A Wave of Nascent Transcription on Activated Human Genes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Análise Integrativa De Perfis Transcricionais De Pacientes Com
UNIVERSIDADE DE SÃO PAULO FACULDADE DE MEDICINA DE RIBEIRÃO PRETO PROGRAMA DE PÓS-GRADUAÇÃO EM GENÉTICA ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Ribeirão Preto – 2012 ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Tese apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo para obtenção do título de Doutor em Ciências. Área de Concentração: Genética Orientador: Prof. Dr. Eduardo Antonio Donadi Co-orientador: Prof. Dr. Geraldo A. S. Passos Ribeirão Preto – 2012 AUTORIZO A REPRODUÇÃO E DIVULGAÇÃO TOTAL OU PARCIAL DESTE TRABALHO, POR QUALQUER MEIO CONVENCIONAL OU ELETRÔNICO, PARA FINS DE ESTUDO E PESQUISA, DESDE QUE CITADA A FONTE. FICHA CATALOGRÁFICA Evangelista, Adriane Feijó Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. Ribeirão Preto, 2012 192p. Tese de Doutorado apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo. Área de Concentração: Genética. Orientador: Donadi, Eduardo Antonio Co-orientador: Passos, Geraldo A. 1. Expressão gênica – microarrays 2. Análise bioinformática por module maps 3. Diabetes mellitus tipo 1 4. Diabetes mellitus tipo 2 5. Diabetes mellitus gestacional FOLHA DE APROVAÇÃO ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. -

A Crosstalk Between the RNA Binding Protein Smaug and the Hedgehog Pathway Links Cell Signaling to Mrna Regulation in Drosophila Lucía Bruzzone
A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila Lucía Bruzzone To cite this version: Lucía Bruzzone. A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila. Cellular Biology. Université Sorbonne Paris Cité, 2018. English. NNT : 2018USPCC234. tel-02899776 HAL Id: tel-02899776 https://tel.archives-ouvertes.fr/tel-02899776 Submitted on 15 Jul 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Thèse de doctorat de l’Université Sorbonne Paris Cité Préparée à l’Université Paris Diderot Ecole doctorale HOB n° 561 Institut Jacques Monod / Equipe Développement, Signalisation et Trafic A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in Drosophila Lucía Bruzzone Thèse de doctorat de Biologie Dirigée par Anne Plessis Présentée et soutenue publiquement à Paris le 19 mars 2018 Président du jury: Alain Zider / Professeur Université Paris Diderot -

Module Analysis Using Single-Patient Differential Expression Signatures
bioRxiv preprint doi: https://doi.org/10.1101/2020.01.05.894931; this version posted January 6, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Module analysis using single-patient differential expression signatures improve the power of association study for Alzheimer's disease Jialan Huang1, Dong Lu1, and Guofeng Meng1,∗ 1Institute of interdisciplinary integrative Medicine Research, shanghai University of Traditional Chinese Medicine, shanghai, China Abstract The causal mechanism of Alzheimer's disease is extremely complex. It usually requires a huge number of samples to achieve a good statistical power in association studies. In this work, we illustrated a different strategy to identify AD risk genes by clustering AD patients into modules based on their single-patient differential expression signatures. Evaluation suggested that our method could enrich AD patients with common clinical manifestations. Applying it to a cohort of only 310 AD patients, we identified 175 AD risk loci at a strict threshold of empirical p < 0:05 while only two loci were identified using all the AD patients. As an evaluation, we collected 23 AD risk genes reported in a recent large-scale meta-analysis and found that 18 of them were re-discovered by association studies using clustered AD patients, while only three of them were re-discovered using all AD patients. Functional annotation suggested that AD associated genetic variants mainly disturbed neuronal/synaptic function. -

Nonspecific Bridging-Induced Attraction Drives Clustering of DNA
Nonspecific bridging-induced attraction drives PNAS PLUS clustering of DNA-binding proteins and genome organization Chris A. Brackleya, Stephen Taylorb, Argyris Papantonisc,d, Peter R. Cookc,1, and Davide Marenduzzoa,1 aScottish Universities Physics Alliance (SUPA), School of Physics, University of Edinburgh, Edinburgh EH9 3JZ, United Kingdom; bComputational Research Biology Group and cSir William Dunn School of Pathology, University of Oxford, Oxford OX1 3RE, United Kingdom; and dCenter for Molecular Medicine, University of Cologne, 50931 Cologne, Germany Edited by Charles S. Peskin, New York University, Manhattan, NY, and approved August 5, 2013 (received for review February 19, 2013) Molecular dynamics simulations are used to model proteins that Results diffuse to DNA, bind, and dissociate; in the absence of any explicit Clustering of Bridging Proteins on Binding to Naked DNA. We first interaction between proteins, or between templates, binding spon- consider a solution of spherical DNA-binding proteins (con- taneously induces local DNA compaction and protein aggregation. centration, 0.02% in volume or 42.5 μM and therefore within the Small bivalent proteins form into rows [as on binding of the bacterial range found in vivo) that bind nonspecifically to naked DNA histone-like nucleoid-structuring protein (H-NS)], large proteins (36.7 kbp), modeled as a semiflexible string of spherical mono- into quasi-spherical aggregates (as on nanoparticle binding), and mers (persistence length, 50 nm) (18) confined within a cube cylinders with eight binding sites (representing octameric nucleo- (250 × 250 × 250 nm). Both proteins and monomers have somal cores) into irregularly folded clusters (like those seen in diameters of 2.5 nm, each DNA monomer represents ∼7.35 bp, nucleosomal strings). -

Supplementary Data
Supplemental figures Supplemental figure 1: Tumor sample selection. A total of 98 thymic tumor specimens were stored in Memorial Sloan-Kettering Cancer Center tumor banks during the study period. 64 cases corresponded to previously untreated tumors, which were resected upfront after diagnosis. Adjuvant treatment was delivered in 7 patients (radiotherapy in 4 cases, cyclophosphamide- doxorubicin-vincristine (CAV) chemotherapy in 3 cases). 34 tumors were resected after induction treatment, consisting of chemotherapy in 16 patients (cyclophosphamide-doxorubicin- cisplatin (CAP) in 11 cases, cisplatin-etoposide (PE) in 3 cases, cisplatin-etoposide-ifosfamide (VIP) in 1 case, and cisplatin-docetaxel in 1 case), in radiotherapy (45 Gy) in 1 patient, and in sequential chemoradiation (CAP followed by a 45 Gy-radiotherapy) in 1 patient. Among these 34 patients, 6 received adjuvant radiotherapy. 1 Supplemental Figure 2: Amino acid alignments of KIT H697 in the human protein and related orthologs, using (A) the Homologene database (exons 14 and 15), and (B) the UCSC Genome Browser database (exon 14). Residue H697 is highlighted with red boxes. Both alignments indicate that residue H697 is highly conserved. 2 Supplemental Figure 3: Direct comparison of the genomic profiles of thymic squamous cell carcinomas (n=7) and lung primary squamous cell carcinomas (n=6). (A) Unsupervised clustering analysis. Gains are indicated in red, and losses in green, by genomic position along the 22 chromosomes. (B) Genomic profiles and recurrent copy number alterations in thymic carcinomas and lung squamous cell carcinomas. Gains are indicated in red, and losses in blue. 3 Supplemental Methods Mutational profiling The exonic regions of interest (NCBI Human Genome Build 36.1) were broken into amplicons of 500 bp or less, and specific primers were designed using Primer 3 (on the World Wide Web for general users and for biologist programmers (see Supplemental Table 2) [1]. -

UNIVERSITY of CALIFORNIA Los Angeles a Sterile Alpha Motif
UNIVERSITY OF CALIFORNIA Los Angeles A Sterile Alpha Motif Domain Network Involved in Kidney Development A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Biochemistry and Molecular Biology by Catherine Nicole Leettola 2015 ABSTRACT OF THE DISSERTATION A Sterile Alpha Motif Domain Network Involved in Kidney Development by Catherine Nicole Leettola Doctor of Philosophy in Biochemistry and Molecular Biology University of California, Los Angeles, 2015 Professor James U. Bowie, Chair Cystic kidney diseases including polycystic kidney disease (PKD) and nephronophthisis (NPHP) are the most common genetic disorders leading to end-stage renal failure in humans. Animal models and human cases of PKD and NPHP have implicated the sterile alpha motif (SAM) domain containing proteins bicaudal C homolog 1 (BICC1) and ankyrin repeat and SAM- domain containing protein 6 (ANKS6) as being involved in these conditions and important for renal development. SAM domains are known protein-protein interaction domains that are capable of binding each other to form polymers and heterodimers. Using a negGFP native gel assay, we have identified the SAM domain of the previously uncharacterized protein ankyrin repeat and SAM-domain containing protein 3 (ANKS3) as a direct binding partner of the BICC1 and ANKS6 SAM domains. We found the ANKS3 SAM domain to polymerize with moderate affinity and determined the ANKS6 SAM domain can bind to a single end of this polymer. Crystal structures of the ANKS3 SAM domain polymer and the ANKS3 SAM-ANKS6 SAM ii heterodimer are presented to reveal typical ML-EH SAM domain interaction interfaces with a pronounced charge complementarity. -

Heterogeneity Between Primary Colon Carcinoma and Paired Lymphatic and Hepatic Metastases
MOLECULAR MEDICINE REPORTS 6: 1057-1068, 2012 Heterogeneity between primary colon carcinoma and paired lymphatic and hepatic metastases HUANRONG LAN1, KETAO JIN2,3, BOJIAN XIE4, NA HAN5, BINBIN CUI2, FEILIN CAO2 and LISONG TENG3 Departments of 1Gynecology and Obstetrics, and 2Surgical Oncology, Taizhou Hospital, Wenzhou Medical College, Linhai, Zhejiang; 3Department of Surgical Oncology, First Affiliated Hospital, College of Medicine, Zhejiang University, Hangzhou, Zhejiang; 4Department of Surgical Oncology, Sir Run Run Shaw Hospital, College of Medicine, Zhejiang University, Hangzhou, Zhejiang; 5Cancer Chemotherapy Center, Zhejiang Cancer Hospital, Zhejiang University of Chinese Medicine, Hangzhou, Zhejiang, P.R. China Received January 26, 2012; Accepted May 8, 2012 DOI: 10.3892/mmr.2012.1051 Abstract. Heterogeneity is one of the recognized characteris- Introduction tics of human tumors, and occurs on multiple levels in a wide range of tumors. A number of studies have focused on the Intratumor heterogeneity is one of the recognized charac- heterogeneity found in primary tumors and related metastases teristics of human tumors, which occurs on multiple levels, with the consideration that the evaluation of metastatic rather including genetic, protein and macroscopic, in a wide range than primary sites could be of clinical relevance. Numerous of tumors, including breast, colorectal cancer (CRC), non- studies have demonstrated particularly high rates of hetero- small cell lung cancer (NSCLC), prostate, ovarian, pancreatic, geneity between primary colorectal tumors and their paired gastric, brain and renal clear cell carcinoma (1). Over the past lymphatic and hepatic metastases. It has also been proposed decade, a number of studies have focused on the heterogeneity that the heterogeneity between primary colon carcinomas and found in primary tumors and related metastases with the their paired lymphatic and hepatic metastases may result in consideration that the evaluation of metastatic rather than different responses to anticancer therapies. -

CREB-Dependent Transcription in Astrocytes: Signalling Pathways, Gene Profiles and Neuroprotective Role in Brain Injury
CREB-dependent transcription in astrocytes: signalling pathways, gene profiles and neuroprotective role in brain injury. Tesis doctoral Luis Pardo Fernández Bellaterra, Septiembre 2015 Instituto de Neurociencias Departamento de Bioquímica i Biologia Molecular Unidad de Bioquímica y Biologia Molecular Facultad de Medicina CREB-dependent transcription in astrocytes: signalling pathways, gene profiles and neuroprotective role in brain injury. Memoria del trabajo experimental para optar al grado de doctor, correspondiente al Programa de Doctorado en Neurociencias del Instituto de Neurociencias de la Universidad Autónoma de Barcelona, llevado a cabo por Luis Pardo Fernández bajo la dirección de la Dra. Elena Galea Rodríguez de Velasco y la Dra. Roser Masgrau Juanola, en el Instituto de Neurociencias de la Universidad Autónoma de Barcelona. Doctorando Directoras de tesis Luis Pardo Fernández Dra. Elena Galea Dra. Roser Masgrau In memoriam María Dolores Álvarez Durán Abuela, eres la culpable de que haya decidido recorrer el camino de la ciencia. Que estas líneas ayuden a conservar tu recuerdo. A mis padres y hermanos, A Meri INDEX I Summary 1 II Introduction 3 1 Astrocytes: physiology and pathology 5 1.1 Anatomical organization 6 1.2 Origins and heterogeneity 6 1.3 Astrocyte functions 8 1.3.1 Developmental functions 8 1.3.2 Neurovascular functions 9 1.3.3 Metabolic support 11 1.3.4 Homeostatic functions 13 1.3.5 Antioxidant functions 15 1.3.6 Signalling functions 15 1.4 Astrocytes in brain pathology 20 1.5 Reactive astrogliosis 22 2 The transcription -
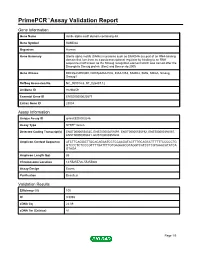
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name sterile alpha motif domain containing 4A Gene Symbol SAMD4A Organism Human Gene Summary Sterile alpha motifs (SAMs) in proteins such as SAMD4A are part of an RNA-binding domain that functions as a posttranscriptional regulator by binding to an RNA sequence motif known as the Smaug recognition element which was named after the Drosophila Smaug protein (Baez and Boccaccio 2005 Gene Aliases DKFZp434H0350, DKFZp686A1532, KIAA1053, SAMD4, SMG, SMGA, Smaug, Smaug1 RefSeq Accession No. NC_000014.8, NT_026437.12 UniGene ID Hs.98259 Ensembl Gene ID ENSG00000020577 Entrez Gene ID 23034 Assay Information Unique Assay ID qHsaCED0003246 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000554335, ENST00000251091, ENST00000555192, ENST00000392067, ENST00000305831, ENST00000357634 Amplicon Context Sequence ATCTTCAGGGTTGCACAGAATCCTCCAAGATACTTTGCAGCCTTTTTTCCCCCTG GTCCCTCTCCCGTTTTGATTTTGTGAGAGCGTAGGTCATCCTCGTAAACATATCA GTAGA Amplicon Length (bp) 85 Chromosome Location 14:55255726-55255840 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 100 R2 0.9998 cDNA Cq 22.09 cDNA Tm (Celsius) 81 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq 23.37 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report SAMD4A, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard -

MAFB Determines Human Macrophage Anti-Inflammatory
MAFB Determines Human Macrophage Anti-Inflammatory Polarization: Relevance for the Pathogenic Mechanisms Operating in Multicentric Carpotarsal Osteolysis This information is current as of October 4, 2021. Víctor D. Cuevas, Laura Anta, Rafael Samaniego, Emmanuel Orta-Zavalza, Juan Vladimir de la Rosa, Geneviève Baujat, Ángeles Domínguez-Soto, Paloma Sánchez-Mateos, María M. Escribese, Antonio Castrillo, Valérie Cormier-Daire, Miguel A. Vega and Ángel L. Corbí Downloaded from J Immunol 2017; 198:2070-2081; Prepublished online 16 January 2017; doi: 10.4049/jimmunol.1601667 http://www.jimmunol.org/content/198/5/2070 http://www.jimmunol.org/ Supplementary http://www.jimmunol.org/content/suppl/2017/01/15/jimmunol.160166 Material 7.DCSupplemental References This article cites 69 articles, 22 of which you can access for free at: http://www.jimmunol.org/content/198/5/2070.full#ref-list-1 by guest on October 4, 2021 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2017 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. -

Exploiting Native Forces to Capture Chromosome Conformation in Mammalian Cell Nuclei
Edinburgh Research Explorer Exploiting native forces to capture chromosome conformation in mammalian cell nuclei Citation for published version: Brant, L, Georgomanolis, T, Nikolic, M, Brackley, CA, Kolovos, P, van Ijcken, W, Grosveld, FG, Marenduzzo, D & Papantonis, A 2016, 'Exploiting native forces to capture chromosome conformation in mammalian cell nuclei', Molecular Systems Biology, vol. 12, no. 12, pp. 891. https://doi.org/10.15252/msb.20167311 Digital Object Identifier (DOI): 10.15252/msb.20167311 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Molecular Systems Biology General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 07. Oct. 2021 Published online: December 9, 2016 Report Exploiting native forces to capture chromosome conformation in mammalian cell nuclei Lilija Brant1, Theodore Georgomanolis1,†, Milos Nikolic1,†, Chris A Brackley2, Petros Kolovos3, Wilfred van Ijcken4 , Frank G Grosveld3, Davide Marenduzzo2 & Argyris Papantonis1,* Abstract et al, 2013; Pombo & Dillon, 2015; Denker & de Laat, 2016) and allowed us to address diverse biological questions (e.g., Tolhuis et al, Mammalian interphase chromosomes fold into a multitude of 2002; Papantonis et al, 2012; Zhang et al, 2012; Naumova et al, loops to fit the confines of cell nuclei, and looping is tightly 2013; Rao et al, 2014). -

UTR Shortening Has a Limited Effect on Protein Abundance in Proliferating T Cells
ARTICLE Received 17 Apr 2014 | Accepted 3 Oct 2014 | Published 21 Nov 2014 DOI: 10.1038/ncomms6465 Global 30 UTR shortening has a limited effect on protein abundance in proliferating T cells Andreas R. Gruber1,*, Georges Martin1,*, Philipp Mu¨ller2,3, Alexander Schmidt4, Andreas J. Gruber1, Rafal Gumienny1, Nitish Mittal1, Rajesh Jayachandran3, Jean Pieters3, Walter Keller1, Erik van Nimwegen1 & Mihaela Zavolan1 Alternative polyadenylation is a cellular mechanism that generates mRNA isoforms differing in their 30 untranslated regions (30 UTRs). Changes in polyadenylation site usage have been described upon induction of proliferation in resting cells, but the underlying mechanism and functional significance of this phenomenon remain largely unknown. To understand the functional consequences of shortened 30 UTR isoforms in a physiological setting, we used 30 end sequencing and quantitative mass spectrometry to determine polyadenylation site usage, mRNA and protein levels in murine and human naive and activated T cells. Although 30 UTR shortening in proliferating cells is conserved between human and mouse, orthologous genes do not exhibit similar expression of alternative 30 UTR isoforms. We generally find that 30 UTR shortening is not accompanied by a corresponding change in mRNA and protein levels. This suggests that although 30 UTR shortening may lead to changes in the RNA-binding protein interactome, it has limited effects on protein output. 1 Computational and Systems Biology, Biozentrum, University of Basel and Swiss Institute of Bioinformatics, Basel CH-4056, Switzerland. 2 Department of Biomedicine, University of Basel and University Hospital Basel, Hebelstrasse 20, Basel CH-4031, Switzerland. 3 Infection Biology, Biozentrum, University of Basel, Basel CH-4056, Switzerland.