Effects of Overexpression of HBP1 Upon Growth and Differentiation of Leukemic Myeloid Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The P63 Target HBP-1 Is Required for Keratinocyte Differentiation and Stratification
The p63 target HBP-1 is required for keratinocyte differentiation and stratification. Roberto Mantovani, Serena Borrelli, Eleonora Candi, Diletta Dolfini, Olì Maria Victoria Grober, Alessandro Weisz, Gennaro Melino, Alessandra Viganò, Bing Hu, Gian Paolo Dotto To cite this version: Roberto Mantovani, Serena Borrelli, Eleonora Candi, Diletta Dolfini, Olì Maria Victoria Grober, et al.. The p63 target HBP-1 is required for keratinocyte differentiation and stratification.. Cell Death and Differentiation, Nature Publishing Group, 2010, 10.1038/cdd.2010.59. hal-00542927 HAL Id: hal-00542927 https://hal.archives-ouvertes.fr/hal-00542927 Submitted on 4 Dec 2010 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Running title: HBP1 in skin differentiation. The p63 target HBP-1 is required for skin differentiation and stratification. Serena Borrelli1, Eleonora Candi2, Bing Hu3, Diletta Dolfini1, Maria Ravo4, Olì Maria Victoria Grober4, Alessandro Weisz4,5, GianPaolo Dotto3, Gerry Melino2,6, Maria Alessandra Viganò1 and Roberto Mantovani1*. 1) Dipartimento di Scienze Biomolecolari e Biotecnologie. Università degli Studi di Milano. Via Celoria 26, 20133 Milano, Italy. 2) Biochemistry IDI-IRCCS laboratory, c/o University of Rome "Tor Vergata", Via Montpellier 1, 00133 Roma, Italy. -

Identification of SNP-Containing Regulatory Motifs in the Myelodysplastic Syndromes Model Using SNP Arrays Ad Gene Expression Arrays" (2013)
University of Central Florida STARS Faculty Bibliography 2010s Faculty Bibliography 1-1-2013 Identification of SNP-containing egulatr ory motifs in the myelodysplastic syndromes model using SNP arrays ad gene expression arrays Jing Fan Jennifer G. Dy Chung-Che Chang University of Central Florida Xiaoboo Zhou Find similar works at: https://stars.library.ucf.edu/facultybib2010 University of Central Florida Libraries http://library.ucf.edu This Article is brought to you for free and open access by the Faculty Bibliography at STARS. It has been accepted for inclusion in Faculty Bibliography 2010s by an authorized administrator of STARS. For more information, please contact [email protected]. Recommended Citation Fan, Jing; Dy, Jennifer G.; Chang, Chung-Che; and Zhou, Xiaoboo, "Identification of SNP-containing regulatory motifs in the myelodysplastic syndromes model using SNP arrays ad gene expression arrays" (2013). Faculty Bibliography 2010s. 3960. https://stars.library.ucf.edu/facultybib2010/3960 Chinese Journal of Cancer Original Article Jing Fan 1, Jennifer G. Dy 1, Chung鄄Che Chang 2 and Xiaobo Zhou 3 Abstract Myelodysplastic syndromes have increased in frequency and incidence in the American population, but patient prognosis has not significantly improved over the last decade. Such improvements could be realized if biomarkers for accurate diagnosis and prognostic stratification were successfully identified. In this study, we propose a method that associates two state鄄of 鄄th e鄄ar t array technologies single nucleotide polymor鄄 要 phism (SNP) array and gene expression array with gene motifs considered transcription factor -binding 要 sites (TFBS). We are particularly interested in SNP鄄co ntaining motifs introduced by genetic variation and mutation as TFBS. -

The Ire1a-XBP1 Pathway Promotes T Helper Cell Differentiation by Resolving Secretory Stress and Accelerating Proliferation
bioRxiv preprint doi: https://doi.org/10.1101/235010; this version posted December 15, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The IRE1a-XBP1 pathway promotes T helper cell differentiation by resolving secretory stress and accelerating proliferation Jhuma Pramanik1, Xi Chen1, Gozde Kar1,2, Tomás Gomes1, Johan Henriksson1, Zhichao Miao1,2, Kedar Natarajan1, Andrew N. J. McKenzie3, Bidesh Mahata1,2*, Sarah A. Teichmann1,2,4* 1. Wellcome Trust Sanger Institute, Wellcome Genome Campus, Hinxton, Cambridge, CB10 1SA, United Kingdom 2. EMBL-European Bioinformatics Institute, Wellcome Genome Campus, Hinxton, Cambridge, CB10 1SD, United Kingdom 3. MRC Laboratory of Molecular Biology, Cambridge Biomedical Campus, Francis Crick Avenue, Cambridge CB2 OQH, United Kingdom 4. Theory of Condensed Matter, Cavendish Laboratory, 19 JJ Thomson Ave, Cambridge CB3 0HE, United Kingdom. *To whom correspondence should be addressed: [email protected] and [email protected] Keywords: Th2 lymphocyte, XBP1, Genome wide XBP1 occupancy, Th2 lymphocyte proliferation, ChIP-seq, RNA-seq, Th2 transcriptome Summary The IRE1a-XBP1 pathway, a conserved adaptive mediator of the unfolded protein response, is indispensable for the development of secretory cells. It maintains endoplasmic reticulum homeostasis by facilitating protein folding and enhancing secretory capacity of the cells. Its role in immune cells is emerging. It is involved in dendritic cell, plasma cell and eosinophil development and differentiation. Using genome-wide approaches, integrating ChIPmentation and mRNA-sequencing data, we have elucidated the regulatory circuitry governed by the IRE1a-XBP1 pathway in type-2 T helper cells (Th2). -

Homeobox Transcription Factor Prox1 in Sympathetic Ganglia of Vertebrate Embryos: Correlation with Human Stage 4S Neuroblastoma
0031-3998/10/6802-0112 Vol. 68, No. 2, 2010 PEDIATRIC RESEARCH Printed in U.S.A. Copyright © 2010 International Pediatric Research Foundation, Inc. Homeobox Transcription Factor Prox1 in Sympathetic Ganglia of Vertebrate Embryos: Correlation With Human Stage 4s Neuroblastoma JU¨ RGEN BECKER, BAIGANG WANG, HELENA PAVLAKOVIC, KERSTIN BUTTLER, AND JO¨ RG WILTING Department of Anatomy and Cell Biology, University Medicine Goettingen, 37075 Goettingen, Germany ABSTRACT: Previously, we observed expression of the homeobox progression of tumors derived from these tissues has not been transcription factor Prox1 in neuroectodermal embryonic tissues. investigated. We have studied Prox1 expression in the sym- Besides essential functions during embryonic development, Prox1 pathetic nervous system of avian and murine embryos, and in has been implicated in both progression and suppression of malig- childhood tumors derived from this tissue: neuroblastoma nancies. Here, we show that Prox1 is expressed in embryonic sym- (NB). We show that Prox1 is expressed in sympathetic neu- pathetic trunk ganglia of avian and murine embryos. Prox1 protein is rons during early stages of development at similar levels as in localized in the nucleus of neurofilament-positive sympathetic neu- rons. Sympathetic progenitors represent the cell of origin of neuro- lymphatic endothelial cells (LECs), but at greatly reduced blastoma (NB), the most frequent solid extracranial malignancy of levels in human NB cell lines. Studies of primary NB of all children. NB may progress to life-threatening stage 4, or regress stages (stages 1–4s) show significantly higher amounts of spontaneously in the special stage 4s. By qRT-PCR, we show that Prox1 mRNA in stage 4s. -

Mice Lacking Hbp1 Function Are Viable and Fertile
RESEARCH ARTICLE Mice Lacking Hbp1 Function Are Viable and Fertile Cassy M. Spiller1¤a*, Dagmar Wilhelm1¤b, David A. Jans2, Josephine Bowles3, Peter Koopman1 1 Institute for Molecular Bioscience, The University of Queensland, Brisbane, Queensland, Australia, 2 School of Biomedical Sciences, Monash University, Clayton, Victoria, Australia, 3 School of Biomedical Sciences, The University of Queensland, Brisbane, Queensland, Australia ¤a Current address: School of Biomedical Sciences, The University of Queensland, Brisbane, Queensland, Australia ¤b Current address: Department of Anatomy & Neuroscience, The University of Melbourne, Melbourne, Victoria, Australia a1111111111 * [email protected] a1111111111 a1111111111 a1111111111 a1111111111 Abstract Fetal germ cell development is tightly regulated by the somatic cell environment, and is char- acterised by cell cycle states that differ between XY and XX gonads. In the testis, gonocytes enter G1/G0 arrest from 12.5 days post coitum (dpc) in mice and maintain cell cycle arrest OPEN ACCESS until after birth. Failure to correctly maintain G1/G0 arrest can result in loss of germ cells or, Citation: Spiller CM, Wilhelm D, Jans DA, Bowles conversely, germ cell tumours. High mobility group box containing transcription factor 1 J, Koopman P (2017) Mice Lacking Hbp1 Function (HBP1) is a transcription factor that was previously identified in fetal male germ cells at the Are Viable and Fertile. PLoS ONE 12(1): e0170576. doi:10.1371/journal.pone.0170576 time of embryonic cell cycle arrest. In somatic cells, HBP1 is classified as a tumour suppres- sor protein, known to regulate proliferation and senescence. We therefore investigated the Editor: Stefan Schlatt, University Hospital of MuÈnster, GERMANY possible role of HBP1 in the initiation and maintenance of fetal germ cell G1/G0 arrest using the mouse model. -

Diversification Pattern of the HMG and SOX Family Members During Evolution
J Mol Evol (1999) 48:517–527 © Springer-Verlag New York Inc. 1999 Diversification Pattern of the HMG and SOX Family Members During Evolution Ste´phan Soullier,1 Philippe Jay,1 Francis Poulat,1 Jean-Marc Vanacker,2 Philippe Berta,1 Vincent Laudet2 1 ERS155 du CNRS, Centre de Recherche en Biochimie Macromole´culaire, CNRS, BP5051, Route de Mende, 34293 Montpellier Cedex 5, France 2 UMR 319 du CNRS, Oncologie Mole´culaire, Institut de Biologie de Lille, Institut Pasteur de Lille, 1 rue Calmette, 59019 Lille Cedex, France Received: 20 July 1998 / Accepted: 19 October 1998 Abstract. From a database containing the published brate HMG box 2, insect SSRP, and plant HMG. The HMG protein sequences, we constructed an alignment of various UBF boxes cannot be clustered together and their the HMG box functional domain based on sequence diversification appears to be extremely ancient, probably identity. Due to the large number of sequences (more before the appearance of metazoans. than 250) and the short size of this domain, several data sets were used. This analysis reveals that the HMG box Key words: HMG box — SOX proteins — Sry — superfamily can be separated into two clearly defined Molecular phylogeny subfamilies: (i) the SOX/MATA/TCF family, which clusters proteins able to bind to specific DNA sequences; and (ii) the HMG/UBF family, which clusters members Introduction which bind non specifically to DNA. The appearance and diversification of these subfamilies largely predate the split between the yeast and the metazoan lineages. Par- The high-mobility group (HMG) proteins were first de- ticular emphasis was placed on the analysis of the SOX fined by their electrophoretic behavior on SDS-PAGE, as subfamily. -
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
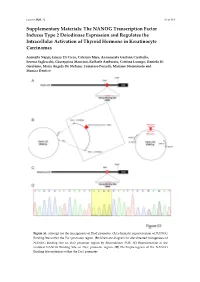
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

The Tumor Suppressor HHEX Inhibits Axon Growth When Prematurely Expressed in Developing Central Nervous System Neurons
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by epublications@Marquette Marquette University e-Publications@Marquette Biological Sciences Faculty Research and Biological Sciences, Department of Publications 9-1-2015 The umorT Suppressor HHEX Inhibits Axon Growth when Prematurely Expressed in Developing Central Nervous System Neurons Matthew .T Simpson Marquette University Ishwariya Venkatesh Marquette University Ben L. Callif Marquette University Laura K. Thiel Marquette University Denise M. Coley Marquette University See next page for additional authors Accepted version. Molecular and Cellular Neuroscience, Vol 68 )September 2015): 272-283. DOI. © 2015 Elsevier Inc. Used with permission. NOTICE: this is the author’s version of a work that was accepted for publication in Molecular and Cellular Neuroscience. Changes resulting from the publishing process, such as peer review, editing, corrections, structural formatting, and other quality control mechanisms may not be reflected in this document. Changes may have been made to this work since it was submitted for publication. A definitive version was subsequently published in Molecular and Cellular Neuroscience, Vol 68 )September 2015): 272-283. DOI. Authors Matthew T. Simpson, Ishwariya Venkatesh, Ben L. Callif, Laura K. Thiel, Denise M. Coley, Kristen N. Winsor, Zimei Wang, Audra A. Kramer, Jessica K. Lerch, and Murray G. Blackmore This article is available at e-Publications@Marquette: https://epublications.marquette.edu/bio_fac/515 NOT THE PUBLISHED VERSION; this is the author’s final, peer-reviewed manuscript. The published version may be accessed by following the link in the citation at the bottom of the page. The Tumor Suppressor HHEX Inhibits Axon Growth When Prematurely Expressed in Developing Central Nervous System Neurons Matthew T. -

The P63 Target HBP1 Is Required for Skin Differentiation and Stratification
Cell Death and Differentiation (2010) 17, 1896–1907 & 2010 Macmillan Publishers Limited All rights reserved 1350-9047/10 $32.00 www.nature.com/cdd The p63 target HBP1 is required for skin differentiation and stratification S Borrelli1, E Candi2,BHu3, D Dolfini1, M Ravo4, OMV Grober4, A Weisz4,5, GP Dotto3, G Melino2,6, MA Vigano` 1 and R Mantovani*,1 Genetic experiments established that p63 is crucial for the development and maintenance of pluristratified epithelia. In the RNA interference (RNAi) screening for targets of p63 in keratinocytes, we identified the transcription factor, High Mobility Group (HMG) box protein 1 (HBP1). HBP1 is an HMG-containing repressor transiently induced during differentiation of several cell lineages. We investigated the relationship between the two factors: using RNAi, overexpression, chromatin immunoprecipita- tions and transient transfections with reporter constructs, we established that HBP1 is directly repressed by p63. This was further confirmed in vivo by evaluating expression in p63 knockout mice and in transgenics expressing p63 in basal keratinocytes. Consistent with these findings, expression of HBP1 increases upon differentiation of primary keratinocytes and HaCaT cells in culture, and it is higher in the upper layers of human skin. Inactivation of HBP1 by RNAi prevents differentiation of keratinocytes and stratification of organotypic skin cultures. Finally, we analyzed the keratinocyte transcriptomes after HBP1 RNAi; in addition to repression of growth-promoting genes, unexpected activation of differentiation genes was uncovered, coexisting with repression of other genes involved in epithelial cornification. Our data indicate that suppression of HBP1 is part of the growth-promoting strategy of p63 in the lower layers of epidermis and that HBP1 temporally coordinates expression of genes involved in stratification, leading to the formation of the skin barrier. -

Maintenance of Mammary Epithelial Phenotype by Transcription Factor Runx1 Through Mitotic Gene Bookmarking Joshua Rose University of Vermont
University of Vermont ScholarWorks @ UVM Graduate College Dissertations and Theses Dissertations and Theses 2019 Maintenance Of Mammary Epithelial Phenotype By Transcription Factor Runx1 Through Mitotic Gene Bookmarking Joshua Rose University of Vermont Follow this and additional works at: https://scholarworks.uvm.edu/graddis Part of the Biochemistry Commons, and the Genetics and Genomics Commons Recommended Citation Rose, Joshua, "Maintenance Of Mammary Epithelial Phenotype By Transcription Factor Runx1 Through Mitotic Gene Bookmarking" (2019). Graduate College Dissertations and Theses. 998. https://scholarworks.uvm.edu/graddis/998 This Thesis is brought to you for free and open access by the Dissertations and Theses at ScholarWorks @ UVM. It has been accepted for inclusion in Graduate College Dissertations and Theses by an authorized administrator of ScholarWorks @ UVM. For more information, please contact [email protected]. MAINTENANCE OF MAMMARY EPITHELIAL PHENOTYPE BY TRANSCRIPTION FACTOR RUNX1 THROUGH MITOTIC GENE BOOKMARKING A Thesis Presented by Joshua Rose to The Faculty of the Graduate College of The University of Vermont In Partial Fulfillment of the Requirements for the Degree of Master of Science Specializing in Cellular, Molecular, and Biomedical Sciences January, 2019 Defense Date: November 12, 2018 Thesis Examination Committee: Sayyed Kaleem Zaidi, Ph.D., Advisor Gary Stein, Ph.D., Advisor Seth Frietze, Ph.D., Chairperson Janet Stein, Ph.D. Jonathan Gordon, Ph.D. Cynthia J. Forehand, Ph.D. Dean of the Graduate College ABSTRACT Breast cancer arises from a series of acquired mutations that disrupt normal mammary epithelial homeostasis and create multi-potent cancer stem cells that can differentiate into clinically distinct breast cancer subtypes. Despite improved therapies and advances in early detection, breast cancer remains the leading diagnosed cancer in women. -

Exosomal Micrornas: Pleiotropic Impacts on Breast Cancer Metastasis and Their Clinical Perspectives
biology Review Exosomal microRNAs: Pleiotropic Impacts on Breast Cancer Metastasis and Their Clinical Perspectives Li-Bo Tang 1,2,† , Shu-Xin Ma 3,†, Zhuo-Hui Chen 2, Qi-Yuan Huang 1,2 , Long-Yuan Wu 1,4, Yi Wang 1, Rui-Chen Zhao 1,3 and Li-Xia Xiong 1,5,* 1 Department of Pathophysiology, Basic Medical College, Nanchang University, Nanchang 330006, China; [email protected] (L.-B.T.); [email protected] (Q.-Y.H.); [email protected] (L.-Y.W.); [email protected] (Y.W.); [email protected] (R.-C.Z.) 2 Second Clinical Medical College, Nanchang University, Nanchang 330006, China; [email protected] 3 Queen Mary School, Jiangxi Medical College, Nanchang University, Nanchang 330006, China; [email protected] 4 First Clinical Medical College, Nanchang University, Nanchang 330006, China 5 Jiangxi Province Key Laboratory of Tumor Pathogenesis and Molecular Pathology, Nanchang 330006, China * Correspondence: [email protected]; Tel.: +86-791-8636-0556 † These authors contributed equally to this work. Simple Summary: This review has comprehensively summarized the most recent studies in the last few years about exosomal microRNAs on metastasis of breast cancer (BC), systematically outlined and elucidated the pleiotropic roles that exosomal microRNAs take, and discussed the specific underlying mechanisms related. Besides, we also clearly demonstrate the clinical implications of exosomal microRNAs in various aspects, including early-stage discovery of BC, systematic and Citation: Tang, L.-B.; Ma, S.-X.; Chen, targeted therapy, and the selection of anti-cancer chemo-agents. This review clarifies the scope and Z.-H.; Huang, Q.-Y.; Wu, L.-Y.; Wang, extent of current studies about the relationship between exosomal microRNAs and metastasis of Y.; Zhao, R.-C.; Xiong, L.-X. -

Mir-155: a Novel Target in Allergic Asthma
Int. J. Mol. Sci. 2016, 17, 1773; doi: 10.3390/ijms17101773 S1 of S2 Supplementary Materials: miR-155: A Novel Target in Allergic Asthma Hong Zhou, Junyao Li, Peng Gao, Qi Wang and Jie Zhang Table S1. miR-155 targets identified by TargetScan, miRTarBase and both bioinformatic analyses. Names Total Elements LCORL CHD8 TOMM20 SLC33A1 CHD9 ZNF248 IRF2BP2 DNAJB1 C10orf12 PALLD CARD11 GNAS ZBTB38 RAPH1 ETNK2 MSH6 ARL5B CCDC41 MMP16 RHEB TOMM34 MEF2A RICTOR RAB11FIP2 FAM135A ZBTB18 TMEM33 TCF12 KRAS TM6SF1 DHX40 PICALM MYO10 TCF4 FUBP1 ATP6V1C1 SERTAD2 SH3PXD2A UBQLN2 YWHAZ AGO4 CHAF1A ZNF236 MORC3 MEIS1 WWC1 TAB2 NAA50 PRKAR1A CSNK1G2 PHC2 HBP1 SPRED1 ADAM10 KANSL1 MIDN ZNF644 NFAT5 IL17RB STRN3 MAP3K10 ZSWIM6 DMTF1 ITK PDE3A ZIC3 PELI1 CSNK1A1 ARID2 GSK3B SPIN1 TSPAN14 PTAR1 FOXK1 WEE1 PKN2 TPD52 CARHSP1 MYBL1 WBP1L SAP30L VEZF1 EEF2 FLT1 PHF17 RCOR1 SMAD2 CBFB RORA HIVEP2 CHD7 RAP1B TargetScan 190 SPI1 PEA15 FGF7 RREB1 CBL MYLK S1PR1 TMEM136 PIK3CA NKX3-1 CTLA4 miRTarBase RAB3B SMAD1 ANKFY1 FOS SKIV2L2 SMARCA4 TP53INP1 TSHZ3 PSMG1 FGF2 SKI CPEB4 JARID2 MSI2 SWSAP1 LRRC40 ETS1 COPS3 IKBKE SOCS1 TRIM32 LRRC59 CDC73 RAB5C CAB39 LNX2 NSA2 CDC37 MBNL3 MAFB INPP5D E2F2 PKIA RAB30 CEP41 DET1 UBTD2 C3orf18 BACH1 RAPGEF2 CREBRF SHANK2 PAXBP1 BAG5 KBTBD2 KIF3A HHIP EHD1 HERC4 PALD1 HNRNPA3 N4BP1 PIK3R1 PTPRJ NOVA1 GPM6B CKAP5 TAPT1 CLDN1 SIRT1 SEPT11 COLGALT1 HMGCS1 TLE4 TERF1 ZNF703 FOXO3 KCTD3 APC INADL BCAT1 WNK1 CEBPB TRPS1 CSF1R KDM3A MYO1D RNF123 TADA2B AAK1 RBAK USP8 RCN2 SMAD5 PDE12 ZNF652 MYB SMNDC1 RPTOR PLCE1 KIF26B TNIK RTKN2 ZPLD1 ARRB2