Discrete Roles of STAT4 and STAT6 Transcription Factors in Tuning Epigenetic Modifications and Transcription During T Helper Cell Differentiation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Stat4 Consensus and Mutant Oligonucleotides
SANTA CRUZ BIOTECHNOLOGY, INC. Stat4 Consensus and Mutant Oligonucleotides BACKGROUND PRODUCT Electrophoretic mobility shift assays (EMSAs), also known as gel shift assays, Stat4 CONSENSUS OLIGONUCLEOTIDE: sc-2569 provide a relatively straightforward and sensitive method for studying bind- ■ binding site for Stat4 (3) ing interactions between transcription factors and consensus DNA binding elements. For such studies, DNA probes are provided as double-stranded 5’ — GAG CCT GA T TTC CCC GAA AT G ATG AGC TAG — 3’ oligonucleotides designed with 5' OH blunt ends to facilitate labeling to high 3’ — CTC GGA C T A AAG GGG CTT TA C TAC TCG ATC — 5’ specific activity with polynucleotide kinase. These are constructed both with Stat4 MUTANT OLIGONUCLEOTIDE: sc-2570 specific DNA binding consensus sequences for various transcription factors ■ identical to sc-2569 with the exception of a “CCC” “TTT” substitution in and as control or “mutant” probes in which one or more nucleotides mapping → the Stat4 binding motif (3) within the consensus binding site has been substituted. 5’ — GAG CCT GA T TTC TTT GAA AT G ATG AGC TAG — 3’ REFERENCES 3’ — CTC GGA C T A AAG AAA CTT TA C TAC TCG ATC — 5’ 1. Dignam, J.D., et al. 1983. Accurate transcription initiation by RNA poly- merase II in a soluble extract from isolated mammalian nuclei. Nucleic SELECT PRODUCT CITATIONS Acids Res. 11: 1475-1489. 1. Ahn, H.J., et al. 1998. Requirement for distinct Janus kinases and STAT 2. Murre, C., et al. 1991. B cell- and myocyte-specific E2-box-binding factors proteins in T cell proliferation versus IFN-g production following IL-12 contain E12/E47-like subunits. -

RNA-Binding Protein Hnrnpll Regulates Mrna Splicing and Stability During B-Cell to Plasma-Cell Differentiation
RNA-binding protein hnRNPLL regulates mRNA splicing and stability during B-cell to plasma-cell differentiation Xing Changa,b, Bin Lic, and Anjana Raoa,b,d,e,1 Divisions of aSignaling and Gene Expression and cVaccine Discovery, La Jolla Institute for Allergy and Immunology, La Jolla, CA 92037; bSanford Consortium for Regenerative Medicine, La Jolla, CA 92037; and dDepartment of Pharmacology and eMoores Cancer Center, University of California at San Diego, La Jolla, CA 92093 Contributed by Anjana Rao, December 2, 2014 (sent for review July 20, 2014) Posttranscriptional regulation is a major mechanism to rewire the RBP-binding sites, thus validating the specificity of RBP binding transcriptomes during differentiation. Heterogeneous nuclear to coprecipitating RNAs and mapping RBP-binding sites on the RNA-binding protein LL (hnRNPLL) is specifically induced in terminally validated RNAs at close to single-nucleotide resolution (8). differentiated lymphocytes, including effector T cells and plasma Heterogeneous nuclear RNA-binding proteins (hnRNPs) is cells. To study the molecular functions of hnRNPLL at a genome- the term applied to a collection of unrelated nuclear RBPs. wide level, we identified hnRNPLL RNA targets and binding sites in hnRNPLL was identified through a targeted lentiviral shRNA plasma cells through integrated Photoactivatable-Ribonucleoside- screen for regulators of CD45RA to CD45RO switching during Enhanced Cross-Linking and Immunoprecipitation (PAR-CLIP) and memory T-cell development (9) and independently through RNA sequencing. hnRNPLL preferentially recognizes CA dinucleo- two separate screens performed by different groups for exclusion tide-containing sequences in introns and 3′ untranslated regions of CD45 exon 4 in a minigene context (10) and for altered CD44 (UTRs), promotes exon inclusion or exclusion in a context-dependent and CD45R expression on T cells in N-ethyl-N-nitrosourea manner, and stabilizes mRNA when associated with 3′ UTRs. -

The E–Id Protein Axis Modulates the Activities of the PI3K–AKT–Mtorc1
Downloaded from genesdev.cshlp.org on October 6, 2021 - Published by Cold Spring Harbor Laboratory Press The E–Id protein axis modulates the activities of the PI3K–AKT–mTORC1– Hif1a and c-myc/p19Arf pathways to suppress innate variant TFH cell development, thymocyte expansion, and lymphomagenesis Masaki Miyazaki,1,8 Kazuko Miyazaki,1,8 Shuwen Chen,1 Vivek Chandra,1 Keisuke Wagatsuma,2 Yasutoshi Agata,2 Hans-Reimer Rodewald,3 Rintaro Saito,4 Aaron N. Chang,5 Nissi Varki,6 Hiroshi Kawamoto,7 and Cornelis Murre1 1Department of Molecular Biology, University of California at San Diego, La Jolla, California 92093, USA; 2Department of Biochemistry and Molecular Biology, Shiga University of Medical School, Shiga 520-2192, Japan; 3Division of Cellular Immunology, German Cancer Research Center, D-69120 Heidelberg, Germany; 4Department of Medicine, University of California at San Diego, La Jolla, California 92093, USA; 5Center for Computational Biology, Institute for Genomic Medicine, University of California at San Diego, La Jolla, California 92093, USA; 6Department of Pathology, University of California at San Diego, La Jolla, California 92093, USA; 7Department of Immunology, Institute for Frontier Medical Sciences, Kyoto University, Kyoto 606-8507, Japan It is now well established that the E and Id protein axis regulates multiple steps in lymphocyte development. However, it remains unknown how E and Id proteins mechanistically enforce and maintain the naı¨ve T-cell fate. Here we show that Id2 and Id3 suppressed the development and expansion of innate variant follicular helper T (TFH) cells. Innate variant TFH cells required major histocompatibility complex (MHC) class I-like signaling and were associated with germinal center B cells. -

Targeting Il13ralpha2 Activates STAT6-TP63 Pathway to Suppress Breast Cancer Lung Metastasis Panagiotis Papageorgis1,2,3*, Sait Ozturk2, Arthur W
Papageorgis et al. Breast Cancer Research (2015) 17:98 DOI 10.1186/s13058-015-0607-y RESEARCH ARTICLE Open Access Targeting IL13Ralpha2 activates STAT6-TP63 pathway to suppress breast cancer lung metastasis Panagiotis Papageorgis1,2,3*, Sait Ozturk2, Arthur W. Lambert2, Christiana M. Neophytou1, Alexandros Tzatsos4, Chen K. Wong2, Sam Thiagalingam2*† and Andreas I. Constantinou1*† Abstract Introduction: Basal-like breast cancer (BLBC) is an aggressive subtype often characterized by distant metastasis, poor patient prognosis, and limited treatment options. Therefore, the discovery of alternative targets to restrain its metastatic potential is urgently needed. In this study, we aimed to identify novel genes that drive metastasis of BLBC and to elucidate the underlying mechanisms of action. Methods: An unbiased approach using gene expression profiling of a BLBC progression model and in silico leveraging of pre-existing tumor transcriptomes were used to uncover metastasis-promoting genes. Lentiviral- mediated knockdown of interleukin-13 receptor alpha 2 (IL13Ralpha2) coupled with whole-body in vivo bioluminescence imaging was performed to assess its role in regulating breast cancer tumor growth and lung metastasis. Gene expression microarray analysis was followed by in vitro validation and cell migration assays to elucidate the downstream molecular pathways involved in this process. Results: We found that overexpression of the decoy receptor IL13Ralpha2 is significantly enriched in basal compared with luminal primary breast tumors as well as in a subset of metastatic basal-B breast cancer cells. Importantly, breast cancer patients with high-grade tumors and increased IL13Ralpha2 levels had significantly worse prognosis for metastasis-free survival compared with patients with low expression. -

Entinostat Augments NK Cell Functions Via Epigenetic Upregulation of IFIT1-STING-STAT4 Pathway
www.oncotarget.com Oncotarget, 2020, Vol. 11, (No. 20), pp: 1799-1815 Research Paper Entinostat augments NK cell functions via epigenetic upregulation of IFIT1-STING-STAT4 pathway John M. Idso1, Shunhua Lao1, Nathan J. Schloemer1,2, Jeffrey Knipstein2, Robert Burns3, Monica S. Thakar1,2,* and Subramaniam Malarkannan1,2,4,5,* 1Laboratory of Molecular Immunology and Immunotherapy, Blood Research Institute, Versiti, Milwaukee, WI, USA 2Division of Pediatric Hematology-Oncology-BMT, Department of Pediatrics, Medical College of Wisconsin, Milwaukee, WI, USA 3Bioinformatics Core, Blood Research Institute, Versiti, Milwaukee, WI, USA 4Divson of Hematology-Oncology, Department of Medicine, Medical College of Wisconsin, Milwaukee, WI, USA 5Department of Microbiology and Immunology, Medical College of Wisconsin, Milwaukee, WI, USA *Co-senior authors Correspondence to: Monica S. Thakar, email: [email protected] Subramaniam Malarkannan, email: [email protected] Keywords: NK cells; histone deacetylase inhibitor; Ewing sarcoma; rhabdomyosarcoma; immunotherapy Received: September 10, 2019 Accepted: March 03, 2020 Published: May 19, 2020 Copyright: Idso et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Histone deacetylase inhibitors (HDACi) are an emerging cancer therapy; however, their effect on natural killer (NK) cell-mediated anti-tumor responses remain unknown. Here, we evaluated the impact of a benzamide HDACi, entinostat, on human primary NK cells as well as tumor cell lines. Entinostat significantly upregulated the expression of NKG2D, an essential NK cell activating receptor. Independently, entinostat augmented the expression of ULBP1, HLA, and MICA/B on both rhabdomyosarcoma and Ewing sarcoma cell lines. -

Detection of Enhancer-Associated Rearrangements Reveals Mechanisms of Oncogene Dysregulation in B-Cell Lymphoma
Published OnlineFirst July 30, 2015; DOI: 10.1158/2159-8290.CD-15-0370 ReseaRch aRticle Detection of Enhancer-Associated Rearrangements Reveals Mechanisms of Oncogene Dysregulation in B-cell Lymphoma Russell J.H. Ryan1,2, Yotam Drier1,2, Holly Whitton2, M. Joel Cotton1,2, Jasleen Kaur1,2, Robbyn Issner2, Shawn Gillespie1,2, Charles B. Epstein2, Valentina Nardi1, Aliyah R. Sohani1, Ephraim P. Hochberg3, and Bradley E. Bernstein1,2 abstRact B-cell lymphomas frequently contain genomic rearrangements that lead to onco- gene activation by heterologous distal regulatory elements. We used a novel approach called “pinpointing enhancer-associated rearrangements by chromatin immunoprecipitation,” or PEAR- ChIP, to simultaneously map enhancer activity and proximal rearrangements in lymphoma cell lines and patient biopsies. This method detects rearrangements involving known cancer genes, including CCND1, BCL2, MYC, PDCD1LG2, NOTCH1, CIITA, and SGK1, as well as novel enhancer duplication events of likely oncogenic significance. We identify lymphoma subtype–specific enhancers in the MYC locus that are silenced in lymphomas with MYC-activating rearrangements and are associated with germline polymorphisms that alter lymphoma risk. We show that BCL6-locus enhancers are acetylated by the BCL6-activating transcription factor MEF2B, and can undergo genomic duplication, or target the MYC promoter for activation in the context of a “pseudo-double-hit” t(3;8)(q27;q24) rearrangement linking the BCL6 and MYC loci. Our work provides novel insights regarding enhancer-driven oncogene activa- tion in lymphoma. SIGNIFICANCE: We demonstrate a novel approach for simultaneous detection of genomic rearrange- ments and enhancer activity in tumor biopsies. We identify novel mechanisms of enhancer-driven regu- lation of the oncogenes MYC and BCL6, and show that the BCL6 locus can serve as an enhancer donor in an “enhancer hijacking” translocation. -

IL-17-Secreting Th Cells Stat3 and Stat4 Direct Development Of
Stat3 and Stat4 Direct Development of IL-17-Secreting Th Cells Anubhav N. Mathur, Hua-Chen Chang, Dimitrios G. Zisoulis, Gretta L. Stritesky, Qing Yu, John T. O'Malley, This information is current as Reuben Kapur, David E. Levy, Geoffrey S. Kansas and Mark of September 29, 2021. H. Kaplan J Immunol 2007; 178:4901-4907; ; doi: 10.4049/jimmunol.178.8.4901 http://www.jimmunol.org/content/178/8/4901 Downloaded from References This article cites 38 articles, 15 of which you can access for free at: http://www.jimmunol.org/content/178/8/4901.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 29, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2007 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Stat3 and Stat4 Direct Development of IL-17-Secreting Th Cells1 Anubhav N. Mathur,2*† Hua-Chen Chang,2*† Dimitrios G. Zisoulis,‡ Gretta L. -

A Dissertation Entitled the Androgen Receptor
A Dissertation entitled The Androgen Receptor as a Transcriptional Co-activator: Implications in the Growth and Progression of Prostate Cancer By Mesfin Gonit Submitted to the Graduate Faculty as partial fulfillment of the requirements for the PhD Degree in Biomedical science Dr. Manohar Ratnam, Committee Chair Dr. Lirim Shemshedini, Committee Member Dr. Robert Trumbly, Committee Member Dr. Edwin Sanchez, Committee Member Dr. Beata Lecka -Czernik, Committee Member Dr. Patricia R. Komuniecki, Dean College of Graduate Studies The University of Toledo August 2011 Copyright 2011, Mesfin Gonit This document is copyrighted material. Under copyright law, no parts of this document may be reproduced without the expressed permission of the author. An Abstract of The Androgen Receptor as a Transcriptional Co-activator: Implications in the Growth and Progression of Prostate Cancer By Mesfin Gonit As partial fulfillment of the requirements for the PhD Degree in Biomedical science The University of Toledo August 2011 Prostate cancer depends on the androgen receptor (AR) for growth and survival even in the absence of androgen. In the classical models of gene activation by AR, ligand activated AR signals through binding to the androgen response elements (AREs) in the target gene promoter/enhancer. In the present study the role of AREs in the androgen- independent transcriptional signaling was investigated using LP50 cells, derived from parental LNCaP cells through extended passage in vitro. LP50 cells reflected the signature gene overexpression profile of advanced clinical prostate tumors. The growth of LP50 cells was profoundly dependent on nuclear localized AR but was independent of androgen. Nevertheless, in these cells AR was unable to bind to AREs in the absence of androgen. -

Interferons Inhibit Activation of STAT6 by Interleukin 4 in Human Monocytes by Inducing SOCS-1 Gene Expression
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 10800–10805, September 1999 Immunology Interferons inhibit activation of STAT6 by interleukin 4 in human monocytes by inducing SOCS-1 gene expression HAROLD L. DICKENSHEETS*, CHANDRASEKAR VENKATARAMAN†,ULRIKE SCHINDLER†, AND RAYMOND P. DONNELLY*‡ *Division of Cytokine Biology, Center for Biologics Evaluation and Research, Food and Drug Administration, Bethesda, MD 20892; and †Tularik, Inc., South San Francisco, CA 94080 Edited by William E. Paul, National Institutes of Health, Bethesda, MD, and approved July 8, 1999 (received for review February 16, 1999) ABSTRACT Interferons (IFNs) inhibit induction by IL-4 12), 15-lipoxygenase (15-LO) (13, 14), IL-1 receptor antago- of multiple genes in human monocytes. However, the mecha- nist (IL-1ra) (15–17), and types I and II IL-1 receptors (IL-1R) nism by which IFNs mediate this inhibition has not been (18, 19). Type I IFNs (IFN-␣ and IFN-) and type II IFN defined. IL-4 activates gene expression by inducing tyrosine (IFN-␥) inhibit IL-4͞IL-13-induced gene expression in mono- phosphorylation, homodimerization, and nuclear transloca- cytes and B cells. For example, IFN-␥ suppresses IgE synthesis tion of the latent transcription factor, STAT6 (signal trans- by IL-4-stimulated B cells (20–22). IFNs also inhibit IL-4- ducer and activator of transcription-6). STAT6-responsive induced CD23 expression in both B cells (23, 24) and mono- elements are characteristically present in the promoters of cytes (25, 26). Furthermore, IFN-␥ inhibits IL-4- and IL-13- IL-4-inducible genes. Because STAT6 activation is essential induced expression of 15-LO and IL-1ra in monocytes (13, 14, for IL-4-induced gene expression, we examined the ability of 27). -

Whole-Genome Cartography of Estrogen Receptor a Binding Sites
Whole-Genome Cartography of Estrogen Receptor a Binding Sites Chin-Yo Lin1[¤, Vinsensius B. Vega1[, Jane S. Thomsen1, Tao Zhang1, Say Li Kong1, Min Xie1, Kuo Ping Chiu1, Leonard Lipovich1, Daniel H. Barnett2, Fabio Stossi2, Ailing Yeo3, Joshy George1, Vladimir A. Kuznetsov1, Yew Kok Lee1, Tze Howe Charn1, Nallasivam Palanisamy1, Lance D. Miller1, Edwin Cheung1,3, Benita S. Katzenellenbogen2, Yijun Ruan1, Guillaume Bourque1, Chia-Lin Wei1, Edison T. Liu1* 1 Genome Institute of Singapore, Singapore, Republic of Singapore, 2 Department of Molecular and Integrative Physiology, University of Illinois at Urbana-Champaign, Urbana, Illinois, United States of America, 3 Department of Biochemistry, Yong Loo Lin School of Medicine, National University of Singapore, Singapore, Republic of Singapore Using a chromatin immunoprecipitation-paired end diTag cloning and sequencing strategy, we mapped estrogen receptor a (ERa) binding sites in MCF-7 breast cancer cells. We identified 1,234 high confidence binding clusters of which 94% are projected to be bona fide ERa binding regions. Only 5% of the mapped estrogen receptor binding sites are located within 5 kb upstream of the transcriptional start sites of adjacent genes, regions containing the proximal promoters, whereas vast majority of the sites are mapped to intronic or distal locations (.5 kb from 59 and 39 ends of adjacent transcript), suggesting transcriptional regulatory mechanisms over significant physical distances. Of all the identified sites, 71% harbored putative full estrogen response elements (EREs), 25% bore ERE half sites, and only 4% had no recognizable ERE sequences. Genes in the vicinity of ERa binding sites were enriched for regulation by estradiol in MCF-7 cells, and their expression profiles in patient samples segregate ERa-positive from ERa-negative breast tumors. -
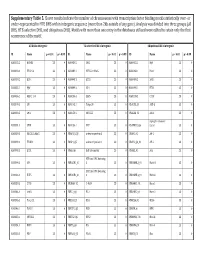
Supplementary Table 5. Clover Results Indicate the Number Of
Supplementary Table 5. Clover results indicate the number of chromosomes with transcription factor binding motifs statistically over‐ or under‐represented in HTE DHS within intergenic sequence (more than 2kb outside of any gene). Analysis was divided into three groups (all DHS, HTE‐selective DHS, and ubiquitous DHS). Motifs with more than one entry in the databases utilized were edited to retain only the first occurrence of the motif. All DHS x Intergenic TEselective DHS x Intergenic Ubiquitous DHS x Intergenic ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 MA0002.2 RUNX1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0003.1 TFAP2A 23 0 MA0089.1 NFE2L1::MafG 23 0 MA0068.1 Pax4 23 0 MA0039.2 Klf4 23 0 MA0098.1 ETS1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0099.2 AP1 23 0 MA0098.1 ETS1 23 0 MA0056.1 MZF1_1‐4 23 0 MA0136.1 ELF5 23 0 MA0139.1 CTCF 23 0 MA0079.2 SP1 23 0 MA0145.1 Tcfcp2l1 23 0 V$ALX3_01 ALX‐3 23 0 MA0080.2 SPI1 23 0 MA0150.1 NFE2L2 23 0 V$ALX4_02 Alx‐4 23 0 myocyte enhancer MA0081.1 SPIB 23 0 MA0156.1 FEV 23 0 V$AMEF2_Q6 factor 23 0 MA0089.1 NFE2L1::MafG 23 0 V$AP1FJ_Q2 activator protein 1 23 0 V$AP1_01 AP‐1 23 0 MA0090.1 TEAD1 23 0 V$AP4_Q5 activator protein 4 23 0 V$AP2_Q6_01 AP‐2 23 0 MA0098.1 ETS1 23 0 V$AR_Q6 half‐site matrix 23 0 V$ARX_01 Arx 23 0 BTB and CNC homolog MA0099.2 AP1 23 0 V$BACH1_01 1 23 0 V$BARHL1_01 Barhl‐1 23 0 BTB and CNC homolog MA0136.1 ELF5 23 0 V$BACH2_01 2 23 0 V$BARHL2_01 Barhl2 23 0 MA0139.1 CTCF 23 0 V$CMAF_02 C‐MAF 23 0 V$BARX1_01 Barx1 23 0 MA0144.1 Stat3 23 0 -

Coexpression and Relative Abundance Defined by B Cell
Stages of Germinal Center Transit Are Defined by B Cell Transcription Factor Coexpression and Relative Abundance This information is current as Giorgio Cattoretti, Rita Shaknovich, Paula M. Smith, of September 27, 2021. Hans-Martin Jäck, Vundavalli V. Murty and Bachir Alobeid J Immunol 2006; 177:6930-6939; ; doi: 10.4049/jimmunol.177.10.6930 http://www.jimmunol.org/content/177/10/6930 Downloaded from References This article cites 63 articles, 28 of which you can access for free at: http://www.jimmunol.org/content/177/10/6930.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 27, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2006 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Stages of Germinal Center Transit Are Defined by B Cell Transcription Factor Coexpression and Relative Abundance Giorgio Cattoretti,1*† Rita Shaknovich,‡ Paula M. Smith,† Hans-Martin Ja¨ck,§ Vundavalli V. Murty,*† and Bachir Alobeid* The transit of T cell-activated B cells through the germinal center (GC) is controlled by sequential activation and repression of key transcription factors, executing the pre- and post-GC B cell program.