Page 1 of 41 RSC Advances
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Macellibacteroides Fermentans Gen. Nov., Sp. Nov., a Member of the Family Porphyromonadaceae Isolated from an Upflow Anaerobic Filter Treating Abattoir Wastewaters
International Journal of Systematic and Evolutionary Microbiology (2012), 62, 2522–2527 DOI 10.1099/ijs.0.032508-0 Macellibacteroides fermentans gen. nov., sp. nov., a member of the family Porphyromonadaceae isolated from an upflow anaerobic filter treating abattoir wastewaters Linda Jabari,1,2 Hana Gannoun,2 Jean-Luc Cayol,1 Abdeljabbar Hedi,1 Mitsuo Sakamoto,3 Enevold Falsen,4 Moriya Ohkuma,3 Moktar Hamdi,2 Guy Fauque,1 Bernard Ollivier1 and Marie-Laure Fardeau1 Correspondence 1Aix-Marseille Universite´ du Sud Toulon-Var, CNRS/INSU, IRD, MIO, UM 110, Case 925, Marie-Laure Fardeau 163 Avenue de Luminy, 13288 Marseille Cedex 9, France [email protected] 2Laboratoire d’Ecologie et de Technologie Microbienne, Institut National des Sciences Applique´es et de Technologie, Centre Urbain Nord, BP 676, 1080 Tunis Cedex, Tunisia 3Microbe Division/Japan Collection of Microorganisms, RIKEN BioResource Center 2-1 Hirosawa, Wako, Saitama 351-0198, Japan 4CCUG, Culture Collection, Department of Clinical Bacteriology, University of Go¨teborg, 41346 Go¨teborg, Sweden A novel obligately anaerobic, non-spore-forming, rod-shaped mesophilic bacterium, which stained Gram-positive but showed the typical cell wall structure of Gram-negative bacteria, was isolated from an upflow anaerobic filter treating abattoir wastewaters in Tunisia. The strain, designated LIND7HT, grew at 20–45 6C (optimum 35–40 6C) and at pH 5.0–8.5 (optimum pH 6.5–7.5). It did not require NaCl for growth, but was able to grow in the presence of up to 2 % NaCl. Sulfate, thiosulfate, elemental sulfur, sulfite, nitrate and nitrite were not used as terminal electron acceptors. -

Sequencing Batch Reactor and Bacterial Community in Aerobic Granular Sludge for Wastewater Treatment of Noodle-Manufacturing Sector
applied sciences Article Sequencing Batch Reactor and Bacterial Community in Aerobic Granular Sludge for Wastewater Treatment of Noodle-Manufacturing Sector Tang Thi Chinh 1,3,*, Phung Duc Hieu 1, Bui Van Cuong 1, Nguyen Nhat Linh 2, Nguyen Ngoc Lan 2,3, Nguyen Sy Nguyen 1, Nguyen Quang Hung 1,3 and Le Thi Thu Hien 2,3,* ID 1 Institute of Environmental Technology (IET), Vietnam Academy of Science and Technology (VAST), 18 Hoang Quoc Viet, Cau Giay, Hanoi 100000, Vietnam; [email protected] (P.D.H.); [email protected] (B.V.C.); [email protected] (N.S.N.); [email protected] (N.Q.H.) 2 Institute of Genome Research (IGR), Vietnam Academy of Science and Technology (VAST), 18 Hoang Quoc Viet, Cau Giay, Hanoi 100000, Vietnam; [email protected] (N.N.L.); [email protected] (N.N.L.) 3 Graduate University of Science and Technology (GUST), Vietnam Academy of Science and Technology (VAST), 18 Hoang Quoc Viet, Cau Giay, Hanoi 100000, Vietnam * Correspondence: [email protected] (T.T.C.); [email protected] (L.T.T.H.); Tel.: +84-904-187-106 (T.T.C.); +84-989-019-691 (L.T.T.H.) Received: 1 March 2018; Accepted: 26 March 2018; Published: 27 March 2018 Abstract: The sequencing batch reactor (SBR) has been increasingly applied in the control of high organic wastewater. In this study, SBR with aerobic granular sludge was used for wastewater treatment in a noodle-manufacturing village in Vietnam. The results showed that after two months of operation, the chemical oxygen demand, total nitrogen and total phosphorous removal efficiency of aerobic granular SBR reached 92%, 83% and 75%, respectively. -

Revised Supplement 1: Reference List for Figure 1
Revised Supplement 1: Reference list for Figure 1. Manuscript title: Process disturbances in agricultural biogas production – causes, mechanisms and effects on the biogas microbiome: A review Susanne Theuerl 1,*, Johanna Klang 1, Annette Prochnow 1,2 1 Leibniz Institute for Agricultural Engineering and Bioeconomy, Max-Exth-Allee 100, 14469 Potsdam, Germany, [email protected] (ST), [email protected] (JK), [email protected] (AP) 2 Humboldt-Universität zu Berlin, Albrecht-Daniel-Thaer-Institute for Agricultural and Horticultural Sciences, Hinter der Reinhardtstr. 6-8, 10115 Berlin, Germany * Correspondence: [email protected] Tel.: +49-331-5699-900 References of Figure 1 1Abt et al. 2010, 2Parizzi et al. 2012, 3Hahnke et al. 2016 and Tomazetto et al. 2018, 4Ueki et al. 2006 and Gronow et al. 2011, 5Grabowski et al. 2005, 6Chen and Dong 2005, 7Avgustin et al. 1997 and Purushe et al. 2010, 8Yamada et al. 2006 and Matsuura et al. 2015, 9Yamada et al. 2007 and Matsuura et al. 2015, 10Sun et al. 2016, 11Suen et al. 2011, 12Hahnke et al. 2014 and Tomazetto et al. 2016, 13Mechichi et al. 1999, 14Koeck et al. 2015a and 2015b, 15Tomazetto et al. 2017, 16Fonknechten et al. 2010, 17Chen et al. 2010, 18Nishiyama et al. 2009, 19Sieber et al. 2010, 20Plerce et al. 2008, 21Westerholm et al. 2011 and Müller et al. 2015, 22Ueki et al. 2014, 23Jackson et al. 1999 and McInerney et al. 2007, 24Ma et al. 2017, 25Harmsen et al. 1998 and Plugge et al. 2012, 26Menes and Muxi 2002, Mavromatis et al. 2013 and Hania et al. -

Lascolabacillus Massiliensis'': a New Species Isolated
NEW MICROBES IN HUMANS “Lascolabacillus massiliensis”: a new consent, and the agreement of the National Ethics Committee of Senegal and the local ethics committee of the IFR48 species isolated from the human gut (Marseille, France) were obtained under numbers 11-017 and 09-022, respectively. The initial growth was obtained after 10 days of culture in a 5% sheep blood-enriched sheep rumen M. Beye, S. Bakour, S. I. Traore, D. Raoult and P.-E. Fournier medium in aerobic atmosphere at 37°C. The bacterium was Unité de Recherche en Maladies Infectieuses et Tropicales Emergentes, sub-cultured on 5% sheep blood-enriched Columbia agar Institut Hospitalo-Universitaire Méditerranée-Infection, Aix-Marseille (bioMérieux, Marcy l’Etoile, France) and grew in 24 hours at Université, Faculté de Médecine, Marseille cedex 5, France 37°C in both aerobic and anaerobic conditions. Agar-grown colonies were pale grey and 1.5 mm in diameter. Bacterial cells were Gram-negative, rod-shaped and polymorphic, ranginginlengthfrom1.5to10μm. Strain SIT8 was catalase- Abstract and oxidase-negative. The 16S rRNA gene was sequenced using the fD1-rP2 primers as previously described, using a 3130-XL sequencer (Applied Biosciences, Saint Aubin, We report here the main characteristics of “Lascolabacillus France). Strain SIT8 exhibited a 94.14% sequence identity with massiliensis” strain SIT8 (CSUR P1560) that was isolated from the Proteiniphilum acetatigenes strain TB107 (GenBank accession stool of a healthy 28-month-old boy. NR_043154), the phylogenetically closest species with New Microbes and New Infections © 2016 The Authors. Published standing in nomenclature (Fig. 1), which putatively classifies it by Elsevier Ltd on behalf of European Society of Clinical as a member of a new genus within the family Porphyr- Microbiology and Infectious Diseases. -

Phylogenomic Analysis of 589 Metagenome-Assembled Genomes Encompassing All Major Prokaryotic Lineages from the Gut of Higher Termites
Phylogenomic analysis of 589 metagenome-assembled genomes encompassing all major prokaryotic lineages from the gut of higher termites Vincent Hervé1, Pengfei Liu1, Carsten Dietrich1, David Sillam-Dussès2, Petr Stiblik3, Jan Šobotník3 and Andreas Brune1 1 Research Group Insect Gut Microbiology and Symbiosis, Max Planck Institute for Terrestrial Microbiology, Marburg, Germany 2 Laboratory of Experimental and Comparative Ethology EA 4443, Université Paris 13, Villetaneuse, France 3 Faculty of Forestry and Wood Sciences, Czech University of Life Sciences, Prague, Czech Republic ABSTRACT “Higher” termites have been able to colonize all tropical and subtropical regions because of their ability to digest lignocellulose with the aid of their prokaryotic gut microbiota. Over the last decade, numerous studies based on 16S rRNA gene amplicon libraries have largely described both the taxonomy and structure of the prokaryotic communities associated with termite guts. Host diet and microenvironmental conditions have emerged as the main factors structuring the microbial assemblages in the different gut compartments. Additionally, these molecular inventories have revealed the existence of termite-specific clusters that indicate coevolutionary processes in numerous prokaryotic lineages. However, for lack of representative isolates, the functional role of most lineages remains unclear. We reconstructed 589 metagenome-assembled genomes (MAGs) from the different Submitted 29 August 2019 gut compartments of eight higher termite species that encompass 17 prokaryotic -
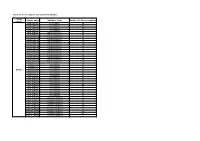
Table S1: List of Samples Included in the Analysis
Table S1: list of samples included in the analysis Study Sample name Inhibitory status Number of days at sampling number DNA.0P2T4 No inhibition 29 DNA.0P2T6 No inhibition 57 DNA.10P2T4 No inhibition 29 DNA.10P2T6 No inhibition 57 DNA.75P2T4 Phenol inhibition 29 DNA.75P2T6 Phenol inhibition 57 DNA.100P2T4 Phenol inhibition 29 DNA.100P2T6 Phenol inhibition 57 DNA.125P1T4 Phenol inhibition 29 DNA.125P1T6 Phenol inhibition 57 DNA.125P2T4 Phenol inhibition 29 DNA.125P2T6 Phenol inhibition 57 DNA.125P3T4 Phenol inhibition 29 DNA.125P3T6 Phenol inhibition 57 DNA.150P2T4 Phenol inhibition 29 DNA.150P2T6 Phenol inhibition 57 DNA.200P2T4 Phenol inhibition 29 DNA.200P2T6 Phenol inhibition 57 DNA.0N2T4 No inhibition 29 DNA.0N2T5 No inhibition 42 DNA.0N2T6 No inhibition 57 Study 1 DNA.5N2T4 No inhibition 29 DNA.5N2T5 No inhibition 42 DNA.5N2T6 No inhibition 57 DNA.10N2T4 No inhibition 29 DNA.10N2T5 No inhibition 42 DNA.10N2T6 No inhibition 57 DNA.15N2T4 No inhibition 29 DNA.15N2T5 No inhibition 42 DNA.15N2T6 No inhibition 57 DNA.25N2T4 No inhibition 29 DNA.25N2T5 No inhibition 42 DNA.25N2T6 No inhibition 57 DNA.75N2T4 Ammonia inhibition 29 DNA.75N2T5 Ammonia inhibition 42 DNA.75N2T6 Ammonia inhibition 57 DNA.100N2T4 Ammonia inhibition 29 DNA.100N2T5 Ammonia inhibition 42 DNA.100N2T6 Ammonia inhibition 57 DNA.250N2T4 Ammonia inhibition 29 DNA.250N2T5 Ammonia inhibition 42 DNA.250N2T6 Ammonia inhibition 57 nono2T3 No inhibition 16 noN2T4 Ammonia inhibition 23 noN2T8 Ammonia inhibition 60 noN2T9 Ammonia inhibition 85 noPhi2T4 Phenol inhibition 23 noPhi2T5 -

Electronic Supplementary Information
Electronic Supplementary Material (ESI) for Environmental Science: Water Research & Technology. This journal is © The Royal Society of Chemistry 2019 Electronic Supplementary Information Microbial community and antibiotic resistance profiles of biomass and effluent are distinctly affected by antibiotic addition to an anaerobic membrane bioreactor Ali Zarei-Baygi*, Moustapha Harb#,*, Phillip Wang*, Lauren Stadler^, and Adam L. Smith*† * Astani Department of Civil and Environmental Engineering, University of Southern California, 3620 South Vermont Avenue, Los Angeles, CA 90089, USA # Department of Civil and Environmental Engineering, Lebanese American University, 309 Bassil Building, Byblos, Lebanon ^ Department of Civil and Environmental Engineering, Rice University, 6100 Main Street, Houston, TX 77005, USA †Corresponding author (Adam L. Smith) Phone: +1 213.740.0473 Email: [email protected] Number of pages: 15 Number of figures: 6 Number of tables: 6 S1 Quantification of antibiotics by LC-MS For antibiotics quantification, 10 mL samples were collected for each sampling time point from the influent and effluent of the AnMBR. Both collected samples and standard solutions were filtered through 0.2 µm PTFE syringe filters (Whatman) using 10 mL syringes with Luer lock tips and stored in certified 2 mL amber LC vials (Agilent) at 4 ºC refrigerator for no more than 3 days prior to analysis. Stock solutions of sulfamethoxazole and erythromycin were prepared in HPLC-grade methanol at concentrations of 20 mg/L and stored at -20 ºC. Ampicillin stock solution was prepared in HPLC-grade water at 4 mg/L due to its lack of solubility in methanol and stored at 4 ºC. For each antibiotic, a six-point standard calibration curve was constructed within the appropriate range (i.e., 0.1-30 µg/L to target effluent antibiotics and 30- 400 µg/L to target influent antibiotics). -

EXPERIMENTAL STUDIES on FERMENTATIVE FIRMICUTES from ANOXIC ENVIRONMENTS: ISOLATION, EVOLUTION, and THEIR GEOCHEMICAL IMPACTS By
EXPERIMENTAL STUDIES ON FERMENTATIVE FIRMICUTES FROM ANOXIC ENVIRONMENTS: ISOLATION, EVOLUTION, AND THEIR GEOCHEMICAL IMPACTS By JESSICA KEE EUN CHOI A dissertation submitted to the School of Graduate Studies Rutgers, The State University of New Jersey In partial fulfillment of the requirements For the degree of Doctor of Philosophy Graduate Program in Microbial Biology Written under the direction of Nathan Yee And approved by _______________________________________________________ _______________________________________________________ _______________________________________________________ _______________________________________________________ New Brunswick, New Jersey October 2017 ABSTRACT OF THE DISSERTATION Experimental studies on fermentative Firmicutes from anoxic environments: isolation, evolution and their geochemical impacts by JESSICA KEE EUN CHOI Dissertation director: Nathan Yee Fermentative microorganisms from the bacterial phylum Firmicutes are quite ubiquitous in subsurface environments and play an important biogeochemical role. For instance, fermenters have the ability to take complex molecules and break them into simpler compounds that serve as growth substrates for other organisms. The research presented here focuses on two groups of fermentative Firmicutes, one from the genus Clostridium and the other from the class Negativicutes. Clostridium species are well-known fermenters. Laboratory studies done so far have also displayed the capability to reduce Fe(III), yet the mechanism of this activity has not been investigated -

Research Article Review Jmb
J. Microbiol. Biotechnol. (2017), 27(0), 1–7 https://doi.org/10.4014/jmb.1707.07027 Research Article Review jmb Methods 20,546 sequences and all the archaeal datasets were normalized to 21,154 sequences by the “sub.sample” Bioinformatics Analysis command. The filtered sequences were classified against The raw read1 and read2 datasets was demultiplexed by the SILVA 16S reference database (Release 119) using a trimming the barcode sequences with no more than 1 naïve Bayesian classifier built in Mothur with an 80% mismatch. Then the sequences with the same ID were confidence score [5]. Sequences passing through all the picked from the remaining read1 and read2 datasets by a filtration were also clustered into OTUs at 6% dissimilarity self-written python script. Bases with average quality score level. Then a “classify.otu” function was utilized to assign lower than 25 over a 25 bases sliding window were the phylogenetic information to each OTU. excluded and sequences which contained any ambiguous base or had a final length shorter than 200 bases were Reference abandoned using Sickle [1]. The paired reads were assembled into contigs and any contigs with an ambiguous 1. Joshi NA, FJ. 2011. Sickle: A sliding-window, adaptive, base, more than 8 homopolymeric bases and fewer than 10 quality-based trimming tool for FastQ files (Version 1.33) bp overlaps were culled. After that, the contigs were [Software]. further trimmed to get rid of the contigs that have more 2. Schloss PD. 2010. The Effects of Alignment Quality, than 1 forward primer mismatch and 2 reverse primer Distance Calculation Method, Sequence Filtering, and Region on the Analysis of 16S rRNA Gene-Based Studies. -

Paludibacter Propionicigenes Type Strain (WB4)
Lawrence Berkeley National Laboratory Recent Work Title Complete genome sequence of Paludibacter propionicigenes type strain (WB4). Permalink https://escholarship.org/uc/item/4kr5k5w1 Journal Standards in genomic sciences, 4(1) ISSN 1944-3277 Authors Gronow, Sabine Munk, Christine Lapidus, Alla et al. Publication Date 2011 DOI 10.4056/sigs.1503846 Peer reviewed eScholarship.org Powered by the California Digital Library University of California Standards in Genomic Sciences (2011) 4:36-44 DOI:10.4056/sigs.1503846 Complete genome sequence of Paludibacter propionicigenes type strain (WB4T) Sabine Gronow1, Christine Munk2,3, Alla Lapidus2, Matt Nolan2, Susan Lucas2, Nancy Hammon2, Shweta Deshpande2, Jan-Fang Cheng2, Roxane Tapia2,3, Cliff Han2,3, Lynne Goodwin2,3, Sam Pitluck2, Konstantinos Liolios2, Natalia Ivanova2, Konstantinos Mavromatis2, Natalia Mikhailova2, Amrita Pati2, Amy Chen4, Krishna Palaniappan4, Miriam Land2,5, Loren Hauser2,5, Yun-Juan Chang2,5, Cynthia D. Jeffries2,5, Evelyne Brambilla1, Manfred Rohde6, Markus Göker1, John C. Detter2,3, Tanja Woyke2, James Bristow2, Jonathan A. Eisen2,7, Victor Markowitz4, Philip Hugenholtz2,8, Nikos C. Kyrpides2, and Hans-Peter Klenk1* 1 DSMZ - German Collection of Microorganisms and Cell Cultures GmbH, Braunschweig, Germany 2 DOE Joint Genome Institute, Walnut Creek, California, USA 3 Los Alamos National Laboratory, Bioscience Division, Los Alamos, New Mexico, USA 4 Biological Data Management and Technology Center, Lawrence Berkeley National Laboratory, Berkeley, California, USA 5 Oak Ridge -

Characterization of Cucumber Fermentation Spoilage Bacteria by Enrichment Culture and 16S Rdna Cloning
Characterization of Cucumber Fermentation Spoilage Bacteria by Enrichment Culture and 16S rDNA Cloning Fred Breidt, Eduardo Medina, Doria Wafa, Ilenys P´erez-D´ıaz, Wendy Franco, Hsin-Yu Huang, Suzanne D. Johanningsmeier, and Jae Ho Kim Abstract: Commercial cucumber fermentations are typically carried out in 40000 L fermentation tanks. A secondary fermentation can occur after sugars are consumed that results in the formation of acetic, propionic, and butyric acids, concomitantly with the loss of lactic acid and an increase in pH. Spoilage fermentations can result in significant economic loss for industrial producers. The microbiota that result in spoilage remain incompletely defined. Previous studies have implicated yeasts, lactic acid bacteria, enterobacteriaceae, and Clostridia as having a role in spoilage fermentations. We report that Propionibacterium and Pectinatus isolates from cucumber fermentation spoilage converted lactic acid to propionic acid, increasing pH. The analysis of 16S rDNA cloning libraries confirmed and expanded the knowledge gained from previous studies using classical microbiological methods. Our data show that Gram-negative anaerobic bacteria supersede Gram-positive Fermincutes species after the pH rises from around 3.2 to pH 5, and propionic and butyric acids are produced. Characterization of the spoilage microbiota is an important first step in efforts to prevent cucumber fermentation spoilage. Keywords: pickled vegetables, Pectinatus, Propionibacteria, secondary cucumber fermentation, spoilage M: Food Microbiology Practical Application: An understanding of the microorganisms that cause commercial cucumber fermentation spoilage & Safety may aid in developing methods to prevent the spoilage from occurring. Introduction cucumbers fermented at 2.3% NaCl (Fleming and others 1989). Commercial cucumber fermentations are typically carried out In this fermentation tank, the initial lactic acid fermentation was in large 40000 L outdoor tanks (reviewed by Breidt and others completed within 2 wk, with 1.2% lactic acid formed (pH 3.6) 2007). -

Microbial Communities Associated with Stable Fly (Diptera: Muscidae) Larvae and Their Developmental Substrates Erin Scully USDA-ARS, [email protected]
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Faculty Publications: Department of Entomology Entomology, Department of 2017 Microbial Communities Associated With Stable Fly (Diptera: Muscidae) Larvae and Their Developmental Substrates Erin Scully USDA-ARS, [email protected] Kristina Friesen USDA-ARS, [email protected] Brian Wienhold USDA-ARS, [email protected] Lisa M. Durso USDA-ARS, [email protected] Follow this and additional works at: http://digitalcommons.unl.edu/entomologyfacpub Part of the Entomology Commons Scully, Erin; Friesen, Kristina; Wienhold, Brian; and Durso, Lisa M., "Microbial Communities Associated With Stable Fly (Diptera: Muscidae) Larvae and Their eD velopmental Substrates" (2017). Faculty Publications: Department of Entomology. 502. http://digitalcommons.unl.edu/entomologyfacpub/502 This Article is brought to you for free and open access by the Entomology, Department of at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in Faculty Publications: Department of Entomology by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln. Annals of the Entomological Society of America, 110(1), 2017, 61–72 doi: 10.1093/aesa/saw087 Special Collection: Filth Fly–Microbe Interactions Research article Microbial Communities Associated With Stable Fly (Diptera: Muscidae) Larvae and Their Developmental Substrates Erin Scully,1 Kristina Friesen,2,3 Brian Wienhold,2 and Lisa M. Durso2 1USDA, ARS, Stored Product