Dinophyceae Can Use Exudates As Weapons Against the Parasite Amoebophrya Sp
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
Molecular Data and the Evolutionary History of Dinoflagellates by Juan Fernando Saldarriaga Echavarria Diplom, Ruprecht-Karls-Un
Molecular data and the evolutionary history of dinoflagellates by Juan Fernando Saldarriaga Echavarria Diplom, Ruprecht-Karls-Universitat Heidelberg, 1993 A THESIS SUBMITTED IN PARTIAL FULFILMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY in THE FACULTY OF GRADUATE STUDIES Department of Botany We accept this thesis as conforming to the required standard THE UNIVERSITY OF BRITISH COLUMBIA November 2003 © Juan Fernando Saldarriaga Echavarria, 2003 ABSTRACT New sequences of ribosomal and protein genes were combined with available morphological and paleontological data to produce a phylogenetic framework for dinoflagellates. The evolutionary history of some of the major morphological features of the group was then investigated in the light of that framework. Phylogenetic trees of dinoflagellates based on the small subunit ribosomal RNA gene (SSU) are generally poorly resolved but include many well- supported clades, and while combined analyses of SSU and LSU (large subunit ribosomal RNA) improve the support for several nodes, they are still generally unsatisfactory. Protein-gene based trees lack the degree of species representation necessary for meaningful in-group phylogenetic analyses, but do provide important insights to the phylogenetic position of dinoflagellates as a whole and on the identity of their close relatives. Molecular data agree with paleontology in suggesting an early evolutionary radiation of the group, but whereas paleontological data include only taxa with fossilizable cysts, the new data examined here establish that this radiation event included all dinokaryotic lineages, including athecate forms. Plastids were lost and replaced many times in dinoflagellates, a situation entirely unique for this group. Histones could well have been lost earlier in the lineage than previously assumed. -

In Situ Determination of Cellular DMSP and Pigment Quotas in a Prorocentrum Minimum Bloom Near the Falkland Islands Tyler Cyronaka, Erin O’Reilly B, Peter A
This article was downloaded by: [MUSC Library], [Peter Lee] On: 20 October 2014, At: 12:18 Publisher: Taylor & Francis Informa Ltd Registered in England and Wales Registered Number: 1072954 Registered office: Mortimer House, 37-41 Mortimer Street, London W1T 3JH, UK Advances in Oceanography and Limnology Publication details, including instructions for authors and subscription information: http://www.tandfonline.com/loi/taol20 In situ determination of cellular DMSP and pigment quotas in a Prorocentrum minimum bloom near the Falkland Islands Tyler Cyronaka, Erin O’Reilly b, Peter A. Leeb & Giacomo R. DiTulliob a Scripps Institution of Oceanography, UC San Diego, 9500 Gilman Drive, 0244, La Jolla, CA 92093-0244 b Grice and Hollings Marine Laboratories, College of Charleston, Charleston, South Carolina 29412, USA Published online: 16 Oct 2014. To cite this article: Tyler Cyronak, Erin O’Reilly, Peter A. Lee & Giacomo R. DiTullio (2014): In situ determination of cellular DMSP and pigment quotas in a Prorocentrum minimum bloom near the Falkland Islands, Advances in Oceanography and Limnology, DOI: 10.1080/19475721.2014.968620 To link to this article: http://dx.doi.org/10.1080/19475721.2014.968620 PLEASE SCROLL DOWN FOR ARTICLE Taylor & Francis makes every effort to ensure the accuracy of all the information (the “Content”) contained in the publications on our platform. However, Taylor & Francis, our agents, and our licensors make no representations or warranties whatsoever as to the accuracy, completeness, or suitability for any purpose of the Content. Any opinions and views expressed in this publication are the opinions and views of the authors, and are not the views of or endorsed by Taylor & Francis. -

The Planktonic Protist Interactome: Where Do We Stand After a Century of Research?
bioRxiv preprint doi: https://doi.org/10.1101/587352; this version posted May 2, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Bjorbækmo et al., 23.03.2019 – preprint copy - BioRxiv The planktonic protist interactome: where do we stand after a century of research? Marit F. Markussen Bjorbækmo1*, Andreas Evenstad1* and Line Lieblein Røsæg1*, Anders K. Krabberød1**, and Ramiro Logares2,1** 1 University of Oslo, Department of Biosciences, Section for Genetics and Evolutionary Biology (Evogene), Blindernv. 31, N- 0316 Oslo, Norway 2 Institut de Ciències del Mar (CSIC), Passeig Marítim de la Barceloneta, 37-49, ES-08003, Barcelona, Catalonia, Spain * The three authors contributed equally ** Corresponding authors: Ramiro Logares: Institute of Marine Sciences (ICM-CSIC), Passeig Marítim de la Barceloneta 37-49, 08003, Barcelona, Catalonia, Spain. Phone: 34-93-2309500; Fax: 34-93-2309555. [email protected] Anders K. Krabberød: University of Oslo, Department of Biosciences, Section for Genetics and Evolutionary Biology (Evogene), Blindernv. 31, N-0316 Oslo, Norway. Phone +47 22845986, Fax: +47 22854726. [email protected] Abstract Microbial interactions are crucial for Earth ecosystem function, yet our knowledge about them is limited and has so far mainly existed as scattered records. Here, we have surveyed the literature involving planktonic protist interactions and gathered the information in a manually curated Protist Interaction DAtabase (PIDA). In total, we have registered ~2,500 ecological interactions from ~500 publications, spanning the last 150 years. -

University of Oklahoma
UNIVERSITY OF OKLAHOMA GRADUATE COLLEGE MACRONUTRIENTS SHAPE MICROBIAL COMMUNITIES, GENE EXPRESSION AND PROTEIN EVOLUTION A DISSERTATION SUBMITTED TO THE GRADUATE FACULTY in partial fulfillment of the requirements for the Degree of DOCTOR OF PHILOSOPHY By JOSHUA THOMAS COOPER Norman, Oklahoma 2017 MACRONUTRIENTS SHAPE MICROBIAL COMMUNITIES, GENE EXPRESSION AND PROTEIN EVOLUTION A DISSERTATION APPROVED FOR THE DEPARTMENT OF MICROBIOLOGY AND PLANT BIOLOGY BY ______________________________ Dr. Boris Wawrik, Chair ______________________________ Dr. J. Phil Gibson ______________________________ Dr. Anne K. Dunn ______________________________ Dr. John Paul Masly ______________________________ Dr. K. David Hambright ii © Copyright by JOSHUA THOMAS COOPER 2017 All Rights Reserved. iii Acknowledgments I would like to thank my two advisors Dr. Boris Wawrik and Dr. J. Phil Gibson for helping me become a better scientist and better educator. I would also like to thank my committee members Dr. Anne K. Dunn, Dr. K. David Hambright, and Dr. J.P. Masly for providing valuable inputs that lead me to carefully consider my research questions. I would also like to thank Dr. J.P. Masly for the opportunity to coauthor a book chapter on the speciation of diatoms. It is still such a privilege that you believed in me and my crazy diatom ideas to form a concise chapter in addition to learn your style of writing has been a benefit to my professional development. I’m also thankful for my first undergraduate research mentor, Dr. Miriam Steinitz-Kannan, now retired from Northern Kentucky University, who was the first to show the amazing wonders of pond scum. Who knew that studying diatoms and algae as an undergraduate would lead me all the way to a Ph.D. -

Metagenomic Characterization of Unicellular Eukaryotes in the Urban Thessaloniki Bay
Metagenomic characterization of unicellular eukaryotes in the urban Thessaloniki Bay George Tsipas SCHOOL OF ECONOMICS, BUSINESS ADMINISTRATION & LEGAL STUDIES A thesis submitted for the degree of Master of Science (MSc) in Bioeconomy Law, Regulation and Management May, 2019 Thessaloniki – Greece George Tsipas ’’Metagenomic characterization of unicellular eukaryotes in the urban Thessaloniki Bay’’ Student Name: George Tsipas SID: 268186037282 Supervisor: Prof. Dr. Savvas Genitsaris I hereby declare that the work submitted is mine and that where I have made use of another’s work, I have attributed the source(s) according to the Regulations set in the Student’s Handbook. May, 2019 Thessaloniki - Greece Page 2 of 63 George Tsipas ’’Metagenomic characterization of unicellular eukaryotes in the urban Thessaloniki Bay’’ 1. Abstract The present research investigates through metagenomics sequencing the unicellular protistan communities in Thermaikos Gulf. This research analyzes the diversity, composition and abundance in this marine environment. Water samples were collected monthly from April 2017 to February 2018 in the port of Thessaloniki (Harbor site, 40o 37’ 55 N, 22o 56’ 09 E). The extraction of DNA was completed as well as the sequencing was performed, before the downstream read processing and the taxonomic classification that was assigned using PR2 database. A total of 1248 Operational Taxonomic Units (OTUs) were detected but only 700 unicellular eukaryotes were analyzed, excluding unclassified OTUs, Metazoa and Streptophyta. In this research-based study the most abundant and diverse taxonomic groups were Dinoflagellata and Protalveolata. Specifically, the most abundant groups of all samples are Dinoflagellata with 190 OTUs (27.70%), Protalveolata with 139 OTUs (20.26%) Ochrophyta with 73 OTUs (10.64%), Cercozoa with 67 OTUs (9.77%) and Ciliophora with 64 OTUs (9.33%). -

Predatory Flagellates – the New Recently Discovered Deep Branches of the Eukaryotic Tree and Their Evolutionary and Ecological Significance
Protistology 14 (1), 15–22 (2020) Protistology Predatory flagellates – the new recently discovered deep branches of the eukaryotic tree and their evolutionary and ecological significance Denis V. Tikhonenkov Papanin Institute for Biology of Inland Waters, Russian Academy of Sciences, Borok, 152742, Russia | Submitted March 20, 2020 | Accepted April 6, 2020 | Summary Predatory protists are poorly studied, although they are often representing important deep-branching evolutionary lineages and new eukaryotic supergroups. This short review/opinion paper is inspired by the recent discoveries of various predatory flagellates, which form sister groups of the giant eukaryotic clusters on phylogenetic trees, and illustrate an ancestral state of one or another supergroup of eukaryotes. Here we discuss their evolutionary and ecological relevance and show that the study of such protists may be essential in addressing previously puzzling evolutionary problems, such as the origin of multicellular animals, the plastid spread trajectory, origins of photosynthesis and parasitism, evolution of mitochondrial genomes. Key words: evolution of eukaryotes, heterotrophic flagellates, mitochondrial genome, origin of animals, photosynthesis, predatory protists, tree of life Predatory flagellates and diversity of eu- of the hidden diversity of protists (Moon-van der karyotes Staay et al., 2000; López-García et al., 2001; Edg- comb et al., 2002; Massana et al., 2004; Richards The well-studied multicellular animals, plants and Bass, 2005; Tarbe et al., 2011; de Vargas et al., and fungi immediately come to mind when we hear 2015). In particular, several prevailing and very abun- the term “eukaryotes”. However, these groups of dant ribogroups such as MALV, MAST, MAOP, organisms represent a minority in the real diversity MAFO (marine alveolates, stramenopiles, opistho- of evolutionary lineages of eukaryotes. -
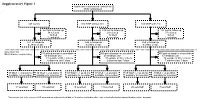
Supplementary Figure 1 Multicenter Randomised Control Trial 2746 Randomised
Supplementary Figure 1 Multicenter randomised control trial 2746 randomised 947 control 910 MNP without zinc 889 MNP with zinc 223 lost to follow up 219 lost to follow up 183 lost to follow up 34 refused 29 refused 37 refused 16 died 12 died 9 died 3 excluded 4 excluded 1 excluded 671 in follow-up 646 in follow-up 659 in follow-up at 24mo of age at 24mo of age at 24mo of age Selection for Microbiome sequencing 516 paired samples unavailable 469 paired samples unavailable 497 paired samples unavailable 69 antibiotic use 63 antibiotic use 67 antibiotic use 31 outside of WLZ criteria 37 outside of WLZ criteria 34 outside of WLZ criteria 6 diarrhea last 7 days 2 diarrhea last 7 days 7 diarrhea last 7 days 39 WLZ > -1 at 24 mo 10 WLZ < -2 at 24mo 58 WLZ > -1 at 24 mo 17 WLZ < -2 at 24mo 48 WLZ > -1 at 24 mo 8 WLZ < -2 at 24mo available for selection available for selection available for selection available for selection available for selection1 available for selection1 14 selected 10 selected 15 selected 14 selected 20 selected1 7 selected1 1 Two subjects (one in the reference WLZ group and one undernourished) had, at 12 months, no diarrhea within 1 day of stool collection but reported diarrhea within 7 days prior. Length, cm kg Weight, Supplementary Figure 2. Length (left) and weight (right) z-scores of children recruited into clinical trial NCT00705445 during the first 24 months of life. Median and quantile values are shown, with medians for participants profiled in current study indicated by red (undernourished) and black (reference WLZ) lines. -

The Mitochondrial Genome and Transcriptome of the Basal
View metadata, citation and similar papers at core.ac.uk brought to you by CORE GBEprovided by PubMed Central The Mitochondrial Genome and Transcriptome of the Basal Dinoflagellate Hematodinium sp.: Character Evolution within the Highly Derived Mitochondrial Genomes of Dinoflagellates C. J. Jackson, S. G. Gornik, and R. F. Waller* School of Botany, University of Melbourne, Australia *Corresponding author: E-mail: [email protected]. Accepted: 12 November 2011 Abstract The sister phyla dinoflagellates and apicomplexans inherited a drastically reduced mitochondrial genome (mitochondrial DNA, mtDNA) containing only three protein-coding (cob, cox1, and cox3) genes and two ribosomal RNA (rRNA) genes. In apicomplexans, single copies of these genes are encoded on the smallest known mtDNA chromosome (6 kb). In dinoflagellates, however, the genome has undergone further substantial modifications, including massive genome amplification and recombination resulting in multiple copies of each gene and gene fragments linked in numerous combinations. Furthermore, protein-encoding genes have lost standard stop codons, trans-splicing of messenger RNAs (mRNAs) is required to generate complete cox3 transcripts, and extensive RNA editing recodes most genes. From taxa investigated to date, it is unclear when many of these unusual dinoflagellate mtDNA characters evolved. To address this question, we investigated the mitochondrial genome and transcriptome character states of the deep branching dinoflagellate Hematodinium sp. Genomic data show that like later-branching dinoflagellates Hematodinium sp. also contains an inflated, heavily recombined genome of multicopy genes and gene fragments. Although stop codons are also lacking for cox1 and cob, cox3 still encodes a conventional stop codon. Extensive editing of mRNAs also occurs in Hematodinium sp. -

Insights Into Alexandrium Minutum Nutrient Acquisition, Metabolism and Saxitoxin Biosynthesis Through Comprehensive Transcriptome Survey
biology Article Insights into Alexandrium minutum Nutrient Acquisition, Metabolism and Saxitoxin Biosynthesis through Comprehensive Transcriptome Survey Muhamad Afiq Akbar 1, Nurul Yuziana Mohd Yusof 2 , Fathul Karim Sahrani 2, Gires Usup 2, Asmat Ahmad 1, Syarul Nataqain Baharum 3 , Nor Azlan Nor Muhammad 3 and Hamidun Bunawan 3,* 1 Department of Biological Sciences and Biotechnology, Faculty of Science and Technology, Universiti Kebangsaan Malaysia, Bangi 43600, Malaysia; muhdafi[email protected] (M.A.A.); [email protected] (A.A.) 2 Department of Earth Science and Environment, Faculty of Science and Technology, Universiti Kebangsaan Malaysia, Bangi 43600, Malaysia; [email protected] (N.Y.M.Y.); [email protected] (F.K.S.); [email protected] (G.U.) 3 Institute of System Biology, Universiti Kebangsaan Malaysia, Bangi 43600, Malaysia; [email protected] (S.N.B.); [email protected] (N.A.N.M.) * Correspondence: [email protected]; Tel.: +60-389-214-570 Simple Summary: Alexandrium minutum is one of the causing organisms for the occurrence of harmful algae bloom (HABs) in marine ecosystems. This species produces saxitoxin, one of the deadliest neurotoxins which can cause human mortality. However, molecular information such as genes and proteins catalog on this species is still lacking. Therefore, this study has successfully Citation: Akbar, M.A.; Yusof, N.Y.M.; characterized several new molecular mechanisms regarding A. minutum environmental adaptation Sahrani, F.K.; Usup, G.; Ahmad, A.; and saxitoxin biosynthesis. Ultimately, this study provides a valuable resource for facilitating future Baharum, S.N.; Muhammad, N.A.N.; dinoflagellates’ molecular response to environmental changes. -

Ellobiopsids of the Genus Thalassomyces Are Alveolates
J. Eukaryot. Microbiol., 51(2), 2004 pp. 246±252 q 2004 by the Society of Protozoologists Ellobiopsids of the Genus Thalassomyces are Alveolates JEFFREY D. SILBERMAN,a,b1 ALLEN G. COLLINS,c,2 LISA-ANN GERSHWIN,d,3 PATRICIA J. JOHNSONa and ANDREW J. ROGERe aDepartment of Microbiology, Immunology, and Molecular Genetics, University of California at Los Angeles, California, USA, and bInstitute of Geophysics and Planetary Physics, University of California at Los Angeles, California, USA, and cEcology, Behavior and Evolution Section, Division of Biology, University of California, La Jolla, California, USA, and dDepartment of Integrative Biology and Museum of Paleontology, University of California, Berkeley, California, USA, and eCanadian Institute for Advanced Research, Program in Evolutionary Biology, Genome Atlantic, Department of Biochemistry and Molecular Biology, Dalhousie University, Halifax, Nova Scotia, Canada ABSTRACT. Ellobiopsids are multinucleate protist parasites of aquatic crustaceans that possess a nutrient absorbing `root' inside the host and reproductive structures that protrude through the carapace. Ellobiopsids have variously been af®liated with fungi, `colorless algae', and dino¯agellates, although no morphological character has been identi®ed that de®nitively allies them with any particular eukaryotic lineage. The arrangement of the trailing and circumferential ¯agella of the rarely observed bi-¯agellated `zoospore' is reminiscent of dino¯agellate ¯agellation, but a well-organized `dinokaryotic nucleus' has never been observed. Using small subunit ribosomal RNA gene sequences from two species of Thalassomyces, phylogenetic analyses robustly place these ellobiopsid species among the alveolates (ciliates, apicomplexans, dino¯agellates and relatives) though without a clear af®liation to any established alveolate lineage. Our trees demonstrate that Thalassomyces fall within a dino¯agellate 1 apicomplexa 1 Perkinsidae 1 ``marine alveolate group 1'' clade, clustering most closely with dino¯agellates. -

Metabarcoding Analysis of Prey Composition of the Copepod Calanus Finmarchicus in Regions of the North Atlantic Ocean Heidi Yeh [email protected]
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by OpenCommons at University of Connecticut University of Connecticut OpenCommons@UConn Master's Theses University of Connecticut Graduate School 7-16-2018 Metabarcoding Analysis of Prey Composition of the Copepod Calanus finmarchicus in Regions of the North Atlantic Ocean Heidi Yeh [email protected] Recommended Citation Yeh, Heidi, "Metabarcoding Analysis of Prey Composition of the Copepod Calanus finmarchicus in Regions of the North Atlantic Ocean" (2018). Master's Theses. 1257. https://opencommons.uconn.edu/gs_theses/1257 This work is brought to you for free and open access by the University of Connecticut Graduate School at OpenCommons@UConn. It has been accepted for inclusion in Master's Theses by an authorized administrator of OpenCommons@UConn. For more information, please contact [email protected]. Metabarcoding Analysis of Prey Composition of the Copepod Calanus finmarchicus in Regions of the North Atlantic Ocean Heidi Yeh B.A., Barnard College, Columbia University, 2014 A Thesis Submitted in Partial Fulfillment of the Requirements for the Degree of Master of Science At the University of Connecticut 2018 Copyright by Heidi Yeh 2018 ii APPROVAL PAGE Masters of Science Thesis Metabarcoding Analysis of Prey Composition of the Copepod Calanus finmarchicus in Regions of the North Atlantic Ocean Presented by Heidi Yeh, B.A. Major Advisor________________________________________________________________ Ann Bucklin Associate Advisor_______________________________________________________________ Senjie Lin Associate Advisor_______________________________________________________________ George McManus University of Connecticut 2018 iii ACKNOWLEDGEMENTS Many people have provided support and encouragement over the course of this research project. I would like to thank my advisor, Ann Bucklin. -

Genetic Diversity and Habitats of Two Enigmatic Marine Alveolate Lineages
AQUATIC MICROBIAL ECOLOGY Vol. 42: 277–291, 2006 Published March 29 Aquat Microb Ecol Genetic diversity and habitats of two enigmatic marine alveolate lineages Agnès Groisillier1, Ramon Massana2, Klaus Valentin3, Daniel Vaulot1, Laure Guillou1,* 1Station Biologique, UMR 7144, CNRS, and Université Pierre & Marie Curie, BP74, 29682 Roscoff Cedex, France 2Department de Biologia Marina i Oceanografia, Institut de Ciències del Mar, CMIMA, CSIC. Passeig Marítim de la Barceloneta 37-49, 08003 Barcelona, Spain 3Alfred Wegener Institute for Polar Research, Am Handelshafen 12, 27570 Bremerhaven, Germany ABSTRACT: Systematic sequencing of environmental SSU rDNA genes amplified from different marine ecosystems has uncovered novel eukaryotic lineages, in particular within the alveolate and stramenopile radiations. The ecological and geographic distribution of 2 novel alveolate lineages (called Group I and II in previous papers) is inferred from the analysis of 62 different environmental clone libraries from freshwater and marine habitats. These 2 lineages have been, up to now, retrieved exclusively from marine ecosystems, including oceanic and coastal waters, sediments, hydrothermal vents, and perma- nent anoxic deep waters and usually represent the most abundant eukaryotic lineages in environmen- tal genetic libraries. While Group I is only composed of environmental sequences (118 clones), Group II contains, besides environmental sequences (158 clones), sequences from described genera (8) (Hema- todinium and Amoebophrya) that belong to the Syndiniales, an atypical order of dinoflagellates exclu- sively composed of marine parasites. This suggests that Group II could correspond to Syndiniales, al- though this should be confirmed in the future by examining the morphology of cells from Group II. Group II appears to be abundant in coastal and oceanic ecosystems, whereas permanent anoxic waters and hy- drothermal ecosystems are usually dominated by Group I.