Structural and Mechanistic Insight Into the Basis of Mucopolysaccharidosis IIIB
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Epidemiology of Mucopolysaccharidoses Update
diagnostics Review Epidemiology of Mucopolysaccharidoses Update Betul Celik 1,2 , Saori C. Tomatsu 2 , Shunji Tomatsu 1 and Shaukat A. Khan 1,* 1 Nemours/Alfred I. duPont Hospital for Children, Wilmington, DE 19803, USA; [email protected] (B.C.); [email protected] (S.T.) 2 Department of Biological Sciences, University of Delaware, Newark, DE 19716, USA; [email protected] * Correspondence: [email protected]; Tel.: +302-298-7335; Fax: +302-651-6888 Abstract: Mucopolysaccharidoses (MPS) are a group of lysosomal storage disorders caused by a lysosomal enzyme deficiency or malfunction, which leads to the accumulation of glycosaminoglycans in tissues and organs. If not treated at an early stage, patients have various health problems, affecting their quality of life and life-span. Two therapeutic options for MPS are widely used in practice: enzyme replacement therapy and hematopoietic stem cell transplantation. However, early diagnosis of MPS is crucial, as treatment may be too late to reverse or ameliorate the disease progress. It has been noted that the prevalence of MPS and each subtype varies based on geographic regions and/or ethnic background. Each type of MPS is caused by a wide range of the mutational spectrum, mainly missense mutations. Some mutations were derived from the common founder effect. In the previous study, Khan et al. 2018 have reported the epidemiology of MPS from 22 countries and 16 regions. In this study, we aimed to update the prevalence of MPS across the world. We have collected and investigated 189 publications related to the prevalence of MPS via PubMed as of December 2020. In total, data from 33 countries and 23 regions were compiled and analyzed. -

Association of Gene Ontology Categories with Decay Rate for Hepg2 Experiments These Tables Show Details for All Gene Ontology Categories
Supplementary Table 1: Association of Gene Ontology Categories with Decay Rate for HepG2 Experiments These tables show details for all Gene Ontology categories. Inferences for manual classification scheme shown at the bottom. Those categories used in Figure 1A are highlighted in bold. Standard Deviations are shown in parentheses. P-values less than 1E-20 are indicated with a "0". Rate r (hour^-1) Half-life < 2hr. Decay % GO Number Category Name Probe Sets Group Non-Group Distribution p-value In-Group Non-Group Representation p-value GO:0006350 transcription 1523 0.221 (0.009) 0.127 (0.002) FASTER 0 13.1 (0.4) 4.5 (0.1) OVER 0 GO:0006351 transcription, DNA-dependent 1498 0.220 (0.009) 0.127 (0.002) FASTER 0 13.0 (0.4) 4.5 (0.1) OVER 0 GO:0006355 regulation of transcription, DNA-dependent 1163 0.230 (0.011) 0.128 (0.002) FASTER 5.00E-21 14.2 (0.5) 4.6 (0.1) OVER 0 GO:0006366 transcription from Pol II promoter 845 0.225 (0.012) 0.130 (0.002) FASTER 1.88E-14 13.0 (0.5) 4.8 (0.1) OVER 0 GO:0006139 nucleobase, nucleoside, nucleotide and nucleic acid metabolism3004 0.173 (0.006) 0.127 (0.002) FASTER 1.28E-12 8.4 (0.2) 4.5 (0.1) OVER 0 GO:0006357 regulation of transcription from Pol II promoter 487 0.231 (0.016) 0.132 (0.002) FASTER 6.05E-10 13.5 (0.6) 4.9 (0.1) OVER 0 GO:0008283 cell proliferation 625 0.189 (0.014) 0.132 (0.002) FASTER 1.95E-05 10.1 (0.6) 5.0 (0.1) OVER 1.50E-20 GO:0006513 monoubiquitination 36 0.305 (0.049) 0.134 (0.002) FASTER 2.69E-04 25.4 (4.4) 5.1 (0.1) OVER 2.04E-06 GO:0007050 cell cycle arrest 57 0.311 (0.054) 0.133 (0.002) -

Bioinformatics Classification of Mutations in Patients with Mucopolysaccharidosis IIIA
Metabolic Brain Disease (2019) 34:1577–1594 https://doi.org/10.1007/s11011-019-00465-6 ORIGINAL ARTICLE Bioinformatics classification of mutations in patients with Mucopolysaccharidosis IIIA Himani Tanwar1 & D. Thirumal Kumar1 & C. George Priya Doss1 & Hatem Zayed2 Received: 30 April 2019 /Accepted: 8 July 2019 /Published online: 5 August 2019 # The Author(s) 2019 Abstract Mucopolysaccharidosis (MPS) IIIA, also known as Sanfilippo syndrome type A, is a severe, progressive disease that affects the central nervous system (CNS). MPS IIIA is inherited in an autosomal recessive manner and is caused by a deficiency in the lysosomal enzyme sulfamidase, which is required for the degradation of heparan sulfate. The sulfamidase is produced by the N- sulphoglucosamine sulphohydrolase (SGSH) gene. In MPS IIIA patients, the excess of lysosomal storage of heparan sulfate often leads to mental retardation, hyperactive behavior, and connective tissue impairments, which occur due to various known missense mutations in the SGSH, leading to protein dysfunction. In this study, we focused on three mutations (R74C, S66W, and R245H) based on in silico pathogenic, conservation, and stability prediction tool studies. The three mutations were further subjected to molecular dynamic simulation (MDS) analysis using GROMACS simulation software to observe the structural changes they induced, and all the mutants exhibited maximum deviation patterns compared with the native protein. Conformational changes were observed in the mutants based on various geometrical parameters, such as conformational stability, fluctuation, and compactness, followed by hydrogen bonding, physicochemical properties, principal component analysis (PCA), and salt bridge analyses, which further validated the underlying cause of the protein instability. Additionally, secondary structure and surrounding amino acid analyses further confirmed the above results indicating the loss of protein function in the mutants compared with the native protein. -

Assessment of Predicted Enzymatic Activity of Alpha-N- Acetylglucosaminidase (NAGLU) Variants of Unknown Significance for CAGI 2016
HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Hum Mutat Manuscript Author . Author manuscript; Manuscript Author available in PMC 2020 April 14. Published in final edited form as: Hum Mutat. 2019 September ; 40(9): 1519–1529. doi:10.1002/humu.23875. Assessment of predicted enzymatic activity of alpha-N- acetylglucosaminidase (NAGLU) variants of unknown significance for CAGI 2016 Wyatt T. Clark1, Laura Kasak2,3, Constantina Bakolitsa2, Zhiqiang Hu2, Gaia Andreoletti2,*, Giulia Babbi4, Yana Bromberg5, Rita Casadio4, Roland Dunbrack6, Lukas Folkman7, Colby T. Ford8, David Jones9, Panagiotis Katsonis10,*, Kunal Kundu11, Olivier Lichtarge12, Pier Luigi Martelli4, Sean D. Mooney13,*, Conor Nodzak8, Lipika R. Pal11, Predrag Radivojac14,*, Castrense Savojardo4, Xinghua Shi8, Yaoqi Zhou15, Aneeta Uppal8, Qifang Xu6, Yizhou Yin11,*, Vikas Pejaver16, Meng Wang17, Liping Wei17, John Moult11, G. Karen Yu1, Steven E. Brenner2, Jonathan H. LeBowitz1 1BioMarin Pharmaceutical, San Rafael, California, USA 2Department of Plant and Microbial Biology, University of California, Berkeley, CA, USA 3Institute of Biomedicine and Translational Medicine, University of Tartu, Tartu, Estonia 4Biocomputing Group, Department of Pharmacy and Biotechnology, University of Bologna, Bologna, Italy 5Department of Biochemistry and Microbiology, Rutgers University, New Brunswick, NJ, USA 6Fox Chase Cancer Center, Philadelphia, PA, USA 7School of Information and Communication Technology, Griffith University, Southport, Australia 8Department of -

Prediction of the Effect of Naturally Occurring Missense Mutations On
bioRxiv preprint doi: https://doi.org/10.1101/598870; this version posted April 5, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. PREDICTION OF THE EFFECT OF NATURALLY OCCURRING MISSENSE MUTATIONS ON CELLULAR N-Acetyl-Glucosaminidase ENZYMATIC ACTIVITY Colby T. Ford∗ Conor M. Nodzak Department of Bioinformatics and Genomics Department of Bioinformatics and Genomics The University of North Carolina at Charlotte The University of North Carolina at Charlotte Charlotte, NC 28223 Charlotte, NC 28223 Aneeta Uppal Xinghua Shi Department of Bioinformatics and Genomics Department of Bioinformatics and Genomics The University of North Carolina at Charlotte The University of North Carolina at Charlotte Charlotte, NC 28223 Charlotte, NC 28223 April 4, 2019 ABSTRACT In 2015, the Critical Assessment of Genome Interpretation (CAGI) proposed a challenge to devise a computational method for predicting the phenotypic consequences of genetic variants of a lysosomal hydrolase enzyme known as α-N-acetylglucosaminidase (NAGLU). In 2014, the Human Gene Mutation Database released that 153 NAGLU mutations associated with MPS IIIB and 90 of them are missense mutations. The ExAC dataset catalogued 189 missense mutations NAGLU based on exome sequence data from about 60,000 individual and 24 of them are known to be disease associated. Biotechnology company, BioMarin, has quantified the relative functionality of NAGLU for the remaining subset of 165 missense mutations. For this particular challenge, we examined the subset missense mutations within the ExAC dataset and predicted the probability of a given mutation being deleterious and relating this measure to the capacity of enzymatic activity. -

Matching Whole Genomes to Rare Genetic Disorders: Identification of Potential
bioRxiv preprint doi: https://doi.org/10.1101/707687; this version posted July 18, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Matching whole genomes to rare genetic disorders: Identification of potential causative variants using phenotype-weighted knowledge in the CAGI SickKids5 clinical genomes challenge Lipika R. Pal 1*, Kunal Kundu 1, 2, Yizhou Yin 1, John Moult 1, 3* 1 Institute for Bioscience and Biotechnology Research, University of Maryland, 9600 Gudelsky Drive, Rockville, MD 20850, USA. 2 Computational Biology, Bioinformatics and Genomics, Biological Sciences Graduate Program, University of Maryland, College Park, MD 20742, USA. 3 Department of Cell Biology and Molecular Genetics, University of Maryland, College Park, MD 20742, USA. *Corresponding authors: [email protected] Phone: (240) 314-6146 [email protected] Phone: (240) 314-6241 FAX: (240) 314-6255 Funding Information: This work was partially supported by NIH R01GM120364 and R01GM104436 to JM. The CAGI experiment coordination is supported by NIH U41 HG007346 and the CAGI conference by NIH R13 HG006650. bioRxiv preprint doi: https://doi.org/10.1101/707687; this version posted July 18, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Keywords: Whole-genome sequencing data, Human Phenotype Ontology (HPO), Eye disorder, Connective-tissue disorder, Neurological diseases, Diagnostic variants, CAGI5. -

Sanfilippo Type B Syndrome (Mucopolysaccharidosis III B)
European Journal of Human Genetics (1999) 7, 34–44 t © 1999 Stockton Press All rights reserved 1018–4813/99 $12.00 http://www.stockton-press.co.uk/ejhg ARTICLES Sanfilippo type B syndrome (mucopolysaccharidosis III B): allelic heterogeneity corresponds to the wide spectrum of clinical phenotypes Birgit Weber1, Xiao-Hui Guo1, Wim J Kleijer2, Jacques JP van de Kamp3, Ben JHM Poorthuis4 and John J Hopwood1 1Lysosomal Diseases Research Unit, Department of Chemical Pathology, Women’s and Children’s Hospital, North Adelaide, Australia 2Department of Clinical Genetics, University Hospital, Rotterdam 3Leiden University Medical Centre, Leiden 4University of Leiden, Department of Pediatrics, Leiden, The Netherlands Sanfilippo B syndrome (mucopolysaccharidosis IIIB, MPS IIIB) is caused by a deficiency of α-N-acetylglucosaminidase, a lysosomal enzyme involved in the degradation of heparan sulphate. Accumulation of the substrate in lysosomes leads to degeneration of the central nervous system with progressive dementia often combined with hyperactivity and aggressive behaviour. Age of onset and rate of progression vary considerably, whilst diagnosis is often delayed due to the absence of the pronounced skeletal changes observed in other mucopolysaccharidoses. Cloning of the gene and cDNA encoding α-N-acetylglucosaminidase enabled a study of the molecular basis of this syndrome. We were able to identify 31 mutations, 25 of them novel, and two polymorphisms in the 40 patients mostly of Australasian and Dutch origin included in this study. The observed allellic heterogeneity reflects the wide spectrum of clinical phenotypes reported for MPS IIIB patients. The majority of changes are missense mutations; also four nonsense and nine frameshift mutations caused by insertions or deletions were identified. -

Recombinant Human Α‑N‑Acetylglucosaminidase/NAGLU Catalog Number: 7096-GH
Recombinant Human α‑N‑acetylglucosaminidase/NAGLU Catalog Number: 7096-GH DESCRIPTION Source Chinese Hamster Ovary cell line, CHO-derived human alpha-N-acetylglucosaminidase/NAGLU protein Met1-Trp743, with a C-terminal 6-His tag Accession # P54802 N-terminal Sequence Asp24 Analysis Predicted Molecular 81 kDa Mass SPECIFICATIONS SDS-PAGE 70-95 kDa, reducing conditions Activity Measured by its ability to hydrolyze 4-Nitrophenyl-N-acetyl-α-D-glucosaminide. The specific activity is >900 pmol/min/μg, as measured under the decribed conditions. Endotoxin Level <1.0 EU per 1 μg of the protein by the LAL method. Purity >95%, by SDS-PAGE visualized with Silver Staining and quantitative densitometry by Coomassie® Blue Staining. Formulation Supplied as a 0.2 μm filtered solution in Tris and NaCl. See Certificate of Analysis for details. Activity Assay Protocol Materials Assay Buffer: 0.1 M Sodium Citrate, 0.25 M NaCl, pH 4.5 Recombinant Human α‑N‑acetylglucosaminidase/NAGLU (rhNAGLU) (Catalog # 7096-GH) Substrate: 4-Nitrophenyl-N-acetyl-α-D-glucosaminide (Sigma, Catalog # N-8759), 5 mM stock in deionized water NaOH, 0.2 M stock in deionized water 96-well Clear Plate (Costar, Catalog # 92592) Plate Reader (Model: SpectraMax Plus by Molecular Devices) or equivalent Assay 1. Dilute rhNAGLU to 3 ng/μL in Assay Buffer. 2. Dilute Substrate to 3 mM in Assay Buffer. 3. In a plate combine 50 μL of rhNAGLU and 50 μL of 3 mM Substrate. Include a Substrate Blank containing 50 μL Assay Buffer and 50 μL of 3 mM Substrate. 4. Seal plate and incubate at 37 °C for 20 minutes. -

Expression of Human Α-N-Acetylglucosaminidase in Sf9 Insect Cells: Effect of Cryptic Splice Site Removal and Native Secretion-Signaling Peptide Addition
Expression of Human α-N-Acetylglucosaminidase in Sf9 Insect Cells: Effect of Cryptic Splice Site Removal and Native Secretion-Signaling Peptide Addition by Roni Rebecca Jantzen B.Sc., University of Victoria, 2009 A Thesis Submitted in Partial Fulfillment of the Requirements for the Degree of MASTER OF SCIENCE in the Department of Biology Roni Rebecca Jantzen, 2011 University of Victoria All rights reserved. This thesis may not be reproduced in whole or in part, by photocopy or other means, without the permission of the author. ii Supervisory Committee Expression of Human α-N-Acetylglucosaminidase in Sf9 Insect Cells: Effect of Cryptic Splice Site Removal and Native Secretion-Signaling Peptide Addition by Roni Rebecca Jantzen B.Sc., University of Victoria, 2009 Supervisory Committee Dr. Francis Y. M. Choy, Supervisor (Department of Biology) Dr. Robert L. Chow, Departmental Member (Department of Biology) Dr. C. Peter Constabel, Departmental Member (Department of Biology) iii Abstract Supervisory Committee Dr. Francis Y. M. Choy, Supervisor (Department of Biology) Dr. Robert L. Chow, Departmental Member (Department of Biology) Dr. C. Peter Constabel, Departmental Member (Department of Biology) Human α-N-Acetylglucosaminidase (Naglu) is a lysosomal acid hydrolase implicated in tthe rare metabolic storage disorder known as mucopolysaccharidosis type IIIB (MPS IIIB; also Sanfilippo syndrome B). Absence of this enzyme results in cytotoxic accumulation of heparan sulphate in the central nervous system, causing mental retardation and a shortened lifespan. Enzyme replacement therapy is not currently effective to treat neurological symptoms due to the inability of exogenous Naglu to access the brain. This laboratory uses a Spodoptera frugiperda (Sf9) insect cell system to express Naglu fused to a synthetic protein transduction domain with the intent to facilitate delivery of Naglu across the blood-brain barrier. -

Mouse Models of Inherited Retinal Degeneration with Photoreceptor Cell Loss
cells Review Mouse Models of Inherited Retinal Degeneration with Photoreceptor Cell Loss 1, 1, 1 1,2,3 1 Gayle B. Collin y, Navdeep Gogna y, Bo Chang , Nattaya Damkham , Jai Pinkney , Lillian F. Hyde 1, Lisa Stone 1 , Jürgen K. Naggert 1 , Patsy M. Nishina 1,* and Mark P. Krebs 1,* 1 The Jackson Laboratory, Bar Harbor, Maine, ME 04609, USA; [email protected] (G.B.C.); [email protected] (N.G.); [email protected] (B.C.); [email protected] (N.D.); [email protected] (J.P.); [email protected] (L.F.H.); [email protected] (L.S.); [email protected] (J.K.N.) 2 Department of Immunology, Faculty of Medicine Siriraj Hospital, Mahidol University, Bangkok 10700, Thailand 3 Siriraj Center of Excellence for Stem Cell Research, Faculty of Medicine Siriraj Hospital, Mahidol University, Bangkok 10700, Thailand * Correspondence: [email protected] (P.M.N.); [email protected] (M.P.K.); Tel.: +1-207-2886-383 (P.M.N.); +1-207-2886-000 (M.P.K.) These authors contributed equally to this work. y Received: 29 February 2020; Accepted: 7 April 2020; Published: 10 April 2020 Abstract: Inherited retinal degeneration (RD) leads to the impairment or loss of vision in millions of individuals worldwide, most frequently due to the loss of photoreceptor (PR) cells. Animal models, particularly the laboratory mouse, have been used to understand the pathogenic mechanisms that underlie PR cell loss and to explore therapies that may prevent, delay, or reverse RD. Here, we reviewed entries in the Mouse Genome Informatics and PubMed databases to compile a comprehensive list of monogenic mouse models in which PR cell loss is demonstrated. -

Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands That Promote Axonal Growth
Research Article: New Research Development Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands that Promote Axonal Growth Jeremy S. Toma,1 Konstantina Karamboulas,1,ª Matthew J. Carr,1,2,ª Adelaida Kolaj,1,3 Scott A. Yuzwa,1 Neemat Mahmud,1,3 Mekayla A. Storer,1 David R. Kaplan,1,2,4 and Freda D. Miller1,2,3,4 https://doi.org/10.1523/ENEURO.0066-20.2020 1Program in Neurosciences and Mental Health, Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada, 2Institute of Medical Sciences University of Toronto, Toronto, Ontario M5G 1A8, Canada, 3Department of Physiology, University of Toronto, Toronto, Ontario M5G 1A8, Canada, and 4Department of Molecular Genetics, University of Toronto, Toronto, Ontario M5G 1A8, Canada Abstract Peripheral nerves provide a supportive growth environment for developing and regenerating axons and are es- sential for maintenance and repair of many non-neural tissues. This capacity has largely been ascribed to paracrine factors secreted by nerve-resident Schwann cells. Here, we used single-cell transcriptional profiling to identify ligands made by different injured rodent nerve cell types and have combined this with cell-surface mass spectrometry to computationally model potential paracrine interactions with peripheral neurons. These analyses show that peripheral nerves make many ligands predicted to act on peripheral and CNS neurons, in- cluding known and previously uncharacterized ligands. While Schwann cells are an important ligand source within injured nerves, more than half of the predicted ligands are made by nerve-resident mesenchymal cells, including the endoneurial cells most closely associated with peripheral axons. At least three of these mesen- chymal ligands, ANGPT1, CCL11, and VEGFC, promote growth when locally applied on sympathetic axons. -
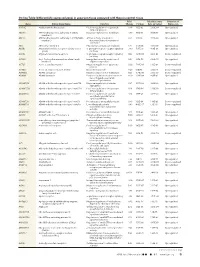
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18