SARS-Cov-2 and Mirna-Like Inhibition Power
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Table S1. Identified Proteins with Exclusive Expression in Cerebellum of Rats of Control, 10Mg F/L and 50Mg F/L Groups
Table S1. Identified proteins with exclusive expression in cerebellum of rats of control, 10mg F/L and 50mg F/L groups. Accession PLGS Protein Name Group IDa Score Q3TXS7 26S proteasome non-ATPase regulatory subunit 1 435 Control Q9CQX8 28S ribosomal protein S36_ mitochondrial 197 Control P52760 2-iminobutanoate/2-iminopropanoate deaminase 315 Control Q60597 2-oxoglutarate dehydrogenase_ mitochondrial 67 Control P24815 3 beta-hydroxysteroid dehydrogenase/Delta 5-->4-isomerase type 1 84 Control Q99L13 3-hydroxyisobutyrate dehydrogenase_ mitochondrial 114 Control P61922 4-aminobutyrate aminotransferase_ mitochondrial 470 Control P10852 4F2 cell-surface antigen heavy chain 220 Control Q8K010 5-oxoprolinase 197 Control P47955 60S acidic ribosomal protein P1 190 Control P70266 6-phosphofructo-2-kinase/fructose-2_6-bisphosphatase 1 113 Control Q8QZT1 Acetyl-CoA acetyltransferase_ mitochondrial 402 Control Q9R0Y5 Adenylate kinase isoenzyme 1 623 Control Q80TS3 Adhesion G protein-coupled receptor L3 59 Control B7ZCC9 Adhesion G-protein coupled receptor G4 139 Control Q6P5E6 ADP-ribosylation factor-binding protein GGA2 45 Control E9Q394 A-kinase anchor protein 13 60 Control Q80Y20 Alkylated DNA repair protein alkB homolog 8 111 Control P07758 Alpha-1-antitrypsin 1-1 78 Control P22599 Alpha-1-antitrypsin 1-2 78 Control Q00896 Alpha-1-antitrypsin 1-3 78 Control Q00897 Alpha-1-antitrypsin 1-4 78 Control P57780 Alpha-actinin-4 58 Control Q9QYC0 Alpha-adducin 270 Control Q9DB05 Alpha-soluble NSF attachment protein 156 Control Q6PAM1 Alpha-taxilin 161 -

Molecular Profiling of Innate Immune Response Mechanisms in Ventilator-Associated 2 Pneumonia
bioRxiv preprint doi: https://doi.org/10.1101/2020.01.08.899294; this version posted January 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Title: Molecular profiling of innate immune response mechanisms in ventilator-associated 2 pneumonia 3 Authors: Khyatiben V. Pathak1*, Marissa I. McGilvrey1*, Charles K. Hu3, Krystine Garcia- 4 Mansfield1, Karen Lewandoski2, Zahra Eftekhari4, Yate-Ching Yuan5, Frederic Zenhausern2,3,6, 5 Emmanuel Menashi3, Patrick Pirrotte1 6 *These authors contributed equally to this work 7 Affiliations: 8 1. Collaborative Center for Translational Mass Spectrometry, Translational Genomics Research 9 Institute, Phoenix, Arizona 85004 10 2. Translational Genomics Research Institute, Phoenix, Arizona 85004 11 3. HonorHealth Clinical Research Institute, Scottsdale, Arizona 85258 12 4. Applied AI and Data Science, City of Hope Medical Center, Duarte, California 91010 13 5. Center for Informatics, City of Hope Medical Center, Duarte, California 91010 14 6. Center for Applied NanoBioscience and Medicine, University of Arizona, Phoenix, Arizona 15 85004 16 Corresponding author: 17 Dr. Patrick Pirrotte 18 Collaborative Center for Translational Mass Spectrometry 19 Translational Genomics Research Institute 20 445 North 5th Street, Phoenix, AZ, 85004 21 Tel: 602-343-8454; 22 [[email protected]] 23 Running Title: Innate immune response in ventilator-associated pneumonia 24 Key words: ventilator-associated pneumonia; endotracheal aspirate; proteome, metabolome; 25 neutrophil degranulation 26 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.01.08.899294; this version posted January 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

Identifying Genetic Variants and Pathways Associated with Extreme Levels of Fetal Hemoglobin in Sickle Cell Disease in Tanzania
Nkya et al. BMC Medical Genetics (2020) 21:125 https://doi.org/10.1186/s12881-020-01059-1 RESEARCH ARTICLE Open Access Identifying genetic variants and pathways associated with extreme levels of fetal hemoglobin in sickle cell disease in Tanzania Siana Nkya1,2*, Liberata Mwita2, Josephine Mgaya2, Happiness Kumburu3, Marco van Zwetselaar3, Stephan Menzel4, Gaston Kuzamunu Mazandu5,6,7* , Raphael Sangeda2,8, Emile Chimusa5 and Julie Makani2 Abstract Background: Sickle cell disease (SCD) is a blood disorder caused by a point mutation on the beta globin gene resulting in the synthesis of abnormal hemoglobin. Fetal hemoglobin (HbF) reduces disease severity, but the levels vary from one individual to another. Most research has focused on common genetic variants which differ across populations and hence do not fully account for HbF variation. Methods: We investigated rare and common genetic variants that influence HbF levels in 14 SCD patients to elucidate variants and pathways in SCD patients with extreme HbF levels (≥7.7% for high HbF) and (≤2.5% for low HbF) in Tanzania. We performed targeted next generation sequencing (Illumina_Miseq) covering exonic and other significant fetal hemoglobin-associated loci, including BCL11A, MYB, HOXA9, HBB, HBG1, HBG2, CHD4, KLF1, MBD3, ZBTB7A and PGLYRP1. Results: Results revealed a range of genetic variants, including bi-allelic and multi-allelic SNPs, frameshift insertions and deletions, some of which have functional importance. Notably, there were significantly more deletions in individuals with high HbF levels (11% vs 0.9%). We identified frameshift deletions in individuals with high HbF levels and frameshift insertions in individuals with low HbF. CHD4 and MBD3 genes, interacting in the same sub-network, were identified to have a significant number of pathogenic or non-synonymous mutations in individuals with low HbF levels, suggesting an important role of epigenetic pathways in the regulation of HbF synthesis. -
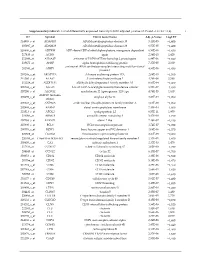
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

Research Article Association Between HBA Locus Copy Number Gains And
INTERNATIONAL JOURNAL OF MEDICAL BIOCHEMISTRY DOI: 10.14744/ijmb.2021.65477 Int J Med Biochem 2021;4(2):91-6 Research Article Association between HBA locus copy number gains and pathogenic HBB gene variants Guven Toksoy1, Nergis Akay2, Agharza Aghayev1, Volkan Karaman1, Sahin Avci1, Tugba Kalayci1, Umut Altunoglu1, Zeynep Karakas2, Zehra Oya Uyguner1 1Department of Medical Genetics, Istanbul University Istanbul Faculty of Medicine, Istanbul, Turkey 2Department of Pediatric Hematology-Oncology, Istanbul University Istanbul Faculty of Medicine, Istanbul, Turkey Abstract Objectives: Alpha (α) and beta (β) thalassemia are the most prevalent genetic hematological disorders. The co-occur- rence of silent β-thalassemia with excess α-globin gene copies is associated with the thalassemia intermedia pheno- type. This study was an investigation of the α-globulin gene dosage and sequence variations in thalassemia patients. Methods: Multiplex ligation-dependent probe amplification and Sanger sequencing were used to identify the hemo- globin subunit alpha 1 (HBA1) and HBA2 gene alterations in 32 patients. Deletion, duplication, and other findings were analyzed in the index cases and family members. Results: Four of the 32 cases (12.5%) were found to have gross duplications. Two cases demonstrated α-globin triplica- tion, and 2 had a quadruplicated HBA1/2 genes. Affected family members revealed genotype-phenotype correlation. In 1 patient, it was observed that quadruplicated HBA genes co-occurrence with hemoglobin subunit beta (HBB) mu- tation was inherited from his mother. Notably, the mother did not demonstrate any thalassemia phenotype. Further investigation showed that the mother was carrying a single copy HBA gene deletion in the trans allele that explained her clinical condition. -

Usbiological Datasheet
HBB (Hemoglobin Subunit beta, Beta-globin, Hemoglobin beta Chain, Cleaved into the following 2 Chains:LVV-hemorphin-7, Spinorphin) Catalog number 223153 Supplier United States Biological Hemoglobin (Hgb) iscoupled to four iron-binding,methene-linked tetrapyrrole rings(heme). The alpha (16p13.3; 5'-zeta-pseudozeta-pseudoalpha2-pseudoalpha1-alpha2-alpha1-θ1-3') and beta (11p15.4) globin loci determine the basic hemoglobin structure. The globin portion of Hgb consists of twoalphachains and two beta chains arranged in pairsforming a tetramer. Each of the four globin chainscovalently associates with a heme group. The bonds between alpha and beta chains are weaker than between similar globin chains, therebyforming a cleavage plane that is important for oxygen binding and release. High affinityfor oxygen occurs upon relaxation of thealpha1-beta2 cleavage plane. When the twoalpha1-beta2 interfaces are closely bound, hemoglobin has a lowaffinityfor oxygen. Hb A,which containstwo alpha chains plustwo beta chains,comprises 97% of totalcirculating hemoglobin. The remaining 3% of totalcirculating hemoglobin iscomprised of Hb A-2,which consists of twoalphachains plustwo delta chains, and fetal hemoglobin (Hb F),which consists of two alpha chainstogetherwith two y chains. Applications Suitable for use in Flow Cytometry, Western Blot, Immunohistochemistry. Other applications not tested. Recommended Dilution Western Blot: 1:500-1:2000 Immunohistochemistry: 1:50-1:200 Flow Cytometry: 1:50-1:200 Optimal dilutions to be determined by the researcher. Storage and Stability May be stored at 4°C for short-term only. Aliquot to avoid repeated freezing and thawing. Store at -20°C. Aliquots are stable for 12 months after receipt. For maximum recovery of product, centrifuge the original vial after thawing and prior to removing the cap. -

Centers for Medicare and Medicaid Services Clinical Laboratory Fee Schedule Annual Laboratory Public Meeting July 31, 2017
Centers for Medicare and Medicaid Services Clinical Laboratory Fee Schedule Annual Laboratory Public Meeting July 31, 2017 Anthony Sireci, MD, Msc Association for Molecular Pathology Expertise that advances patient care through education, innovation, and advocacy. www.amp.org Outline • Pharmacogenomics Procedures (81X30-81X36) • Human Platelet Antigen Procedures (81X15-81X22) • Heme-related Genetic Analysis Procedures, full sequence, common variants, familial variants and del/dup for F9, G6PD, and hemoglobin alpha and beta (81X25, 81X37- 81X40, 81257-81X69, and 813X1-813X4) • Hereditary Peripheral Neuropathy Genomic Sequencing Procedure (814X5) • Oncology-related Procedures • Common Variants (81X23-24) • Targeted Exon or Full Gene Procedures (81X04-05 and 813XX ) • Infectious Disease Procedures (876XX-87X6X and 02X1T) • Reconsidered Procedures (81327) Expertise that advances patient care through education, innovation, and advocacy. www.amp.org Annual Laboratory Public Meeting, July 31, 2017 Basis for Crosswalk Recommendations • Analysis of: • Analytical methods employed • Overall resources utilized • Types of genetic variants tested (e.g., SNPs, deletions, etc.,.) • Amount of genetic material interrogated Expertise that advances patient care through education, innovation, and advocacy. www.amp.org Annual Laboratory Public Meeting, July 31, 2017 Background: 81X30-36 Pharmacogenomics Codes New Code Descriptor 81X30 CYP3A4 (cytochrome P450 family 3 subfamily A member 4) (eg, drug metabolism), gene analysis, common variant(s) (eg, *2, *22) -

Identification of MAZ As a Novel Transcription Factor Regulating Erythropoiesis
Identification of MAZ as a novel transcription factor regulating erythropoiesis Darya Deen1, Falk Butter2, Michelle L. Holland3, Vasiliki Samara4, Jacqueline A. Sloane-Stanley4, Helena Ayyub4, Matthias Mann5, David Garrick4,6,7 and Douglas Vernimmen1,7 1 The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Easter Bush, Midlothian EH25 9RG, United Kingdom. 2 Institute of Molecular Biology (IMB), 55128 Mainz, Germany 3 Department of Medical and Molecular Genetics, School of Basic and Medical Biosciences, King's College London, London SE1 9RT, UK 4 MRC Molecular Haematology Unit, Weatherall Institute for Molecular Medicine, University of Oxford, John Radcliffe Hospital, Oxford OX3 9DS, United Kingdom. 5 Department of Proteomics and Signal Transduction, Max Planck Institute of Biochemistry, 82152 Martinsried, Germany. 6 Current address : INSERM UMRS-976, Institut de Recherche Saint Louis, Université de Paris, 75010 Paris, France. 7 These authors contributed equally Correspondence: [email protected], [email protected] ADDITIONAL MATERIAL Suppl Fig 1. Erythroid-specific hypersensitivity regions DNAse-Seq. (A) Structure of the α-globin locus. Chromosomal position (hg38 genome build) is shown above the genes. The locus consists of the embryonic ζ gene (HBZ), the duplicated foetal/adult α genes (HBA2 and HBA1) together with two flanking pseudogenes (HBM and HBQ1). The upstream, widely-expressed gene, NPRL3 is transcribed from the opposite strand to that of the HBA genes and contains the remote regulatory elements of the α-globin locus (MCS -R1 to -R3) (indicated by red dots). Below is shown the ATAC-seq signal in human erythroblasts (taken from 8) indicating open chromatin structure at promoter and distal regulatory elements (shaded regions). -

View Preprint
Accessing and applying molecular history Joshua G. Stern1 and Eric A. Gaucher2 1Machine Learning Research, Silver Spring, MD 20910; [email protected] 2School of Biology, School of Chemistry and Biochemistry, Georgia Institute of Technology, Atlanta, GA 30332; General Genomics, Atlanta, GA 30332 ABSTRACT Studying the evolutionary history of life’s molecules - DNA, RNA, and protein - reveals nature-based solutions to real-world problems. We discuss an approach to applied molecular evolution that is well-known within the field but may be unfamiliar to a wider audience. Using a case study at the intersection of molecular evolution and medicine, we introduce the fundamental concepts of orthology and paralogy. We also explain a practical entry point to molecular evolution named STORI: Selectable s Taxon Ortholog Retrieval Iteratively. STORI is a machine learning algorithm designed to clear a bottleneck t that researchers encounter when studying evolution. n i r Availability. Existing source code is available for download from GitHub (https://github. P com/jgstern/STORI_singlenode). e r Keywords: molecular evolution, machine learning, synthetic biology P INTRODUCTION In 2013, sickle-cell anemia killed more than 56,000 people [4]. This genetic disorder occurs when someone has a mutation in their beta-hemoglobin gene [6]. This gene is the DNA blueprint for actual beta-hemoglobin proteins: subcellular, nanometer-scale molecular machines made of yet smaller building blocks known as amino acids. Like a jeweler making necklaces from 20 different types of bead, life uses 20 different types of amino acid to build proteins. A single cell contains millions of proteins [38], although many of them share the same sequence of amino acids and thus have the same function. -

Novel Protein Pathways in Development and Progression of Pulmonary Sarcoidosis Maneesh Bhargava1*, K
www.nature.com/scientificreports OPEN Novel protein pathways in development and progression of pulmonary sarcoidosis Maneesh Bhargava1*, K. J. Viken1, B. Barkes2, T. J. Grifn3, M. Gillespie2, P. D. Jagtap3, R. Sajulga3, E. J. Peterson4, H. E. Dincer1, L. Li2, C. I. Restrepo2, B. P. O’Connor5, T. E. Fingerlin5, D. M. Perlman1 & L. A. Maier2 Pulmonary involvement occurs in up to 95% of sarcoidosis cases. In this pilot study, we examine lung compartment-specifc protein expression to identify pathways linked to development and progression of pulmonary sarcoidosis. We characterized bronchoalveolar lavage (BAL) cells and fuid (BALF) proteins in recently diagnosed sarcoidosis cases. We identifed 4,306 proteins in BAL cells, of which 272 proteins were diferentially expressed in sarcoidosis compared to controls. These proteins map to novel pathways such as integrin-linked kinase and IL-8 signaling and previously implicated pathways in sarcoidosis, including phagosome maturation, clathrin-mediated endocytic signaling and redox balance. In the BALF, the diferentially expressed proteins map to several pathways identifed in the BAL cells. The diferentially expressed BALF proteins also map to aryl hydrocarbon signaling, communication between innate and adaptive immune response, integrin, PTEN and phospholipase C signaling, serotonin and tryptophan metabolism, autophagy, and B cell receptor signaling. Additional pathways that were diferent between progressive and non-progressive sarcoidosis in the BALF included CD28 signaling and PFKFB4 signaling. Our studies demonstrate the power of contemporary proteomics to reveal novel mechanisms operational in sarcoidosis. Application of our workfows in well-phenotyped large cohorts maybe benefcial to identify biomarkers for diagnosis and prognosis and therapeutically tenable molecular mechanisms. -

The Long Non-Coding RNA Landscape in Juvenile Myelomonocytic Leukemia
LETTERS TO THE EDITOR in a variety of biological processes, human diseases and The long non-coding RNA landscape in juvenile malignancies.9 myelomonocytic leukemia To explore lncRNA expression in JMML, RNA from isolated mononuclear cell preparations from 44 previous- Juvenile myelomonocytic leukemia (JMML) is a rare ly untreated JMML patients (median age: 2.02 years) and and aggressive myelodysplastic and myeloproliferative 7 bone marrow samples from healthy controls (siblings disorder of early childhood. It is characterized by prolif- screened for transplantation; median age 6.83 years) was 1 eration of granulocytic and monocytic cells. Currently, profiled on a custom designed Agilent (8x60K) microar- hematopoietic stem cell transplantation (HSCT) is the ray platform described by Volders et al.8 The dataset standard of care and results in long-term overall survival (GEO Series accession n. GSE71449) had been previously 2 (OS) of only 50-60%. In approximately 90-95% of used in an mRNA expression analysis, leading to the patients, hyperactivation of the RAS/MAPK pathway can identification of overexpression of LIN28B in a fetal-like be observed through typical mutations in NRAS and subgroup of JMML patients.5 Details of patients' charac- KRAS (20-25%), PTPN11 (35%), NF1 (10-15%) or CBL teristics can be found in the Online Supplementary Table (10-15%) genes. This underlying molecular defect can in S1. part be used to guide patient management (e.g. no HSCT Only probes (n=23,728) representing known lncRNAs in patients with mutations in CBL).2 -

Hemoglobin Subunit Beta Gene Polymorphism Rs33949930 T>C and Risk of Sickle Cell Disease—A Case Control Study from Tabuk (Northwestern Part of Saudi Arabia)
International Journal of Clinical Medicine, 2016, 7, 25-31 Published Online January 2016 in SciRes. http://www.scirp.org/journal/ijcm http://dx.doi.org/10.4236/ijcm.2016.71004 Hemoglobin Subunit Beta Gene Polymorphism rs33949930 T>C and Risk of Sickle Cell Disease—A Case Control Study from Tabuk (Northwestern Part of Saudi Arabia) Rashid Mir1, Fawzia Sharaf2, Abu-Duhier FM1* 1Prince Fahd Bin Sultan Research Chair, Division of Cancer Molecular Genetics, Faculty of Applied Medical Sciences, University of Tabuk, Tabuk, Kingdom of Saudi Arabia 2King Khaled Hospital, Ministry of Health, Tabuk, Kingdom of Saudi Arabia Received 9 December 2015; accepted 19 January 2016; published 22 January 2016 Copyright © 2016 by authors and Scientific Research Publishing Inc. This work is licensed under the Creative Commons Attribution International License (CC BY). http://creativecommons.org/licenses/by/4.0/ Abstract Background: Sickle cell disease and sickle cell trait are common erythrocyte disorders that are most often caused by a point mutation (rs334, designated HbS) in the hemoglobin beta gene (HBB); however of this fact, there is extreme variability in occurrence and clinical presentation of sickle cell disease which may be explained by some other genetic changes associated with the gene. In the present study we examined the association between HBB gene polymorphism rs33949930 T>C in the occurrence of sickle cell disease in Saudi Arabia population. Materials and Methods: A case control study of 100 sickle cell disease patients and 100 healthy controls from Tabuk, Saudi Arabia. HBB gene rs33949930 T>C polymorphism was analyzed using Allele specific polymerase chain reac- tion technique.