The Evolution of Ogres: Cannibalistic Growth in Giant Phagotrophs
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Evolutionary Trajectories Explain the Diversified Evolution of Isogamy And
Evolutionary trajectories explain the diversified evolution of isogamy and anisogamy in marine green algae Tatsuya Togashia,b,c,1, John L. Barteltc,d, Jin Yoshimuraa,e,f, Kei-ichi Tainakae, and Paul Alan Coxc aMarine Biosystems Research Center, Chiba University, Kamogawa 299-5502, Japan; bPrecursory Research for Embryonic Science and Technology, Japan Science and Technology Agency, Kawaguchi 332-0012, Japan; cInstitute for Ethnomedicine, Jackson Hole, WY 83001; dEvolutionary Programming, San Clemente, CA 92673; eDepartment of Systems Engineering, Shizuoka University, Hamamatsu 432-8561, Japan; and fDepartment of Environmental and Forest Biology, State University of New York College of Environmental Science and Forestry, Syracuse, NY 13210 Edited by Geoff A. Parker, University of Liverpool, Liverpool, United Kingdom, and accepted by the Editorial Board July 12, 2012 (received for review March 1, 2012) The evolution of anisogamy (the production of gametes of dif- (11) suggested that the “gamete size model” (Parker, Baker, and ferent size) is the first step in the establishment of sexual dimorphism, Smith’s model) is the most applicable to fungi (11). However, and it is a fundamental phenomenon underlying sexual selection. none of these models successfully account for the evolution of It is believed that anisogamy originated from isogamy (production all known forms of isogamy and anisogamy in extant marine of gametes of equal size), which is considered by most theorists to green algae (12). be the ancestral condition. Although nearly all plant and animal Marine green algae are characterized by a variety of mating species are anisogamous, extant species of marine green algae systems linked to their habitats (Fig. -

Quantitative Evolutionary Analysis of the Life Cycle of Social Amoebae Darja Dubravcic
Quantitative evolutionary analysis of the life cycle of social amoebae Darja Dubravcic To cite this version: Darja Dubravcic. Quantitative evolutionary analysis of the life cycle of social amoebae. Agricultural sciences. Université René Descartes - Paris V, 2013. English. NNT : 2013PA05T033. tel-00914467 HAL Id: tel-00914467 https://tel.archives-ouvertes.fr/tel-00914467 Submitted on 5 Dec 2013 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Université Paris Descartes Ecole doctorale « Interdisciplinaire Européen Frontières de vivant » Laboratory « Ecology & Evolution » UMR7625 Laboratory of Interdisciplinary Physics UMR5588 Quantitative evolutionary analysis of the life cycle of social amoebae By Darja Dubravcic PhD Thesis in: Evolutionary Biology Directed by Minus van Baalen and Clément Nizak Presented on the 15th November 2013 PhD committee: Dr. M. van Baalen, PhD director Dr. C. Nizak, PhD Co-director Prof. V. Nanjundiah, Reviewer Prof. P. Rainey, Reviewer Prof. A. Gardner Prof. J-P. Rieu Prof. J-M Di Meglio Dr. S. de Monte, Invited 2 Abstract Social amoebae are eukaryotic organisms that inhabit soil of almost every climate zone. They are remarkable for their switch from unicellularity to multicellularity as an adaptation to starvation. -

Protozoologica Special Issue: Protists in Soil Processes
Acta Protozool. (2012) 51: 201–208 http://www.eko.uj.edu.pl/ap ActA doi:10.4467/16890027AP.12.016.0762 Protozoologica Special issue: Protists in Soil Processes Review paper Ecology of Soil Eumycetozoans Steven L. STEPHENSON1 and Alan FEEST2 1Department of Biological Sciences, University of Arkansas, Fayetteville, Arkansas, USA; 2Institute of Advanced Studies, University of Bristol and Ecosulis ltd., Newton St Loe, Bath, United Kingdom Abstract. Eumycetozoans, commonly referred to as slime moulds, are common to abundant organisms in soils. Three groups of slime moulds (myxogastrids, dictyostelids and protostelids) are recognized, and the first two of these are among the most important bacterivores in the soil microhabitat. The purpose of this paper is first to provide a brief description of all three groups and then to review what is known about their distribution and ecology in soils. Key words: Amoebae, bacterivores, dictyostelids, myxogastrids, protostelids. INTRODUCTION that they are amoebozoans and not fungi (Bapteste et al. 2002, Yoon et al. 2008, Baudalf 2008). Three groups of slime moulds (myxogastrids, dic- One of the idiosyncratic branches of the eukary- tyostelids and protostelids) are recognized (Olive 1970, otic tree of life consists of an assemblage of amoe- 1975). Members of the three groups exhibit consider- boid protists referred to as the supergroup Amoebozoa able diversity in the type of aerial spore-bearing struc- (Fiore-Donno et al. 2010). The most diverse members tures produced, which can range from exceedingly of the Amoebozoa are the eumycetozoans, common- small examples (most protostelids) with only a single ly referred to as slime moulds. Since their discovery, spore to the very largest examples (certain myxogas- slime moulds have been variously classified as plants, trids) that contain many millions of spores. -

Fold-Change Detection and Scale Invariance of Cell–Cell Signaling In
Fold-change detection and scale invariance of cell–cell PNAS PLUS signaling in social amoeba Keita Kaminoa,1, Yohei Kondoa, Akihiko Nakajimab, Mai Honda-Kitaharaa, Kunihiko Kanekoa,b, and Satoshi Sawaia,b,c,1 aDepartment of Basic Science, Graduate School of Arts and Sciences, University of Tokyo, Tokyo 153-8902, Japan; bResearch Center for Complex Systems Biology, University of Tokyo, Tokyo 153-8902, Japan; and cPrecursory Research for Embryonic Science and Technology, Japan Science and Technology Agency, Saitama 332-0012, Japan Edited by Peter N. Devreotes, The Johns Hopkins University School of Medicine, Baltimore, MD, and approved April 9, 2017 (received for review February 9, 2017) Cell–cell signaling is subject to variability in the extracellular volume, process called “cAMP relay” (13). After prolonged exposure to cell number, and dilution that potentially increase uncertainty in the cAMP, the rise in extracellular cAMP level ceases due to inacti- absolute concentrations of the extracellular signaling molecules. To vation of adenylyl cyclase (14). As extracellular cAMP level is direct cell aggregation, the social amoebae Dictyostelium discoideum lowered by degradation, the cells exit from the state of reduced collectively give rise to oscillations and waves of cyclic adenosine responsivity over the course of several minutes (15, 16), and hence 3′,5′-monophosphate (cAMP) under a wide range of cell density. To the extracellular cAMP level once again starts to elevate. This date, the systems-level mechanism underlying the robustness is un- tendency for the extracellular cAMP level to rise when it is lowered, clear. By using quantitative live-cell imaging, here we show that the and to be lowered when it is raised, essentially renders extracellular magnitude of the cAMP relay response of individual cells is deter- cAMP level unstable and oscillatory. -

Comparative Genomics of the Social Amoebae Dictyostelium Discoideum
Sucgang et al. Genome Biology 2011, 12:R20 http://genomebiology.com/2011/12/2/R20 RESEARCH Open Access Comparative genomics of the social amoebae Dictyostelium discoideum and Dictyostelium purpureum Richard Sucgang1†, Alan Kuo2†, Xiangjun Tian3†, William Salerno1†, Anup Parikh4, Christa L Feasley5, Eileen Dalin2, Hank Tu2, Eryong Huang4, Kerrie Barry2, Erika Lindquist2, Harris Shapiro2, David Bruce2, Jeremy Schmutz2, Asaf Salamov2, Petra Fey6, Pascale Gaudet6, Christophe Anjard7, M Madan Babu8, Siddhartha Basu6, Yulia Bushmanova6, Hanke van der Wel5, Mariko Katoh-Kurasawa4, Christopher Dinh1, Pedro M Coutinho9, Tamao Saito10, Marek Elias11, Pauline Schaap12, Robert R Kay8, Bernard Henrissat9, Ludwig Eichinger13, Francisco Rivero14, Nicholas H Putnam3, Christopher M West5, William F Loomis7, Rex L Chisholm6, Gad Shaulsky3,4, Joan E Strassmann3, David C Queller3, Adam Kuspa1,3,4* and Igor V Grigoriev2 Abstract Background: The social amoebae (Dictyostelia) are a diverse group of Amoebozoa that achieve multicellularity by aggregation and undergo morphogenesis into fruiting bodies with terminally differentiated spores and stalk cells. There are four groups of dictyostelids, with the most derived being a group that contains the model species Dictyostelium discoideum. Results: We have produced a draft genome sequence of another group dictyostelid, Dictyostelium purpureum, and compare it to the D. discoideum genome. The assembly (8.41 × coverage) comprises 799 scaffolds totaling 33.0 Mb, comparable to the D. discoideum genome size. Sequence comparisons suggest that these two dictyostelids shared a common ancestor approximately 400 million years ago. In spite of this divergence, most orthologs reside in small clusters of conserved synteny. Comparative analyses revealed a core set of orthologous genes that illuminate dictyostelid physiology, as well as differences in gene family content. -

The Closest Unicellular Relatives of Animals
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Current Biology, Vol. 12, 1773–1778, October 15, 2002, 2002 Elsevier Science Ltd. All rights reserved. PII S0960-9822(02)01187-9 The Closest Unicellular Relatives of Animals B.F. Lang,1,2 C. O’Kelly,1,3 T. Nerad,4 M.W. Gray,1,5 Results and Discussion and G. Burger1,2,6 1The Canadian Institute for Advanced Research The evolution of the Metazoa from single-celled protists Program in Evolutionary Biology is an issue that has intrigued biologists for more than a 2 De´ partement de Biochimie century. Early morphological and more recent ultra- Universite´ de Montre´ al structural and molecular studies have converged in sup- Succursale Centre-Ville porting the now widely accepted view that animals are Montre´ al, Que´ bec H3C 3J7 related to Fungi, choanoflagellates, and ichthyosporean Canada protists. However, controversy persists as to the spe- 3 Bigelow Laboratory for Ocean Sciences cific evolutionary relationships among these major P.O. Box 475 groups. This uncertainty is reflected in the plethora of 180 McKown Point Road published molecular phylogenies that propose virtually West Boothbay Harbor, Maine 04575 all of the possible alternative tree topologies involving 4 American Type Culture Collection Choanoflagellata, Fungi, Ichthyosporea, and Metazoa. 10801 University Boulevard For example, a monophyletic MetazoaϩChoanoflagel- Manassas, Virginia 20110 lata group has been suggested on the basis of small 5 Department of Biochemistry subunit (SSU) rDNA sequences [1, 6, 7]. Other studies and Molecular Biology using the same sequences have allied Choanoflagellata Dalhousie University with the Fungi [8], placed Choanoflagellata prior to the Halifax, Nova Scotia B3H 4H7 divergence of animals and Fungi [9], or even placed Canada them prior to the divergence of green algae and land plants [10]. -

Ptolemeba N. Gen., a Novel Genus of Hartmannellid Amoebae (Tubulinea, Amoebozoa); with an Emphasis on the Taxonomy of Saccamoeba
The Journal of Published by the International Society of Eukaryotic Microbiology Protistologists Journal of Eukaryotic Microbiology ISSN 1066-5234 ORIGINAL ARTICLE Ptolemeba n. gen., a Novel Genus of Hartmannellid Amoebae (Tubulinea, Amoebozoa); with an Emphasis on the Taxonomy of Saccamoeba Pamela M. Watsona, Stephanie C. Sorrella & Matthew W. Browna,b a Department of Biological Sciences, Mississippi State University, Mississippi State, Mississippi, 39762 b Institute for Genomics, Biocomputing & Biotechnology, Mississippi State University, Mississippi State, Mississippi, 39762 Keywords ABSTRACT 18S rRNA; amoeba; amoeboid; Cashia; cristae; freshwater amoebae; Hartmannella; Hartmannellid amoebae are an unnatural assemblage of amoeboid organisms mitochondrial morphology; SSU rDNA; SSU that are morphologically difficult to discern from one another. In molecular phy- rRNA; terrestrial amoebae; tubulinid. logenetic trees of the nuclear-encoded small subunit rDNA, they occupy at least five lineages within Tubulinea, a well-supported clade in Amoebozoa. The Correspondence polyphyletic nature of the hartmannellids has led to many taxonomic problems, M.W. Brown, Department of Biological in particular paraphyletic genera. Recent taxonomic revisions have alleviated Sciences, Mississippi State University, some of the problems. However, the genus Saccamoeba is paraphyletic and is Mississippi State, MS 39762, USA still in need of revision as it currently occupies two distinct lineages. Here, we Telephone number: +1 662-325-2406; report a new clade on the tree of Tubulinea, which we infer represents a novel FAX number: +1 662-325-7939; genus that we name Ptolemeba n. gen. This genus subsumes a clade of hart- e-mail: [email protected] mannellid amoebae that were previously considered in the genus Saccamoeba, but whose mitochondrial morphology is distinct from Saccamoeba. -

Zygote Gene Expression and Plasmodial Development in Didymium Iridis
DePaul University Via Sapientiae College of Science and Health Theses and Dissertations College of Science and Health Summer 8-25-2019 Zygote gene expression and plasmodial development in Didymium iridis Sean Schaefer DePaul University, [email protected] Follow this and additional works at: https://via.library.depaul.edu/csh_etd Part of the Biology Commons Recommended Citation Schaefer, Sean, "Zygote gene expression and plasmodial development in Didymium iridis" (2019). College of Science and Health Theses and Dissertations. 322. https://via.library.depaul.edu/csh_etd/322 This Thesis is brought to you for free and open access by the College of Science and Health at Via Sapientiae. It has been accepted for inclusion in College of Science and Health Theses and Dissertations by an authorized administrator of Via Sapientiae. For more information, please contact [email protected]. Zygote gene expression and plasmodial development in Didymium iridis A Thesis presented in Partial fulfillment of the Requirements for the Degree of Master of Biology By Sean Schaefer 2019 Advisor: Dr. Margaret Silliker Department of Biological Sciences College of Liberal Arts and Sciences DePaul University Chicago, IL Abstract: Didymium iridis is a cosmopolitan species of plasmodial slime mold consisting of two distinct life stages. Haploid amoebae and diploid plasmodia feed on microscopic organisms such as bacteria and fungi through phagocytosis. Sexually compatible haploid amoebae act as gametes which when fused embark on an irreversible developmental change resulting in a diploid zygote. The zygote can undergo closed mitosis resulting in a multinucleated plasmodium. Little is known about changes in gene expression during this developmental transition. Our principal goal in this study was to provide a comprehensive list of genes likely to be involved in plasmodial development. -

Myxomycetes NMW 2012Orange, Updated KS 2017.Docx
Myxomycete (Slime Mould) Collection Amgueddfa Cymru-National Museum Wales (NMW) Alan Orange (2012), updated by Katherine Slade (2017) Myxomycetes (true or plasmodial slime moulds) belong to the Eumycetozoa, within the Amoebozoa, a group of eukaryotes that are basal to a clade containing animals and fungi. Thus although they have traditionally been studied by mycologists they are distant from the true fungi. Arrangement & Nomenclature Slime Mould specimens in NMW are arranged in alphabetical order of the currently accepted name (as of 2012). Names used on specimen packets that are now synonyms are cross referenced in the list below. The collection currently contains 157 Myxomycete species. Specimens are mostly from Britain, with a few from other parts of Europe or from North America. The current standard work for identification of the British species is: Ing, B. 1999. The Myxomycetes of Britain and Ireland. An Identification Handbook. Slough: Richmond Publishing Co. Ltd. Nomenclature follows the online database of Slime Mould names at www.eumycetozoa.com (accessed 2012). This database is largely in line with Ing (1999). Preservation The feeding stage is a multinucleate motile mass known as a plasmodium. The fruiting stage is a dry, fungus-like structure containing abundant spores. Mature fruiting bodies of Myxomycetes can be collected and dried, and with few exceptions (such as Ceratiomyxa) they preserve well. Plasmodia cannot be preserved, but it is useful to record the colour if possible. Semi-mature fruiting bodies may continue to mature if collected with the substrate and kept in a cool moist chamber. Collected plasmodia are unlikely to fruit. Specimens are stored in boxes to prevent crushing; labels should not be allowed to touch the specimen. -

A Novel Structure Learning Algorithm for Bayesian Networks Inspired by Physarum Polycephalum
Physarum Learner: A Novel Structure Learning Algorithm for Bayesian Networks inspired by Physarum Polycephalum DISSERTATION ZUR ERLANGUNG DES DOKTORGRADES DER NATURWISSENSCHAFTEN (DR. RER. NAT.) DER FAKULTAT¨ FUR¨ BIOLOGIE UND VORKLINISCHE MEDIZIN DER UNIVERSITAT¨ REGENSBURG vorgelegt von Torsten Schon¨ aus Wassertrudingen¨ im Jahr 2013 Der Promotionsgesuch wurde eingereicht am: 21.05.2013 Die Arbeit wurde angeleitet von: Prof. Dr. Elmar W. Lang Unterschrift: Torsten Schon¨ ii iii Abstract Two novel algorithms for learning Bayesian network structure from data based on the true slime mold Physarum polycephalum are introduced. The first algorithm called C- PhyL calculates pairwise correlation coefficients in the dataset. Within an initially fully connected Physarum-Maze, the length of the connections is given by the inverse correla- tion coefficient between the connected nodes. Then, the shortest indirect path between each two nodes is determined using the Physarum Solver. In each iteration, a score of the surviving edges is increased. Based on that score, the highest ranked connections are combined to form a Bayesian network. The novel C-PhyL method is evaluated with different configurations and compared to the LAGD Hill Climber, Tabu Search and Simu- lated Annealing on a set of artificially generated and real benchmark networks of different characteristics, showing comparable performance regarding quality of training results and increased time efficiency for large datasets. The second novel algorithm called SO-PhyL is introduced and shown to be able to out- perform common score based structure learning algorithms for some benchmark datasets. SO-PhyL first initializes a fully connected Physarum-Maze with constant length and ran- dom conductivities. In each Physarum Solver iteration, the source and sink nodes are changed randomly and the conductivities are updated. -

Dictyostelid Cellular Slime Molds from Caves
John C. Landolt, Steven L. Stephenson, and Michael E. Slay – Dictyostelid cellular slime molds from caves. Journal of Cave and Karst Studies, v. 68, no. 1, p. 22–26. DICTYOSTELID CELLULAR SLIME MOLDS FROM CAVES JOHN C. LANDOLT Department of Biology, Shepherd University, Shepherdstown, WV 2544 USA [email protected] STEVEN L. STEPHENSON Department of Biological Sciences, University of Arkansas, Fayetteville, AR 72701 USA [email protected] MICHAEL E. SLAY The Nature Conservancy, 601 North University Avenue, Little Rock, AR 72205 USA [email protected] Dictyostelid cellular slime molds associated with caves in Alabama, Arkansas, Indiana, Missouri, New York, Oklahoma, South Carolina, Tennessee, West Virginia, Puerto Rico, and San Salvador in the Bahamas were investigated during the period of 1990–2005. Samples of soil material collected from more than 100 caves were examined using standard methods for isolating dictyostelids. At least 17 species were recovered, along with a number of isolates that could not be identified completely. Four cos- mopolitan species (Dictyostelium sphaerocephalum, D. mucoroides, D. giganteum and Polysphondylium violaceum) and one species (D. rosarium) with a more restricted distribution were each recorded from more than 25 different caves, but three other species were present in more than 20 caves. The data gen- erated in the present study were supplemented with all known published and unpublished records of dic- tyostelids from caves in an effort to summarize what is known about their occurrence in this habitat. INTRODUCTION also occur on dung and were once thought to be primarily coprophilous (Raper, 1984). However, perhaps the most Dictyostelid cellular slime molds (dictyostelids) are single- unusual microhabitat for dictyostelids is the soil material celled, eukaryotic, phagotrophic bacterivores usually present found in caves. -
Supplementary Material Parameter Unit Average ± Std NO3 + NO2 Nm
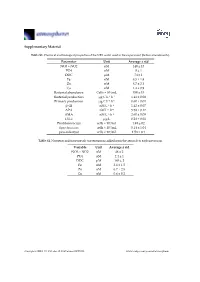
Supplementary Material Table S1. Chemical and biological properties of the NRS water used in the experiment (before amendments). Parameter Unit Average ± std NO3 + NO2 nM 140 ± 13 PO4 nM 8 ± 1 DOC μM 74 ± 1 Fe nM 8.5 ± 1.8 Zn nM 8.7 ± 2.1 Cu nM 1.4 ± 0.9 Bacterial abundance Cells × 104/mL 350 ± 15 Bacterial production μg C L−1 h−1 1.41 ± 0.08 Primary production μg C L−1 h−1 0.60 ± 0.01 β-Gl nM L−1 h−1 1.42 ± 0.07 APA nM L−1 h−1 5.58 ± 0.17 AMA nM L−1·h−1 2.60 ± 0.09 Chl-a μg/L 0.28 ± 0.01 Prochlorococcus cells × 104/mL 1.49 ± 02 Synechococcus cells × 104/mL 5.14 ± 1.04 pico-eukaryot cells × 103/mL 1.58 × 0.1 Table S2. Nutrients and trace metals concentrations added from the aerosols to each mesocosm. Variable Unit Average ± std NO3 + NO2 nM 48 ± 2 PO4 nM 2.4 ± 1 DOC μM 165 ± 2 Fe nM 2.6 ± 1.5 Zn nM 6.7 ± 2.5 Cu nM 0.6 ± 0.2 Atmosphere 2019, 10, 358; doi:10.3390/atmos10070358 www.mdpi.com/journal/atmosphere Atmosphere 2019, 10, 358 2 of 6 Table S3. ANOVA test results between control, ‘UV-treated’ and ‘live-dust’ treatments at 20 h or 44 h, with significantly different values shown in bold. ANOVA df Sum Sq Mean Sq F Value p-value Chl-a 20 H 2, 6 0.03, 0.02 0.02, 0 4.52 0.0634 44 H 2, 6 0.02, 0 0.01, 0 23.13 0.002 Synechococcus Abundance 20 H 2, 7 8.23 × 107, 4.11 × 107 4.11 × 107, 4.51 × 107 0.91 0.4509 44 H 2, 7 5.31 × 108, 6.97 × 107 2.65 × 108, 1.16 × 107 22.84 0.0016 Prochlorococcus Abundance 20 H 2, 8 4.22 × 107, 2.11 × 107 2.11 × 107, 2.71 × 106 7.77 0.0216 44 H 2, 8 9.02 × 107, 1.47 × 107 4.51 × 107, 2.45 × 106 18.38 0.0028 Pico-eukaryote