Advances in Genomics for Drug Development
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

CENTRE for POPULATION GENOMICS Strategic Plan 2021-2022 CENTRE for POPULATION GENOMICS Strategic Plan 2021-2022
CENTRE FOR POPULATION GENOMICS Strategic Plan 2021-2022 CENTRE FOR POPULATION GENOMICS Strategic Plan 2021-2022 • Plan-on-a-page page 3 • Vision page 4 • Purpose page 5 • Goals page 6 • Objectives page 7 • Goal 1 detail page 8 - 10 • Goal 2 detail page 11 - 13 • Goal 3 detail page 14 - 16 • Goal 4 detail page 17 - 19 • Principles page 20 • Operating model page 21 - 25 • Operational plan page 26 • Monitoring & evaluation plan page 27 – 28 • Team values page 29 CENTRE FOR POPULATION GENOMICS Strategic Plan-on-a-page 2021-2022 Vision: A world in which genomic information enables comprehensive disease prediction, accurate diagnosis and effective therapeutics for all people Purpose: To establish respectful partnerships with diverse communities, collect and analyse genomic data at transformative scale and drive genomic discovery and equitable genomic medicine in Australia Goals: Community Computational Genomic Scientific & medical partnerships infrastructure datasets knowledge Principles: Respect Diversity Openness Scalability Connectedness 3 CENTRE FOR POPULATION GENOMICS Strategic Plan 2021-2022 Vision: A world in which genomic information enables comprehensive disease prediction, accurate diagnosis and effective therapeutics for all people • The next decade will see a transformation of medicine and biology, driven in part by an explosion in our understanding of the connections between human genetic variation and individuals’ traits. This knowledge will allow the dissection of the biological basis of human traits, improve the prediction and diagnosis of disease, and accelerate the discovery and validation of new therapeutic targets. Key to these discoveries will be population genomics: the generation and analysis of massive-scale data sets of human genetic variation, combined with information on health and clinical outcomes. -

Pooled CRISPR-Activation Screening Coupled with Single-Cell RNA-Seq in Mouse Embryonic Stem Cells
ll OPEN ACCESS Protocol Pooled CRISPR-activation screening coupled with single-cell RNA-seq in mouse embryonic stem cells Celia Alda-Catalinas, Melanie A. Eckersley-Maslin, Wolf Reik celia.x.aldacatalinas@gsk. com (C.A.-C.) [email protected]. uk (W.R.) Highlights Protocol for CRISPRa screens with single- cell readout to interrogate gene function Detailed description of CRISPRa screening procedures in mouse embryonic stem cells Detailed steps on how to construct derived single-cell sgRNA amplicon libraries CRISPR/Cas9 screens are a powerful approach to identify key regulators of biological processes. By combining pooled CRISPR/Cas9 screening with a single-cell RNA-sequencing readout, individual perturbations can be assessed in parallel both comprehensively and at scale. Importantly, this allows gene function and regulation to be interrogated at a cellular level in an unbiased manner. Here, we present a protocol to perform pooled CRISPR-activation screens in mouse embryonic stem cells using 103 Genomics scRNA-seq as a readout. Alda-Catalinas et al., STAR Protocols 2, 100426 June 18, 2021 ª 2021 The Authors. https://doi.org/10.1016/ j.xpro.2021.100426 ll OPEN ACCESS Protocol Pooled CRISPR-activation screening coupled with single-cell RNA-seq in mouse embryonic stem cells Celia Alda-Catalinas,1,4,7,* Melanie A. Eckersley-Maslin,1,5,6 and Wolf Reik1,2,3,8,* 1Epigenetics Programme, Babraham Institute, Cambridge CB22 3AT, UK 2Wellcome Trust Sanger Institute, Hinxton, Cambridge CB10 1SA, UK 3Centre for Trophoblast Research, University of -

A CRISPR Activation and Interference Toolkit for Industrial Saccharomyces Cerevisiae Strain KE6‑12 Elena Cámara, Ibai Lenitz & Yvonne Nygård*
www.nature.com/scientificreports OPEN A CRISPR activation and interference toolkit for industrial Saccharomyces cerevisiae strain KE6‑12 Elena Cámara, Ibai Lenitz & Yvonne Nygård* Recent advances in CRISPR/Cas9 based genome editing have considerably advanced genetic engineering of industrial yeast strains. In this study, we report the construction and characterization of a toolkit for CRISPR activation and interference (CRISPRa/i) for a polyploid industrial yeast strain. In the CRISPRa/i plasmids that are available in high and low copy variants, dCas9 is expressed alone, or as a fusion with an activation or repression domain; VP64, VPR or Mxi1. The sgRNA is introduced to the CRISPRa/i plasmids from a double stranded oligonucleotide by in vivo homology‑directed repair, allowing rapid transcriptional modulation of new target genes without cloning. The CRISPRa/i toolkit was characterized by alteration of expression of fuorescent protein‑encoding genes under two diferent promoters allowing expression alterations up to ~ 2.5‑fold. Furthermore, we demonstrated the usability of the CRISPRa/i toolkit by improving the tolerance towards wheat straw hydrolysate of our industrial production strain. We anticipate that our CRISPRa/i toolkit can be widely used to assess novel targets for strain improvement and thus accelerate the design‑build‑test cycle for developing various industrial production strains. Te yeast Saccharomyces cerevisiae is one of the most commonly used microorganisms for industrial applications ranging from wine and beer fermentations to the production of biofuels and high-value metabolites1,2. How- ever, some of the current production processes are compromised by low yields and productivities, thus further optimization is required3. -

Spatiotemporal Control of CRISPR/Cas9 Gene Editing
Signal Transduction and Targeted Therapy www.nature.com/sigtrans REVIEW ARTICLE OPEN Spatiotemporal control of CRISPR/Cas9 gene editing Chenya Zhuo1, Jiabin Zhang1, Jung-Hwan Lee2, Ju Jiao3, Du Cheng4, Li Liu5, Hae-Won Kim2,YuTao1 and Mingqiang Li 1,6 The clustered regularly interspaced short palindromic repeats (CRISPR)/associated protein 9 (CRISPR/Cas9) gene editing technology, as a revolutionary breakthrough in genetic engineering, offers a promising platform to improve the treatment of various genetic and infectious diseases because of its simple design and powerful ability to edit different loci simultaneously. However, failure to conduct precise gene editing in specific tissues or cells within a certain time may result in undesirable consequences, such as serious off-target effects, representing a critical challenge for the clinical translation of the technology. Recently, some emerging strategies using genetic regulation, chemical and physical strategies to regulate the activity of CRISPR/Cas9 have shown promising results in the improvement of spatiotemporal controllability. Herein, in this review, we first summarize the latest progress of these advanced strategies involving cell-specific promoters, small-molecule activation and inhibition, bioresponsive delivery carriers, and optical/thermal/ultrasonic/magnetic activation. Next, we highlight the advantages and disadvantages of various strategies and discuss their obstacles and limitations in clinical translation. Finally, we propose viewpoints on directions that can be explored to -

Whole-Genome Analysis of Polymorphism and Divergence in Drosophila Simulans David J
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2007 Population genomics: Whole-genome analysis of polymorphism and divergence in Drosophila simulans David J. Begun University of California - Davis Alisha K. Holloway University of California - Davis Kristian Stevens University of California - Davis LaDeana W. Hillier Washington University School of Medicine in St. Louis Yu-Ping Poh University of California - Davis See next page for additional authors Follow this and additional works at: https://digitalcommons.wustl.edu/open_access_pubs Part of the Medicine and Health Sciences Commons Recommended Citation Begun, David J.; Holloway, Alisha K.; Stevens, Kristian; Hillier, LaDeana W.; Poh, Yu-Ping; Hahn, Matthew W.; Nista, Phillip M.; Jones, Corbin D.; Kern, Andrew D.; Dewey, Colin N.; Pachter, Lior; Myers, Eugene; and Langley, Charles H., ,"Population genomics: Whole-genome analysis of polymorphism and divergence in Drosophila simulans." PLoS Biology.,. e310. (2007). https://digitalcommons.wustl.edu/open_access_pubs/877 This Open Access Publication is brought to you for free and open access by Digital Commons@Becker. It has been accepted for inclusion in Open Access Publications by an authorized administrator of Digital Commons@Becker. For more information, please contact [email protected]. Authors David J. Begun, Alisha K. Holloway, Kristian Stevens, LaDeana W. Hillier, Yu-Ping Poh, Matthew W. Hahn, Phillip M. Nista, Corbin D. Jones, Andrew D. Kern, Colin N. Dewey, Lior Pachter, Eugene Myers, and Charles H. Langley This open access publication is available at Digital Commons@Becker: https://digitalcommons.wustl.edu/open_access_pubs/877 PLoS BIOLOGY Population Genomics: Whole-Genome Analysis of Polymorphism and Divergence in Drosophila simulans David J. -

Population and Comparative Genomics Inform Our Understanding of Bacterial Species Diversity in the Soil
Chapter 11 Population and Comparative Genomics Inform Our Understanding of Bacterial Species Diversity in the Soil Margaret A. Riley 11.1 Introduction The diversity of soil bacteria is simply staggering. According to Torsvik and colleagues (Torsvik et al. 1990; Torsvik and Ovreas 2002), species diversity in soil samples is so high that our most powerful methods of estimation provide only the crudest measure of its magnitude. Nonetheless, many such estimates exist, and they suggest that a single gram of soil may contain over 10 billion microbial cells and more than 1,800 bacterial species (Torsvik and Ovreas 2002; Gans et al. 2005; Zhang et al. 2008). An equally compelling estimate is provided by Dykhuizen (Dykhuizen and Dean 2004), who examined levels of genetic diversity in soil bacteria and predicted that 30 g of forest soil contains over half a million species! As our methods of empirically estimating bacterial diversity improve, so to do our methods of mathematically refining these numbers. For example, some analyti- cal methods now take into account the fact that there are very few common soil species and untold numbers of rare species (Youssef and Elshahed 2009; Schloss and Handelsman 2006). These refinements push our estimates of bacterial diversity to over a million species per gram of soil (Gans et al. 2005). To put these numbers into perspective, similar studies of the human gastrointestinal tract predict a mere 400 distinct bacterial phylotypes (Rajilic-Stojanovic et al. 2007). This staggering level of species diversity is even more remarkable when one considers that the number of prokaryotes reported in the National Center for Biotechnology Informa- tion molecular database is only about 15,111 (Sayers et al. -

A Review on the Population Genomics of Transposable Elements
G C A T T A C G G C A T genes Review On the Population Dynamics of Junk: A Review on the Population Genomics of Transposable Elements Yann Bourgeois and Stéphane Boissinot * New York University Abu Dhabi, P.O. 129188 Saadiyat Island, Abu Dhabi, UAE; [email protected] * Correspondence: [email protected] Received: 4 April 2019; Accepted: 21 May 2019; Published: 30 May 2019 Abstract: Transposable elements (TEs) play an important role in shaping genomic organization and structure, and may cause dramatic changes in phenotypes. Despite the genetic load they may impose on their host and their importance in microevolutionary processes such as adaptation and speciation, the number of population genetics studies focused on TEs has been rather limited so far compared to single nucleotide polymorphisms (SNPs). Here, we review the current knowledge about the dynamics of transposable elements at recent evolutionary time scales, and discuss the mechanisms that condition their abundance and frequency. We first discuss non-adaptive mechanisms such as purifying selection and the variable rates of transposition and elimination, and then focus on positive and balancing selection, to finally conclude on the potential role of TEs in causing genomic incompatibilities and eventually speciation. We also suggest possible ways to better model TEs dynamics in a population genomics context by incorporating recent advances in TEs into the rich information provided by SNPs about the demography, selection, and intrinsic properties of genomes. Keywords: transposable elements; population genetics; selection; drift; coevolution 1. Introduction Transposable elements (TEs) are repetitive DNA sequences that are ubiquitous in the living world and have the ability to replicate and multiply within genomes. -

The Genetic Structure of the Belgian Population Jimmy Van Den Eynden1,2* , Tine Descamps1, Els Delporte1, Nancy H
Van den Eynden et al. Human Genomics (2018) 12:6 DOI 10.1186/s40246-018-0136-8 PRIMARY RESEARCH Open Access The genetic structure of the Belgian population Jimmy Van den Eynden1,2* , Tine Descamps1, Els Delporte1, Nancy H. C. Roosens1, Sigrid C. J. De Keersmaecker1, Vanessa De Wit1, Joris Robert Vermeesch3, Els Goetghebeur4, Jean Tafforeau1, Stefaan Demarest1, Marc Van den Bulcke1 and Herman Van Oyen1,5* Abstract Background: National and international efforts like the 1000 Genomes Project are leading to increasing insights in the genetic structure of populations worldwide. Variation between different populations necessitates access to population-based genetic reference datasets. These data, which are important not only in clinical settings but also to potentiate future transitions towards a more personalized public health approach, are currently not available for the Belgian population. Results: To obtain a representative genetic dataset of the Belgian population, participants in the 2013 National Health Interview Survey (NHIS) were invited to donate saliva samples for DNA analysis. DNA was isolated and single nucleotide polymorphisms (SNPs) were determined using a genome-wide SNP array of around 300,000 sites, resulting in a high-quality dataset of 189 samples that was used for further analysis. A principal component analysis demonstrated the typical European genetic constitution of the Belgian population, as compared to other continents. Within Europe, the Belgian population could be clearly distinguished from other European populations. Furthermore, obvious signs from recent migration were found, mainly from Southern Europe and Africa, corresponding with migration trends from the past decades. Within Belgium, a small north-west to south-east gradient in genetic variability was noted, with differences between Flanders and Wallonia. -

A Population Genomics Approach Shows Widespread Geographical Distribution of Cryptic Genomic Forms of the Symbiotic Fungus Rhizophagus Irregularis
OPEN The ISME Journal (2018) 12, 17–30 www.nature.com/ismej ORIGINAL ARTICLE A population genomics approach shows widespread geographical distribution of cryptic genomic forms of the symbiotic fungus Rhizophagus irregularis Romain Savary1, Frédéric G Masclaux1,2, Tania Wyss1, Germain Droh1,3, Joaquim Cruz Corella1, Ana Paula Machado1, Joseph B Morton4 and Ian R Sanders1 1Department of Ecology and Evolution, Biophore Building, University of Lausanne, Lausanne, Switzerland; 2Vital-IT Group, SIB Swiss Institute of Bioinformatics, Lausanne, Switzerland; 3Laboratoire de Génétique, Unité de Formation et de Recherche en Biosciences, Université Félix Houphouet Boigny, Abidjan, Ivory Coast and 4Division of Plant and Soil Sciences, West Virginia University, Morgantown, WV, USA Arbuscular mycorrhizal fungi (AMF; phylum Gomeromycota) associate with plants forming one of the most successful microbe–plant associations. The fungi promote plant diversity and have a potentially important role in global agriculture. Plant growth depends on both inter- and intra-specific variation in AMF. It was recently reported that an unusually large number of AMF taxa have an intercontinental distribution, suggesting long-distance gene flow for many AMF species, facilitated by either long- distance natural dispersal mechanisms or human-assisted dispersal. However, the intercontinental distribution of AMF species has been questioned because the use of very low-resolution markers may be unsuitable to detect genetic differences among geographically separated AMF, as seen with some other fungi. This has been untestable because of the lack of population genomic data, with high resolution, for any AMF taxa. Here we use phylogenetics and population genomics to test for intra- specific variation in Rhizophagus irregularis, an AMF species for which genome sequence information already exists. -
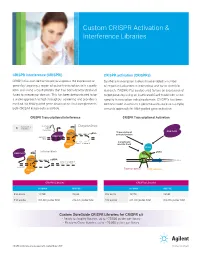
Custom CRISPR Activation & Interference Libraries
Custom CRISPR Activation & Interference Libraries CRISPR interference (CRISPRi) CRISPR activation (CRISPRa) CRISPR/Cas can be harnessed to suppress the expression of Synthetic transcription factors have enabled a number genes by targeting a region of active transcription with a guide of important advances in biomedical and basic scientific RNA and using a Cas9 protein that has been deactivated and research. CRISPR/Cas can be used to turn on expression of fused to a repressor domain. This has been demonstrated to be target genes by using an inactivated Cas9 fused with a non- a viable approach to high throughput screening and provides a specific transcription inducing domain. CRISPRa has been method for RNA-guided gene deactivation that complements demonstrated to work on a genome-wide scale as a simple, both CRISPR knock-outs and RNAi. versatile approach for RNA-guided gene activation. CRISPR Transcriptional Interference CRISPR Transcriptional Activation inactivating interrupted Elongation Block mutations mRNA x transcript RNA Pol II sgRNA Transcriptional activator protein genomic RNA Pol II DNA Catalytically dCAS9 Exon 1 of gene X inactive Cas9 VP64 Initiation Block gRNA RNA Pol II dCAS9 x sgRNA Gene of interest Exon 1 of gene X promoter Target sequence dCAS9 PAM sequence CRISPRi Libraries CRISPRa Libraries HUMAN MOUSE HUMAN MOUSE # of Genes 18,730 19,846 # of Genes 18,574 19,949 # of Guides 205,648 guides total 212,376 guides total # of Guides 201,530 guides total 208,066 guides total Custom SureGuide CRISPR Libraries for CRISPR a/i • Ready-to-Amplify libraries, up to ~70,000 guides per library • Ready-to-Clone libraries, up to ~70,000 guides per library CRISPR a/i libraries are designed with content from UCSF. -

A Novel Eukaryote‐Like CRISPR Activation Tool in Bacteria
METHODS, MODELS & TECHNIQUES Prospects & Overviews www.bioessays-journal.com A Novel Eukaryote-Like CRISPR Activation Tool in Bacteria: Features and Capabilities Yang Liu and Baojun Wang* friend or foe” system, scientists are CRISPR (clustered regularly interspaced short palindromic repeats) activation able to guide the endonucleases to (CRISPRa) in bacteria is an attractive method for programmable gene their desired DNA or RNA targets.[5–8] activation. Recently, a eukaryote-like, 54-dependent CRISPRa system has CRISPR regulation relies on inactivated been reported. It exhibits high dynamic ranges and permits flexible target site CRISPR endonucleases. The nuclease- deficient CRISPR DNA endonucleases, selection. Here, an overview of the existing strategies of CRISPRa in bacteria for instances dCas9 and ddCpf1 (dCas12), is presented, and the characteristics and design principles of the CRISPRa are effectively programmable DNA bind- system are introduced. Possible scenarios for applying the eukaryote-like ing domains. When these domains are CRISPRa system is discussed with corresponding suggestions for tethered to transactivation domains or performance optimization and future functional expansion. The authors subunits of RNA polymerase, they ac- envision the new eukaryote-like CRISPRa system enabling novel designs in tivate the promoters near the CRISPR target sites. This strategy has been 54 multiplexed gene regulation and promoting research in the -dependent widely utilized in both eukaryotes and gene regulatory networks among a variety of biotechnology relevant or prokaryotes,[9–17] particularly in the former, disease-associated bacterial species. where the transcription activation mecha- nisms and the activators are well-studied. While CRISPRa in eukaryotes enjoys much success and is continuously im- 1. -

Human Population Genomics Heritability & Environment
ACGTTTGACTGAGGAGTTTACGGGAGCAAAGCGGCGTCATTGCTATTCGTATCTGTTTAG 010101100010010100001010101010011011100110001100101000100101 Human Population Genomics Heritability & Environment Bienvenu OJ, Davydow DS, & Kendler KS (2011). Psychological medicine, 41 (1), 33-40 PMID: Heritability & Environment Heritability & Environment Heritability & Environment Feasibility of identifying genetic variants by risk allele frequency and strength of genetic effect (odds ratio). TA Manolio et al. Nature 461, 747-753 (2009) doi:10.1038/nature08494 Where is the missing heritability? Some plausible explanations • Rare variants not captured in genotyping microarrays • Many variants of small effect • Structural variants not captured in short read sequencing • Epistatic effects: non-linear gene-gene interactions • ??? TA Manolio et al. Nature 461, 747-753 (2009) doi: 10.1038/nature08494 Global Ancestry Inference Nature. 2008 November 6; 456(7218): 98–101. Modeling population haplotypes – VLMC G , …, G ; G = g … g ; g {0, 1, 2} 0000 1 N i i1 in ij ∈ 0001 Gi = Hi1 + Hi2, where, 0011 0110 Hi = hij1 … hijn ; hijk ∈ {0, 1} 1000 1001 1011 Browning, 2006 Phasing Browning & Browning, 2007 Identity By Descent . { { IBD detection IBD = F IBD = T FastIBD: sample haplotypes for Parente each individual, check for IBD Rodriguez et al. 2013 Browning & Browining 2011 Mexican Ancestry The genetics of Mexico recapitulates Native American substructure and affects biomedical traits, Moreno-Estrada et al. Science, 2014. Fixation, Positive & Negative Selection How can we How can we detect