Comparative Proteomic Analysis Applied for Study of Morphogenic Competence During Somatic Embryogenesis in Sugarcane
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Folic Acid Antagonists: Antimicrobial and Immunomodulating Mechanisms and Applications
International Journal of Molecular Sciences Review Folic Acid Antagonists: Antimicrobial and Immunomodulating Mechanisms and Applications Daniel Fernández-Villa 1, Maria Rosa Aguilar 1,2 and Luis Rojo 1,2,* 1 Instituto de Ciencia y Tecnología de Polímeros, Consejo Superior de Investigaciones Científicas, CSIC, 28006 Madrid, Spain; [email protected] (D.F.-V.); [email protected] (M.R.A.) 2 Consorcio Centro de Investigación Biomédica en Red de Bioingeniería, Biomateriales y Nanomedicina, 28029 Madrid, Spain * Correspondence: [email protected]; Tel.: +34-915-622-900 Received: 18 September 2019; Accepted: 7 October 2019; Published: 9 October 2019 Abstract: Bacterial, protozoan and other microbial infections share an accelerated metabolic rate. In order to ensure a proper functioning of cell replication and proteins and nucleic acids synthesis processes, folate metabolism rate is also increased in these cases. For this reason, folic acid antagonists have been used since their discovery to treat different kinds of microbial infections, taking advantage of this metabolic difference when compared with human cells. However, resistances to these compounds have emerged since then and only combined therapies are currently used in clinic. In addition, some of these compounds have been found to have an immunomodulatory behavior that allows clinicians using them as anti-inflammatory or immunosuppressive drugs. Therefore, the aim of this review is to provide an updated state-of-the-art on the use of antifolates as antibacterial and immunomodulating agents in the clinical setting, as well as to present their action mechanisms and currently investigated biomedical applications. Keywords: folic acid antagonists; antifolates; antibiotics; antibacterials; immunomodulation; sulfonamides; antimalarial 1. -

Integrated Molecular Profiling for Analyzing and Predicting Therapeutic Mechanism, Response, Biomarker and Target
INTEGRATED MOLECULAR PROFILING FOR ANALYZING AND PREDICTING THERAPEUTIC MECHANISM, RESPONSE, BIOMARKER AND TARGET Jia Jia (B. Sci & M. Sci, Zhejiang University) A THESIS SUBMITTED FOR THE DEGREE OF DOCTOR OF PHILOSOPHY DEPARTMENT OF PHARMACY NATIONAL UNIVERSITY OF SINGAPORE 2010 Acknowledgements ACKNOWLEDGEMENTS I would like to deeply thank Professor Chen Yu Zong, for his constant encouragement and advice during the entire period of my postgraduate studies. In particular, he has guided me to make my research applicable to the real world problem. This work would not have been possible without his kindness in supporting me to shape up the manuscript for publication. I am also tremendously benefited from his profound knowledge, expertise in scientific research, as well as his enormous support, which will inspire and motivate me to go further in my future professional career. I am also grateful to our BIDD group members for their insight suggestions and collaborations in my research work: Dr. Tang Zhiqun, Ms. Ma Xiaohua, Mr. Zhu Feng, Ms. Liu Xin, Ms. Shi Zhe, Dr. Cui Juan, Mr. Tu Weimin, Dr. Zhang Hailei, Dr. Lin Honghuang, Dr. Liu Xianghui, Dr. Pankaj Kumar, Dr Yap Chun wei, Ms. Wei Xiaona, Ms. Huang Lu, Mr. Zhang Jinxian, Mr. Han Bucong, Mr. Tao Lin, Dr. Wang Rong, Dr. Yan Kun. I thank them for their valuable support and encouragement in my work. Finally, I owe my gratitude to my parents for their forever love, constant support, understanding, encouragement and strength throughout my life. A special appreciation goes to all for love and support. Jia Jia August 2010 I Table of Contents TABLE OF CONTENTS 1.1 Overview of mechanism and strategies of molecular-targeted therapeutics ................................... -
Generate Metabolic Map Poster
Authors: Pallavi Subhraveti Anamika Kothari Quang Ong Ron Caspi An online version of this diagram is available at BioCyc.org. Biosynthetic pathways are positioned in the left of the cytoplasm, degradative pathways on the right, and reactions not assigned to any pathway are in the far right of the cytoplasm. Transporters and membrane proteins are shown on the membrane. Ingrid Keseler Peter D Karp Periplasmic (where appropriate) and extracellular reactions and proteins may also be shown. Pathways are colored according to their cellular function. Csac1394711Cyc: Candidatus Saccharibacteria bacterium RAAC3_TM7_1 Cellular Overview Connections between pathways are omitted for legibility. Tim Holland TM7C00001G0420 TM7C00001G0109 TM7C00001G0953 TM7C00001G0666 TM7C00001G0203 TM7C00001G0886 TM7C00001G0113 TM7C00001G0247 TM7C00001G0735 TM7C00001G0001 TM7C00001G0509 TM7C00001G0264 TM7C00001G0176 TM7C00001G0342 TM7C00001G0055 TM7C00001G0120 TM7C00001G0642 TM7C00001G0837 TM7C00001G0101 TM7C00001G0559 TM7C00001G0810 TM7C00001G0656 TM7C00001G0180 TM7C00001G0742 TM7C00001G0128 TM7C00001G0831 TM7C00001G0517 TM7C00001G0238 TM7C00001G0079 TM7C00001G0111 TM7C00001G0961 TM7C00001G0743 TM7C00001G0893 TM7C00001G0630 TM7C00001G0360 TM7C00001G0616 TM7C00001G0162 TM7C00001G0006 TM7C00001G0365 TM7C00001G0596 TM7C00001G0141 TM7C00001G0689 TM7C00001G0273 TM7C00001G0126 TM7C00001G0717 TM7C00001G0110 TM7C00001G0278 TM7C00001G0734 TM7C00001G0444 TM7C00001G0019 TM7C00001G0381 TM7C00001G0874 TM7C00001G0318 TM7C00001G0451 TM7C00001G0306 TM7C00001G0928 TM7C00001G0622 TM7C00001G0150 TM7C00001G0439 TM7C00001G0233 TM7C00001G0462 TM7C00001G0421 TM7C00001G0220 TM7C00001G0276 TM7C00001G0054 TM7C00001G0419 TM7C00001G0252 TM7C00001G0592 TM7C00001G0628 TM7C00001G0200 TM7C00001G0709 TM7C00001G0025 TM7C00001G0846 TM7C00001G0163 TM7C00001G0142 TM7C00001G0895 TM7C00001G0930 Detoxification Carbohydrate Biosynthesis DNA combined with a 2'- di-trans,octa-cis a 2'- Amino Acid Degradation an L-methionyl- TM7C00001G0190 superpathway of pyrimidine deoxyribonucleotides de novo biosynthesis (E. -

Coordinated Slowing of Metabolism in Enteric Bacteria Under Nitrogen
Coordinated Slowing of Metabolism in Enteric Bacteria under Nitrogen Limitation: A Perspective Ned S. Wingreen NEC Research Institute, 4 Independence Way Princeton, New Jersey 08540 and Department of Physics, University of California Berkeley, CA 94720 Sydney Kustu Department of Plant Biology, Molecular and Cell Biology University of California, Berkeley, CA 94720 Abstract It is natural to ask how bacteria coordinate metabolism when depletion of an essential nutrient limits their growth, and they must slow their entire rate of biosyn- thesis. A major nutrient with a fluctuating abundance is nitrogen. The growth rate of enteric bacteria under nitrogen-limiting conditions is known to correlate with the internal concentration of free glutamine, the glutamine pool. Here we compare the patterns of utilization of L-glutamine and L-glutamate, the two central inter- mediates of nitrogen metabolism. Monomeric precursors of all of the cell’s macro- molecules – proteins, nucleic acids, and surface polymers – require the amide group of glutamine at the first dedicated step of biosynthesis. This is the case even though only a minority (∼12%) of total cell nitrogen derives from glutamine. In contrast, the amino group of glutamate, which provides the remainder of cell nitrogen, is arXiv:physics/0110037v1 [physics.bio-ph] 12 Oct 2001 generally required late in biosynthetic pathways, e.g. in transaminase reactions for amino acid synthesis. We propose that the pattern of glutamine dependence coor- dinates the decrease in biosynthesis under conditions of nitrogen limitation. Hence, the glutamine pool plays a global regulatory role in the cell. 1 INTRODUCTION Enteric bacteria are notable for their varying environment. -

DHFR Inhibitors: Reading the Past for Discovering Novel Anticancer Agents
molecules Review DHFR Inhibitors: Reading the Past for Discovering Novel Anticancer Agents Maria Valeria Raimondi 1,*,† , Ornella Randazzo 1,†, Mery La Franca 1 , Giampaolo Barone 1 , Elisa Vignoni 2, Daniela Rossi 2 and Simona Collina 2,* 1 Department of Biological, Chemical and Pharmaceutical Sciences and Technologies (STEBICEF), University of Palermo, via Archirafi 32, 90123 Palermo, Italy; [email protected] (O.R.); [email protected] (M.L.F.); [email protected] (G.B.) 2 Drug Sciences Department, Medicinal Chemistry and Pharmaceutical Technology Section, University of Pavia, via Taramelli 12, 27100 Pavia, Italy; [email protected] (E.V.); [email protected] (D.R.) * Correspondence: [email protected] (M.V.R.); [email protected] (S.C.); Tel.: +390-912-389-1915 (M.V.R.); +390-382-987-379 (S.C.) † These Authors contributed equally to this work. Academic Editors: Simona Collina and Mariarosaria Miloso Received: 25 February 2019; Accepted: 20 March 2019; Published: 22 March 2019 Abstract: Dihydrofolate reductase inhibitors are an important class of drugs, as evidenced by their use as antibacterial, antimalarial, antifungal, and anticancer agents. Progress in understanding the biochemical basis of mechanisms responsible for enzyme selectivity and antiproliferative effects has renewed the interest in antifolates for cancer chemotherapy and prompted the medicinal chemistry community to develop novel and selective human DHFR inhibitors, thus leading to a new generation of DHFR inhibitors. This work summarizes the mechanism of action, chemical, and anticancer profile of the DHFR inhibitors discovered in the last six years. New strategies in DHFR drug discovery are also provided, in order to thoroughly delineate the current landscape for medicinal chemists interested in furthering this study in the anticancer field. -

Metabolic Characterization of Folate Precursor Paba Uncovers Its Folate Independent Activity on Root Growth of Arabidopsis Thaliana
Inaugural-Dissertation zur Erlangung der Doktorwürde der Albert-Ludwigs-Universität Freiburg im Breisgau Metabolic characterization of folate precursor pABA uncovers its folate independent activity on root growth of Arabidopsis thaliana . Philip Kochersperger Institut für Biologie II (Botanik) September 2011 Dekan: Prof. Dr. Gunther Neuhaus Promotionsvorsitzender: Prof. Dr. Samuel Rossel Betreuer der Arbeit: Prof. Dr. Klaus Palme und Dr. Franck Ditengou Kogutachter: Prof. Dr. Thomas Laux Drittprüfer: Prof. Dr. Ralf Reski Tag der Verkündigung des Prüfungsergebnisses: 04.05.2012 2 I Table of contents I Table of contents ...................................................................................................... 3 II Figure index ............................................................................................................. 6 III Table index ............................................................................................................. 6 IV Abbreviations ......................................................................................................... 7 V Zusammenfassung auf Deutsch ............................................................................ 11 1 Introduction ............................................................................................................ 12 1.1 Abstract ........................................................................................................... 12 1.2 Folates are essential cofactors for almost all living organisms ....................... -

PRT6/At5g02310 Encodes an Arabidopsis Ubiquitin Ligase of the N-End Rule Pathway with Arginine Specificity and Is Not the CER3 Locus
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector FEBS Letters 581 (2007) 3189–3196 PRT6/At5g02310 encodes an Arabidopsis ubiquitin ligase of the N-end rule pathway with arginine specificity and is not the CER3 locus Marcus Garzo´na, Karolin Eiflera, Andrea Faustb, Hartmut Scheelc, Kay Hofmannc, Csaba Koncza, Alexander Yephremovb, Andreas Bachmaira,* a Department of Plant Developmental Biology, Max Planck Institute for Plant Breeding Research, Carl-von-Linne´-Weg 10, D-50829 Cologne, Germany b Department of Molecular Plant Genetics, Max Planck Institute for Plant Breeding Research, Carl-von-Linne´-Weg 10, D-50829 Cologne, Germany c Bioinformatics Group, Miltenyi Biotec GmbH, Stockheimer Weg 1, D-50829 Cologne, Germany Received 10 May 2007; revised 4 June 2007; accepted 5 June 2007 Available online 12 June 2007 Edited by Ulf-Ingo Flu¨gge mini of cellular proteins, do not target these proteins for deg- Abstract The eukaryotic N-end rule pathway mediates ubiqui- tin- and proteasome-dependent turnover of proteins with a bulky radation. Substrates of the eukaryotic N-end rule pathway amino-terminal residue. Arabidopsis locus At5g02310 shows sig- include components of NO signaling in animals [11], and pro- nificant similarity to the yeast N-end rule ligase Ubr1. We dem- teins important for pathogen defense and senescence in plants onstrate that At5g02310 is a ubiquitin ligase and mediates [12,13]. degradation of proteins with amino-terminal Arg residue. Unlike Saccharomyces cerevisiae was found to have a single ubiqui- Ubr1, the Arabidopsis protein does not participate in degrada- tin ligase devoted to the N-end rule, termed Ubr1 [14], whereas tion of proteins with amino-terminal Phe or Leu. -

Regulation of Translation by One-Carbon Metabolism in Bacteria
REVIEWS Regulation of translation by one-carbon metabolism in bacteria and eukaryotic organelles Received for publication, May 12, 2020, and in revised form, November 15, 2020 Published, Papers in Press, November 16, 2020, https://doi.org/10.1074/jbc.REV120.011985 Sunil Shetty1 and Umesh Varshney2,3,* From the 1Biozentrum, University of Basel, Basel, Switzerland; 2Department of Microbiology and Cell Biology, Indian Institute of Science, Bangalore, India; 3Jawaharlal Nehru Centre for Advanced Scientific Studies, Jakkur, Bangalore, India Edited by Karin Musier-Forsyth Protein synthesis is an energetically costly cellular activity. It also employed as antibiotics. The sulfonamides (sulfa drugs) is therefore important that the process of mRNA translation such as sulfachloropyridazine (SCP) and sulfamethoxazole remains in excellent synchrony with cellular metabolism and its (SMX) and trimethoprim (TMP) are perhaps the most promi- energy reserves. Unregulated translation could lead to the pro- nent examples of OCM inhibitors (3). Interestingly, treatment of duction of incomplete, mistranslated, or misfolded proteins, Escherichia coli with TMP inhibited translation within 15 min squandering the energy needed for cellular sustenance and (4). Supplementation of the growth media with the products of causing cytotoxicity. One-carbon metabolism (OCM), an inte- OCM such as methionine, purines, and thymidine could not gral part of cellular intermediary metabolism, produces a num- rescue translation in the presence of TMP, indicating a direct ber of one-carbon unit intermediates (formyl, methylene, contribution of OCM in translation (5, 6). Unfortunately, methenyl, methyl). These OCM intermediates are required for resistance to sulfa drugs arose rapidly (7). However, recent the production of amino acids such as methionine and other studies have shown that OCM inhibitors could serve as effective biomolecules such as purines, thymidylate, and redox regulators. -
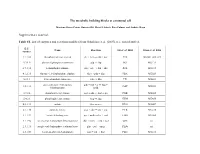
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

Comparative Analysis of the Nodule Transcriptomes of Ceanothus Thyrsiflorus (Rhamnaceae, Rosales) and Datisca Glomerata (Datiscaceae, Cucurbitales)
fpls-09-01629 November 12, 2018 Time: 18:56 # 1 ORIGINAL RESEARCH published: 14 November 2018 doi: 10.3389/fpls.2018.01629 Comparative Analysis of the Nodule Transcriptomes of Ceanothus thyrsiflorus (Rhamnaceae, Rosales) and Datisca glomerata (Datiscaceae, Cucurbitales) Marco G. Salgado1, Robin van Velzen2, Thanh Van Nguyen1, Kai Battenberg3, Alison M. Berry3, Daniel Lundin4,5 and Katharina Pawlowski1* 1 Department of Ecology, Environment and Plant Sciences, Stockholm University, Stockholm, Sweden, 2 Laboratory of Molecular Biology, Department of Plant Sciences, Wageningen University, Wageningen, Netherlands, 3 Department of Plant Sciences, University of California, Davis, Davis, CA, United States, 4 Centre for Ecology and Evolution in Microbial Model Systems, Linnaeus University, Kalmar, Sweden, 5 Department of Biochemistry and Biophysics, Stockholm University, Stockholm, Sweden Edited by: Stefan de Folter, Two types of nitrogen-fixing root nodule symbioses are known, rhizobial and actinorhizal Centro de Investigación y de Estudios symbioses. The latter involve plants of three orders, Fagales, Rosales, and Cucurbitales. Avanzados (CINVESTAV), Mexico To understand the diversity of plant symbiotic adaptation, we compared the nodule Reviewed by: Luis Wall, transcriptomes of Datisca glomerata (Datiscaceae, Cucurbitales) and Ceanothus Universidad Nacional de Quilmes thyrsiflorus (Rhamnaceae, Rosales); both species are nodulated by members of the (UNQ), Argentina Costas Delis, uncultured Frankia clade, cluster II. The analysis focused on various features. In Technological Educational Institute both species, the expression of orthologs of legume Nod factor receptor genes of Peloponnese, Greece was elevated in nodules compared to roots. Since arginine has been postulated as *Correspondence: export form of fixed nitrogen from symbiotic Frankia in nodules of D. glomerata, the Katharina Pawlowski [email protected] question was whether the nitrogen metabolism was similar in nodules of C. -

Thesematthieuvillegente2013annexes
Supplemental table 1: list of the proteins identifed from the total protein fraction of Amborella trichopoda isolated embryo by shot gun Proteins have been analyzed by mono-dimensional electrophoresis and identified by mass spectrometry LC/MS-MS. The protein spots were analysed by LC-MS/MS on the PAPPSO plateform (Benoit Valot, Thierry Balliau, Michel Zivy, INRA Moulon, France; http://pappso.inra.fr). Based on the spectrum generated, proteins were identified using the X-Tandem software. "NCBI accession number" is the accession number in NCBI database. "Protein name" . "Organism" relates to the organism from which the identied protein comes from for functional analysis - efforts were focus on Arabidopsis thaliana as a plant model. " "AGI" . "Function category" and "Function description" relate to the functional categories defined according to the ontological classification of Bevan et al . [Bevan et al . (1998) Nature 391:485-488]. "log(Evalue) identification" reflects the statistical power of the identification by BLAST (performed on TAIR or BLAST). "Identity" indicates in % the recovery of the Amborella protein sequence against the identified protein. "Compartement" indicates if the identification is found in the embryo (emb.), the endosperm (end.) or both (emb./end.). "Description" was taken from the Amborella EVM 27 Predicted Proteins (http://www.amborella.org). The "log (E value)" is a statistical parameter that represents the number of peptides present at random in the database. It was calculated by the product of the Evalue of unique peptides identified in the protein spot (Valot et al., 2011). "Coverage" refers to the recovery rate of the protein by the identified peptides, expressed in %. -

Genetic and Molecular Analysis of Two New Loci Controlling Flowering in Garden Pea
Genetic and molecular analysis of two new loci controlling flowering in garden pea By A S M Mainul Hasan School of Natural Sciences Submitted in fulfilment of the requirements for the degree of Doctor of Philosophy University of Tasmania, July 2018 Declaration of originality This thesis contains no material which has been accepted for a degree or diploma by the University or any other institution, except by way of background information and duly acknowledge in the thesis, and to the best of my knowledge and belief no material previously published or written by another person except where due acknowledgement is made in the text of the thesis, nor does the thesis contain any material that infringes copyright. Authority of access This thesis may be made available for loan. Copying and communication of any part of this thesis is prohibited for two years from the date this statement was signed; after that time limited copying and communication is permitted in accordance with the Copyright Act 1968. Date: 6-07-2018 A S M Mainul Hasan i Abstract Flowering is one of the key developmental process associated with the life cycle of plant and it is regulated by different environmental factors and endogenous cues. In the model species Arabidopsis thaliana a mobile protein, FLOWERING LOCUS T (FT) plays central role to mediate flowering time and expression of FT is regulated by photoperiod. While flowering mechanisms are well-understood in A. thaliana, knowledge about this process is limited in legume (family Fabaceae) which are the second major group of crops after cereals in satisfying the global demand for food and fodder.