Differences in Gene Expression and Variable Splicing Events of Ovaries
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

ACAT) in Cholesterol Metabolism: from Its Discovery to Clinical Trials and the Genomics Era
H OH metabolites OH Review Acyl-Coenzyme A: Cholesterol Acyltransferase (ACAT) in Cholesterol Metabolism: From Its Discovery to Clinical Trials and the Genomics Era Qimin Hai and Jonathan D. Smith * Department of Cardiovascular & Metabolic Sciences, Cleveland Clinic, Cleveland, OH 44195, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-216-444-2248 Abstract: The purification and cloning of the acyl-coenzyme A: cholesterol acyltransferase (ACAT) enzymes and the sterol O-acyltransferase (SOAT) genes has opened new areas of interest in cholesterol metabolism given their profound effects on foam cell biology and intestinal lipid absorption. The generation of mouse models deficient in Soat1 or Soat2 confirmed the importance of their gene products on cholesterol esterification and lipoprotein physiology. Although these studies supported clinical trials which used non-selective ACAT inhibitors, these trials did not report benefits, and one showed an increased risk. Early genetic studies have implicated common variants in both genes with human traits, including lipoprotein levels, coronary artery disease, and Alzheimer’s disease; however, modern genome-wide association studies have not replicated these associations. In contrast, the common SOAT1 variants are most reproducibly associated with testosterone levels. Keywords: cholesterol esterification; atherosclerosis; ACAT; SOAT; inhibitors; clinical trial Citation: Hai, Q.; Smith, J.D. Acyl-Coenzyme A: Cholesterol Acyltransferase (ACAT) in Cholesterol Metabolism: From Its 1. Introduction Discovery to Clinical Trials and the The acyl-coenzyme A:cholesterol acyltransferase (ACAT; EC 2.3.1.26) enzyme family Genomics Era. Metabolites 2021, 11, consists of membrane-spanning proteins, which are primarily located in the endoplasmic 543. https://doi.org/10.3390/ reticulum [1]. -
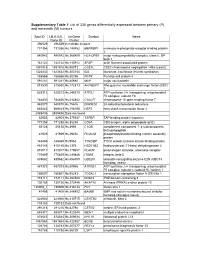
Supplementary Table 1 List of 335 Genes Differentially Expressed Between Primary (P) and Metastatic (M) Tumours
Supplementary Table 1 List of 335 genes differentially expressed between primary (P) and metastatic (M) tumours Spot ID I.M.A.G.E. UniGene Symbol Name Clone ID Cluster 296529 296529 In multiple clusters 731356 731356 Hs.140452 M6PRBP1 mannose-6-phosphate receptor binding protein 1 840942 840942 Hs.368409 HLA-DPB1 major histocompatibility complex, class II, DP beta 1 142122 142122 Hs.115912 AFAP actin filament associated protein 1891918 1891918 Hs.90073 CSE1L CSE1 chromosome segregation 1-like (yeast) 1323432 1323432 Hs.303154 IDS iduronate 2-sulfatase (Hunter syndrome) 788566 788566 Hs.80296 PCP4 Purkinje cell protein 4 591281 591281 Hs.80680 MVP major vault protein 815530 815530 Hs.172813 ARHGEF7 Rho guanine nucleotide exchange factor (GEF) 7 825312 825312 Hs.246310 ATP5J ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F6 784830 784830 Hs.412842 C10orf7 chromosome 10 open reading frame 7 840878 840878 Hs.75616 DHCR24 24-dehydrocholesterol reductase 669443 669443 Hs.158195 HSF2 heat shock transcription factor 2 2485436 2485436 Data not found 82903 82903 Hs.370937 TAPBP TAP binding protein (tapasin) 771258 771258 Hs.85258 CD8A CD8 antigen, alpha polypeptide (p32) 85128 85128 Hs.8986 C1QB complement component 1, q subcomponent, beta polypeptide 41929 41929 Hs.39252 PICALM phosphatidylinositol binding clathrin assembly protein 148469 148469 Hs.9963 TYROBP TYRO protein tyrosine kinase binding protein 415145 415145 Hs.1376 HSD11B2 hydroxysteroid (11-beta) dehydrogenase 2 810017 810017 Hs.179657 PLAUR plasminogen activator, -

Health-Relevant Phenotypes in the Offspring of Mice Given CAR Activators Prior to Pregnancy S
Supplemental material to this article can be found at: http://dmd.aspetjournals.org/content/suppl/2018/08/28/dmd.118.082925.DC1 1521-009X/46/11/1827–1835$35.00 https://doi.org/10.1124/dmd.118.082925 DRUG METABOLISM AND DISPOSITION Drug Metab Dispos 46:1827–1835, November 2018 Copyright ª 2018 by The American Society for Pharmacology and Experimental Therapeutics Health-Relevant Phenotypes in the Offspring of Mice Given CAR Activators Prior to Pregnancy s Karin Dietrich, Jan Baumgart, Leonid Eshkind, Lea Reuter, Ute Gödtel-Armbrust, Elke Butt, Michael Musheev, Federico Marini, Piyush More, Tanja Grosser, Christof Niehrs, Leszek Wojnowski, and Marianne Mathäs Department of Pharmacology (K.D., L.R., U.G.-A., P.M., T.G., L.W., M.Ma.) and Institute of Medical Biostatistics, Epidemiology and Informatics (F.M.), University Medical Center Mainz, Mainz, Germany; Translational Animal Research Center (J.B., L.E.), University Medical Center, Johannes Gutenberg University Mainz, Mainz, Germany; Institute of Experimental Biomedicine II, University Hospital Würzburg, Würzburg, Germany (E.B.); Institute of Molecular Biology, Mainz, Germany (M.Mu., C.N.); and Division of Molecular Embryology, German Cancer Research Center (DKFZ-ZMBH Alliance), Heidelberg, Germany (C.N.) Received June 6, 2018; accepted August 22, 2018 Downloaded from ABSTRACT Hepatic induction in response to drugs and environmental chem- analyses. The important environmental pollutant PCB153 acti- icals affects drug therapies and energy metabolism. We investigated vated CAR in the F1 generation in a manner similar to TCPOBOP. whether the induction is transmitted to the offspring. We injected Our findings indicate that chemicals accumulating and persisting 3-day- and 6-week-old F0 female mice with TCPOBOP, an activator in adipose tissue may exert liver-mediated, health-relevant effects dmd.aspetjournals.org of the nuclear receptor constitutive androstane receptor (CAR, on F1 offspring simply via physical transmission in milk. -
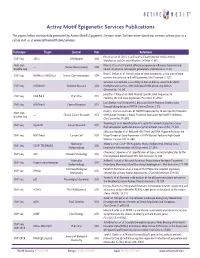
Active Motif Epigenetic Services Publications the Papers Below Contain Data Generated by Active Motif’S Epigenetic Services Team
Active Motif Epigenetic Services Publications The papers below contain data generated by Active Motif’s Epigenetic Services team. To learn more about our services, please give us a call or visit us at www.activemotif.com/services. Technique Target Journal Year Reference Erin Stashi et al. SRC-2 Is an Essential Coactivator for Orchestrating ChIP-Seq SRC-2 Cell Reports 2014 Metabolism and Circadian Rhythm. Cell Rep. 6: 663 Next-Gen Brian G. Dias et al. Parental olfactory experience influences behavior and — Nature Neuroscience 2014 Bisulfite Seq neural structure in subsequent generations. Nat Neurosci. 17: 89 Brian C. Belyea et al. Identification of renin progenitors in the mouse bone ChIP-Seq H3K4me3, H3K27me3 Nature Communications 2014 marrow that give rise to B-cell leukaemia. Nat Commun. 5: 3273 Variation in chromatin accessibility in human kidney cancer links H3K36 ChIP-Seq H3K36me3 Genome Research 2014 methyltransferase loss with widespread RNA processing defects. Genome Res. 24: 241 Jonathan P. Riley et al. PARP-14 Binds Specific DNA Sequences to ChIP-Seq RNA Pol II PLoS One 2013 Promote Th2 Cell Gene Expression. PLoS One. 8: e83127 Lynn Bjerke et al. Histone H3.3 Mutations Drive Pediatric Glioblastoma ChIP-Seq H3K36me3 Cancer Discovery 2013 through Upregulation of MYCN. Cancer Discov. 3: 512 David S. Shames et al. Loss of NAPRT1 Expression by Tumor-Specific Promoter Next-Gen — Clinical Cancer Research 2013 Methylation Provides a Novel Predictive Biomarker for NAMPT Inhibitors. Bisulfite Seq Clin Cancer Res. 19: 6912 Xiaoming Ju et al. Identification of a cyclin D1 network in prostate cancer ChIP-Seq Cyclin D1 Cancer Research 2013 that antagonizes epithelial-mesenchymal restraint. -

Transcriptional and Post-Transcriptional Regulation of ATP-Binding Cassette Transporter Expression
Transcriptional and Post-transcriptional Regulation of ATP-binding Cassette Transporter Expression by Aparna Chhibber DISSERTATION Submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILOSOPHY in Pharmaceutical Sciences and Pbarmacogenomies in the Copyright 2014 by Aparna Chhibber ii Acknowledgements First and foremost, I would like to thank my advisor, Dr. Deanna Kroetz. More than just a research advisor, Deanna has clearly made it a priority to guide her students to become better scientists, and I am grateful for the countless hours she has spent editing papers, developing presentations, discussing research, and so much more. I would not have made it this far without her support and guidance. My thesis committee has provided valuable advice through the years. Dr. Nadav Ahituv in particular has been a source of support from my first year in the graduate program as my academic advisor, qualifying exam committee chair, and finally thesis committee member. Dr. Kathy Giacomini graciously stepped in as a member of my thesis committee in my 3rd year, and Dr. Steven Brenner provided valuable input as thesis committee member in my 2nd year. My labmates over the past five years have been incredible colleagues and friends. Dr. Svetlana Markova first welcomed me into the lab and taught me numerous laboratory techniques, and has always been willing to act as a sounding board. Michael Martin has been my partner-in-crime in the lab from the beginning, and has made my days in lab fly by. Dr. Yingmei Lui has made the lab run smoothly, and has always been willing to jump in to help me at a moment’s notice. -

Epigenetic Services Citations
Active Motif Epigenetic Services Publications The papers below contain data generated by Active Motif’s Epigenetic Services team. To learn more about our services, please give us a call or visit us at www.activemotif.com/services. Technique Target Journal Year Reference Justin C. Boucher et al. CD28 Costimulatory Domain- ATAC-Seq, Cancer Immunol. Targeted Mutations Enhance Chimeric Antigen Receptor — 2021 RNA-Seq Res. T-cell Function. Cancer Immunol. Res. doi: 10.1158/2326- 6066.CIR-20-0253. Satvik Mareedu et al. Sarcolipin haploinsufficiency Am. J. Physiol. prevents dystrophic cardiomyopathy in mdx mice. RNA-Seq — Heart Circ. 2021 Am J Physiol Heart Circ Physiol. doi: 10.1152/ Physiol. ajpheart.00601.2020. Gabi Schutzius et al. BET bromodomain inhibitors regulate Nature Chemical ChIP-Seq BRD4 2021 keratinocyte plasticity. Nat. Chem. Biol. doi: 10.1038/ Biology s41589-020-00716-z. Siyun Wang et al. cMET promotes metastasis and ChIP-qPCR FOXO3 J. Cell Physiol. 2021 epithelial-mesenchymal transition in colorectal carcinoma by repressing RKIP. J. Cell Physiol. doi: 10.1002/jcp.30142. Sonia Iyer et al. Genetically Defined Syngeneic Mouse Models of Ovarian Cancer as Tools for the Discovery of ATAC-Seq — Cancer Discovery 2021 Combination Immunotherapy. Cancer Discov. doi: doi: 10.1158/2159-8290 Vinod Krishna et al. Integration of the Transcriptome and Genome-Wide Landscape of BRD2 and BRD4 Binding BRD2, BRD4, RNA Motifs Identifies Key Superenhancer Genes and Reveals ChIP-Seq J. Immunol. 2021 Pol II the Mechanism of Bet Inhibitor Action in Rheumatoid Arthritis Synovial Fibroblasts. J. Immunol. doi: doi: 10.4049/ jimmunol.2000286. Daniel Haag et al. -
Integrative Bulk and Single-Cell Profiling of Premanufacture T-Cell Populations Reveals Factors Mediating Long-Term Persistence of CAR T-Cell Therapy
Published OnlineFirst April 5, 2021; DOI: 10.1158/2159-8290.CD-20-1677 RESEARCH ARTICLE Integrative Bulk and Single-Cell Profiling of Premanufacture T-cell Populations Reveals Factors Mediating Long-Term Persistence of CAR T-cell Therapy Gregory M. Chen1, Changya Chen2,3, Rajat K. Das2, Peng Gao2, Chia-Hui Chen2, Shovik Bandyopadhyay4, Yang-Yang Ding2,5, Yasin Uzun2,3, Wenbao Yu2, Qin Zhu1, Regina M. Myers2, Stephan A. Grupp2,5, David M. Barrett2,5, and Kai Tan2,3,5 Downloaded from cancerdiscovery.aacrjournals.org on October 1, 2021. © 2021 American Association for Cancer Research. Published OnlineFirst April 5, 2021; DOI: 10.1158/2159-8290.CD-20-1677 ABSTRACT The adoptive transfer of chimeric antigen receptor (CAR) T cells represents a breakthrough in clinical oncology, yet both between- and within-patient differences in autologously derived T cells are a major contributor to therapy failure. To interrogate the molecular determinants of clinical CAR T-cell persistence, we extensively characterized the premanufacture T cells of 71 patients with B-cell malignancies on trial to receive anti-CD19 CAR T-cell therapy. We performed RNA-sequencing analysis on sorted T-cell subsets from all 71 patients, followed by paired Cellular Indexing of Transcriptomes and Epitopes (CITE) sequencing and single-cell assay for transposase-accessible chromatin sequencing (scATAC-seq) on T cells from six of these patients. We found that chronic IFN signaling regulated by IRF7 was associated with poor CAR T-cell persistence across T-cell subsets, and that the TCF7 regulon not only associates with the favorable naïve T-cell state, but is maintained in effector T cells among patients with long-term CAR T-cell persistence. -

Human 3B-Hydroxysteroid Dehydrogenase Deficiency Seems
M-A Burckhardt and others Histology of HSD3B2 deficiency 173:5 K1–K12 Case Report Human 3b-hydroxysteroid dehydrogenase deficiency seems to affect fertility but may not harbor a tumor risk: lesson from an experiment of nature Marie-Anne Burckhardt1, Sameer S Udhane1,†, Nesa Marti1,†, Isabelle Schnyder2, Coya Tapia3, John E Nielsen4, Primus E Mullis1, Ewa Rajpert-De Meyts4 and Christa E Flu¨ ck1 1Pediatric Endocrinology and Diabetology, Departments of Pediatrics and Clinical Research, 2Pediatric Surgery and Correspondence 3Institute of Pathology, University of Bern, CH-3010 Bern, Switzerland and 4Department of Growth and should be addressed Reproduction, Rigshospitalet, Copenhagen University Hospital, Copenhagen, Denmark to C E Flu¨ ck †S S Udhane and N Marti are now at Graduate School Bern, University of Bern, Bern Switzerland Email christa.fl[email protected] Abstract Context:3b-hydroxysteroid dehydrogenase deficiency (3bHSD) is a rare disorder of sexual development and steroidogenesis. There are two isozymes of 3bHSD, HSD3B1 and HSD3B2. Human mutations are known for the HSD3B2 gene which is expressed in the gonads and the adrenals. Little is known about testis histology, fertility and malignancy risk. Objective: To describe the molecular genetics, the steroid biochemistry, the (immuno-)histochemistry and the clinical implications of a loss-of-function HSD3B2 mutation. Methods: Biochemical, genetic and immunohistochemical investigations on human biomaterials. Results: A 46,XY boy presented at birth with severe undervirilization of the external genitalia. Steroid profiling showed low steroid production for mineralocorticoids, glucocorticoids and sex steroids with typical precursor metabolites for HSD3B2 European Journal of Endocrinology deficiency. The genetic analysis of the HSD3B2 gene revealed a homozygous c.687del27 deletion. -

WO 2019/089592 Al 09 May 2019 (09.05.2019) W 1 P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization III International Bureau (10) International Publication Number (43) International Publication Date WO 2019/089592 Al 09 May 2019 (09.05.2019) W 1 P O PCT (51) International Patent Classification: (72) Inventors; and A61K 35/12 (2015.01) C07K 16/30 (2006.01) (71) Applicants: JAN, Max [US/US]; c/o The Broad Institute, A61K 5/7 7 (2015.01) C07K 14/705 (2006.01) Inc., 45 1 Main Street, Cambridge, Massachusetts 02142 (US). SIEVERS, Quinlan [US/US]; c/o The Broad In¬ (21) International Application Number: stitute, Inc., 451 Main Street, Cambridge, Massachusetts PCT/US2018/058210 02142 (US). EBERT, Benjamin [US/US]; c/o The Broad (22) International Filing Date: Institute, Inc., 451 Main Street, Cambridge, Massachusetts 30 October 2018 (30. 10.2018) 02142 (US). MAUS, Marcela [US/US]; c/o The Broad In¬ stitute, Inc., 451 Main Street, Cambridge, Massachusetts (25) Filing Language: English 02142 (US). (26) Publication Language: English (74) Agent: COWLES, Christopher R. et al; Burns & Levin- (30) Priority Data: son, LLP, 125 Summer Street, Boston, Massachusetts 62/579,454 3 1 October 2017 (3 1. 10.2017) U S 021 10 (US). 62/633,725 22 February 2018 (22.02.2018) U S (81) Designated States (unless otherwise indicated, for every (71) Applicants: THE GENERAL HOSPITAL CORPO¬ kind of national protection available): AE, AG, AL, AM, RATION [US/US]; 55 Fruit Street, Boston, Massachu¬ AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, setts 021 14 (US) BRIGHAM & WOMEN'S HOSPITAL CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, [US/US]; 75 Francis Street, Boston, Massachusetts 021 15 DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, (US) PRESIDENT AND FELLOWS OF HARVARD HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, COLLEGE [US/US]; 17 Quincy Street, Cambridge, Mass¬ KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, achusetts 02138 (US). -

Identification of Transcriptional Mechanisms Downstream of Nf1 Gene Defeciency in Malignant Peripheral Nerve Sheath Tumors Daochun Sun Wayne State University
Wayne State University DigitalCommons@WayneState Wayne State University Dissertations 1-1-2012 Identification of transcriptional mechanisms downstream of nf1 gene defeciency in malignant peripheral nerve sheath tumors Daochun Sun Wayne State University, Follow this and additional works at: http://digitalcommons.wayne.edu/oa_dissertations Recommended Citation Sun, Daochun, "Identification of transcriptional mechanisms downstream of nf1 gene defeciency in malignant peripheral nerve sheath tumors" (2012). Wayne State University Dissertations. Paper 558. This Open Access Dissertation is brought to you for free and open access by DigitalCommons@WayneState. It has been accepted for inclusion in Wayne State University Dissertations by an authorized administrator of DigitalCommons@WayneState. IDENTIFICATION OF TRANSCRIPTIONAL MECHANISMS DOWNSTREAM OF NF1 GENE DEFECIENCY IN MALIGNANT PERIPHERAL NERVE SHEATH TUMORS by DAOCHUN SUN DISSERTATION Submitted to the Graduate School of Wayne State University, Detroit, Michigan in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY 2012 MAJOR: MOLECULAR BIOLOGY AND GENETICS Approved by: _______________________________________ Advisor Date _______________________________________ _______________________________________ _______________________________________ © COPYRIGHT BY DAOCHUN SUN 2012 All Rights Reserved DEDICATION This work is dedicated to my parents and my wife Ze Zheng for their continuous support and understanding during the years of my education. I could not achieve my goal without them. ii ACKNOWLEDGMENTS I would like to express tremendous appreciation to my mentor, Dr. Michael Tainsky. His guidance and encouragement throughout this project made this dissertation come true. I would also like to thank my committee members, Dr. Raymond Mattingly and Dr. John Reiners Jr. for their sustained attention to this project during the monthly NF1 group meetings and committee meetings, Dr. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in