, pGlu and pCys. Most of the effort in studying protein phosphorylation has been focused on Ser, Thr and Tyr phosphorylation. The recent development of 1- and 3-pHis monoclonal antibodies promises to increase our understanding of His phosphorylation and the kinases and phosphatases involved. Several His kinases are well defined in prokaryotes, especially those involved in two-component system (TCS) signaling. However, in higher eukaryotes, NM23, a protein originally characterized as a nucleoside diphosphate kinase, is the only characterized protein–histidine kinase. This ubiquitous and conserved His kinase autophosphorylates its active site His, and transfers this phosphate either onto a nucleoside diphosphate or onto a protein His residue. Studies of NM23 protein targets using newly developed anti-pHis antibodies will surely help illuminate the elusive His phosphorylation-based signaling pathways. This review discusses the role that the NM23/NME/NDPK phosphotransferase has, how the addition of the pHis phosphoproteome will expand the phosphoproteome and make His phosphorylation part of the global phosphorylation world. -

Regulation of Fructose-6-Phosphate 2-Kinase By
Proc. Natt Acad. Sci. USA Vol. 79, pp. 325-329, January 1982 Biochemistry Regulation of fructose-6-phosphate 2-kinase by phosphorylation and dephosphorylation: Possible mechanism for coordinated control of glycolysis and glycogenolysis (phosphofructokinase) EISUKE FURUYA*, MOTOKO YOKOYAMA, AND KOSAKU UYEDAt Pre-Clinical Science Unit of the Veterans Administration Medical Center, 4500 South Lancaster Road, Dallas, Texas 75216; and Biochemistry Department of the University ofTexas Health Science Center, 5323 Harry Hines Boulevard, Dallas, Texas 75235 Communicated by Jesse C. Rabinowitz, September 28, 1981 ABSTRACT The kinetic properties and the control mecha- Fructose 6-phosphate + ATP nism of fructose-6-phosphate 2-kinase (ATP: D-fructose-6-phos- -3 Fructose + ADP. [1] phate 2-phosphotransferase) were investigated. The molecular 2,6-bisphosphate weight of the enzyme is -100,000 as determined by gel filtration. The plot of initial velocity versus ATP concentration is hyperbolic We have shown that the administration of extremely low con- with a K. of 1.2 mM. However, the plot of enzyme activity as a centrations of glucagon (0.1 fM) or high concentrations of epi- function of fructose 6-phosphate is sigmoidal. The apparent K0.5 nephrine (10 ,uM) to hepatocytes results in inactivation offruc- for fructose 6-phosphate is 20 ,IM. Fructose-6-phosphate 2-kinase tose-6-phosphate 2-kinase and concomitant decrease in the is inactivated by -the catalytic subunit of cyclic AMP-dependent fructose 2,6-bisphosphate level (12). These results, as well as protein kinase, and the inactivation is closely correlated with phos- more recent data using Ca2+ and the Ca2+ ionophore A23187 phorylation. -

Overview of the Synthesis of Nucleoside Phosphates and Polyphosphates 13.1.6
Overview of the Synthesis of Nucleoside UNIT 13.1 Phosphates and Polyphosphates Phosphorylated nucleosides play a domi- ity to the synthesis. Side reactions can occur, nant role in biochemistry. Primary metabolism, such as depurination of the nucleoside, phos- DNA replication and repair, RNA synthesis, phorylation of the nucleobase, as well as chemi- protein synthesis, signal transduction, polysac- cal alteration of nucleobase analogs. Due to charide biosynthesis, and enzyme regulation their intrinsic reactivity, the synthesis of phos- are just a handful of processes involving these phoanhydride bonds is also synthetically chal- molecules. Literally thousands of enzymes use lenging. Phosphate anhydrides are phosphory- these compounds as substrates and/or regula- lating reagents that are readily degraded under tors. The need to obtain such compounds in acidic conditions. Finally, purification of syn- both labeled and unlabeled forms, as well as a thetic nucleotides can be problematic. Ionic burgeoning need for analogs, has driven the reagents, starting materials, and mixtures of development of a myriad of chemical and en- regioisomers (2′-, 3′-, 5′-phosphates) can be zymatic synthetic approaches. As chemical en- particularly difficult to separate from the de- tities, few molecules possess the wide array of sired product. densely packed functionality present in phos- In spite of the many potential difficulties phorylated nucleosides. This poses a formida- associated with nucleoside phosphorylation ble challenge to the synthetic chemist, one that and polyphosphorylation, a certain amount of has not yet been fully overcome. This overview success has been achieved in these areas. Given will address some common methods (synthetic the wealth of phosphorylating reagents avail- and enzymatic) used to construct phosphory- able, simple phosphorylation of nucleosides at lated nucleosides. -

The Glycogen Phosphorylase
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 1999 The glycogen phosphorylase/ phosphorylase kinase interaction: effects of mutations in the amino-terminal region of glycogen phosphorylase Alyssa Christine Biorn Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Biochemistry Commons, and the Molecular Biology Commons Recommended Citation Biorn, Alyssa Christine, "The glycogen phosphorylase/ phosphorylase kinase interaction: effects of mutations in the amino-terminal region of glycogen phosphorylase " (1999). Retrospective Theses and Dissertations. 12198. https://lib.dr.iastate.edu/rtd/12198 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. INFORMATION TO USERS This manuscript has been reproduced from the microfilm master. UMI films the text directly from the original or copy submitted. Thus, some thesis and dissertation copies are in typewriter face, while others may be from any type of computer printer. The quality of this reproduction is dependent upon the quality of the copy submitted. Broken or indistinct print, colored or poor quality illustrations and photographs, print bleedthrough, substandard margins, and improper alignment can adversely affect reproduction. In the unlikely event that the author did not send UMI a complete manuscript and there are missing pages, these will be noted. Also, if unauthorized copyright material had to be removed, a note will indicate the deletion. -

Increased Potency and Efficacy of Combined Phosphorylase
Increased Potency and Efficacy of Combined Phosphorylase Inactivation and Glucokinase Activation in Control of Hepatocyte Glycogen Metabolism Laura J. Hampson and Loranne Agius Glucokinase and phosphorylase both have a high control Activators of glucokinase lower blood glucose in animal strength over hepatocyte glycogen metabolism and are models of type 2 diabetes by potentiating glucose-induced potential therapeutic targets for type 2 diabetes. We insulin secretion (6) and stimulating hepatic glucose me- tested whether combined phosphorylase inactivation tabolism (7). Their efficacy in lowering blood glucose and glucokinase activation is a more effective strategy concentrations is consistent with the effect of activating for controlling hepatic glycogen metabolism than single- mutations of the glucokinase gene that cause hyperinsu- site targeting. Activation of glucokinase by enzyme linemia and hypoglycemia in humans (8). The high control overexpression combined with selective dephosphoryla- tion of phosphorylase-a by an indole carboxamide that strength of glucokinase on liver metabolism is explained favors the T conformation of phosphorylase caused a by the reciprocal control of glucokinase activity by its greater stimulation of glycogen synthesis than the sum regulatory protein (9) and the role of glucose-6-phosphate of either treatment alone. This result is explained by (G6P), the product of the glucokinase-catalyzed reaction, the complementary roles of elevated glucose-6-phos- as an activator of glycogen synthase (10,11) and inactiva- phate (G6P; a positive modulator) and depleted phos- tor of phosphorylase (12,13). phorylase-a (a negative modulator) in activating Inhibitors of liver phosphorylase also lower blood glycogen synthase and also by synergistic inactivation glucose in animal models of type 2 diabetes (14–16). -

Energy Metabolism of Thehuman Fallopian Tube
Energy metabolism of the human Fallopian tube I. A. Brewis, R. M. L. Winston and H. J. Leese department of Biology, University of York, Heslington, York Y01 5DD, UK; and 2 Institute of Obstetrics and Gynaecology, Royal Postgraduate Medical School, Hammersmith Hospital, DuCane Road, London W12 ONN, UK Summary. The consumption of oxygen (QO2), the production of lactate and the profile of four key metabolic enzymes were measured in small samples of human oviductal mucosa (endosalpinx) removed at surgery. The QO2 in the absence of substrate was 3\m=.\4\g=m\1O2 (mg dry wt) \m=-\1h\m=-\1,a value typical of quiescent tissue. The QO2 was stimulated by glucose, but diminished by glutamine and acetoacetate. Tissue lactate production was low and not increased by glucose. Hexokinase had the highest activity of the enzymes measured, followed by 2-oxoglutarate dehydrogenase; 6-phosphofructokinase and glycogen phosphorylase had low activities. The data are consistent with the proposition that glucose is a major metabolic fuel for human endosalpinx. Keywords: Fallopian tube; oxygen uptake; metabolism; glucose; lactate; human Introduction The human Fallopian tube consists of an outer muscle layer, the myosalpinx, which surrounds an inner mucosal lining, or endosalpinx. This paper reports data on the energy metabolism of the endosalpinx, the two specialized energy-requiring functions of which are the secretion of molecules into oviductal fluid (Leese, 1988) and the mechanical action of cilia. Endosalpinx from a variety of species including humans is being widely used in the 'co- culture' of preimplantation embryos (Bongso et ai, 1990). Information on the metabolism of the endosalpinx in vitro is likely to be important in defining its role in sustaining embryo development.