Polypyrimidine Tract Binding Proteins Are Essential for B Cell Development
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Ptbp2 Represses Adult-Specific Splicing to Regulate the Generation of Neuronal Precursors in the Embryonic Brain
Downloaded from genesdev.cshlp.org on October 4, 2021 - Published by Cold Spring Harbor Laboratory Press Ptbp2 represses adult-specific splicing to regulate the generation of neuronal precursors in the embryonic brain Donny D. Licatalosi,1,4 Masato Yano,1,5 John J. Fak,1 Aldo Mele,1 Sarah E. Grabinski,2 Chaolin Zhang,1 and Robert B. Darnell1,3,6 1Laboratory of Molecular Neuro-Oncology, The Rockefeller University, New York, New York 10065, USA; 2Center for RNA Molecular Biology, Case Western Reserve University, Cleveland, Ohio 44106, USA; 3Howard Hughes Medical Institute, The Rockefeller University, New York, New York 10065, USA Two polypyrimidine tract RNA-binding proteins (PTBs), one near-ubiquitously expressed (Ptbp1) and another highly tissue-restricted (Ptbp2), regulate RNA in interrelated but incompletely understood ways. Ptbp1, a splicing regulator, is replaced in the brain and differentiated neuronal cell lines by Ptbp2. To define the roles of Ptbp2 in the nervous system, we generated two independent Ptbp2-null strains, unexpectedly revealing that Ptbp2 is expressed in neuronal progenitors and is essential for postnatal survival. A HITS-CLIP (high-throughput sequencing cross- linking immunoprecipitation)-generated map of reproducible Ptbp2–RNA interactions in the developing mouse neocortex, combined with results from splicing-sensitive microarrays, demonstrated that the major action of Ptbp2 is to inhibit adult-specific alternative exons by binding pyrimidine-rich sequences upstream of and/or within them. These regulated exons are present in mRNAs encoding proteins associated with control of cell fate, proliferation, and the actin cytoskeleton, suggesting a role for Ptbp2 in neurogenesis. Indeed, neuronal progenitors in the Ptbp2-null brain exhibited an aberrant polarity and were associated with regions of premature neurogenesis and reduced progenitor pools. -

The PTB Interacting Protein Raver1 Regulates A-Tropomyosin Alternative Splicing
The EMBO Journal Vol. 22 No. 23 pp. 6356±6364, 2003 The PTB interacting protein raver1 regulates a-tropomyosin alternative splicing Natalia Gromak, Alexis Rideau, alternative splicing, can be achieved by activation or Justine Southby, A.D.J.Scadden, inhibition (Smith and ValcaÂrcel, 2000; Caceres and Clare Gooding, Stefan HuÈ ttelmaier1, Kornblihtt, 2002). Spliceosome assembly can be assisted Robert H.Singer1 and by additional interactions of `SR' splicing factors with Christopher W.J.Smith2 splicing enhancer sequences, which are commonly found within exons (Blencowe, 2000; Cartegni et al., 2002). SR Department of Biochemistry, University of Cambridge, 80 Tennis proteins have a number of distinct roles in splicing and 1 Court Road, Cambridge CB2 1GA, UK and Department of Anatomy have been characterized as both constitutive and alterna- & Structural Biology, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA tive splicing factors (Manley and Tacke, 1996; Graveley, 2000). A number of negative regulators have been 2 Corresponding author identi®ed, many of which are members of the hetero- e-mail: [email protected] geneous nuclear ribonucleoprotein (hnRNP) family Regulated switching of the mutually exclusive exons 2 (Krecic and Swanson, 1999; Smith and ValcaÂrcel, 2000; and3ofa-tropomyosin (TM) involves repression of Dreyfuss et al., 2002). These proteins contain RNA- exon 3 in smooth muscle cells. Polypyrimidine tract- binding domains as well as various accessory domains, but binding protein (PTB) is necessary but not suf®cient they do not contain the arginine-serine rich domains for regulation of TM splicing. Raver1 was identi®ed in characteristic of the SR family. -

An Exon That Prevents Transport of a Mature Mrna
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 3813–3818, March 1999 Genetics An exon that prevents transport of a mature mRNA MARGARET A. MACMORRIS*, DIEGO A. R. ZORIO*†, AND THOMAS BLUMENTHAL*‡ *Department of Biochemistry and Molecular Genetics, University of Colorado Health Sciences Center, Denver, CO 80262; and †Department of Biology, Indiana University, Bloomington, IN 47405 Communicated by William B. Wood III, University of Colorado, Boulder, CO, January 25, 1999 (received for review November 18, 1998) ABSTRACT In Caenorhabditis elegans, pre-mRNA for the recognizes (6). Here we show that uaf-1 exon 3-containing essential splicing factor U2AF65 sometimes is spliced to pro- RNA is not subject to Smg-mediated decay in spite of its duce an RNA that includes an extra 216-bp internal exon, exon containing a PTC. Although it is not yet understood how the 3. Inclusion of exon 3 inserts an in-frame stop codon, yet this components of the pathway act to recognize and degrade RNA is not subject to SMG-mediated RNA surveillance. To aberrant mRNAs, Smg is likely to operate at the site of protein test whether exon 3 causes RNA to remain nuclear and thereby synthesis. Because the yeast homolog of SMG-2, UPF1, has escape decay, we inserted it into the 3* untranslated region of been localized to the cytoplasm at the site of polyribosomes a gfp reporter gene. Although exon 3 did not affect accumu- (7), an mRNA that fails to leave the nucleus might well be lation or processing of the mRNA, it dramatically suppressed unaffected by the Smg pathway. -

Mrna Editing, Processing and Quality Control in Caenorhabditis Elegans
| WORMBOOK mRNA Editing, Processing and Quality Control in Caenorhabditis elegans Joshua A. Arribere,*,1 Hidehito Kuroyanagi,†,1 and Heather A. Hundley‡,1 *Department of MCD Biology, UC Santa Cruz, California 95064, †Laboratory of Gene Expression, Medical Research Institute, Tokyo Medical and Dental University, Tokyo 113-8510, Japan, and ‡Medical Sciences Program, Indiana University School of Medicine-Bloomington, Indiana 47405 ABSTRACT While DNA serves as the blueprint of life, the distinct functions of each cell are determined by the dynamic expression of genes from the static genome. The amount and specific sequences of RNAs expressed in a given cell involves a number of regulated processes including RNA synthesis (transcription), processing, splicing, modification, polyadenylation, stability, translation, and degradation. As errors during mRNA production can create gene products that are deleterious to the organism, quality control mechanisms exist to survey and remove errors in mRNA expression and processing. Here, we will provide an overview of mRNA processing and quality control mechanisms that occur in Caenorhabditis elegans, with a focus on those that occur on protein-coding genes after transcription initiation. In addition, we will describe the genetic and technical approaches that have allowed studies in C. elegans to reveal important mechanistic insight into these processes. KEYWORDS Caenorhabditis elegans; splicing; RNA editing; RNA modification; polyadenylation; quality control; WormBook TABLE OF CONTENTS Abstract 531 RNA Editing and Modification 533 Adenosine-to-inosine RNA editing 533 The C. elegans A-to-I editing machinery 534 RNA editing in space and time 535 ADARs regulate the levels and fates of endogenous dsRNA 537 Are other modifications present in C. -

Landscape of the Spliced Leader Trans-Splicing Mechanism In
www.nature.com/scientificreports OPEN Landscape of the spliced leader trans-splicing mechanism in Schistosoma mansoni Received: 12 July 2017 Mariana Boroni1,2, Michael Sammeth 3,4, Sandra Grossi Gava 5, Natasha Andressa Accepted: 12 February 2018 Nogueira Jorge2, Andréa Mara Macedo1, Carlos Renato Machado1, Marina Moraes Mourão5 & Published: xx xx xxxx Glória Regina Franco1 Spliced leader dependent trans-splicing (SLTS) has been described as an important RNA regulatory process that occurs in diferent organisms, including the trematode Schistosoma mansoni. We identifed more than seven thousand putative SLTS sites in the parasite, comprising genes with a wide spectrum of functional classes, which underlines the SLTS as a ubiquitous mechanism in the parasite. Also, SLTS gene expression levels span several orders of magnitude, showing that SLTS frequency is not determined by the expression level of the target gene, but by the presence of particular gene features facilitating or hindering the trans-splicing mechanism. Our in-depth investigation of SLTS events demonstrates widespread alternative trans-splicing (ATS) acceptor sites occurring in diferent regions along the entire gene body, highlighting another important role of SLTS generating alternative RNA isoforms in the parasite, besides the polycistron resolution. Particularly for introns where SLTS directly competes for the same acceptor substrate with cis-splicing, we identifed for the frst time additional and important features that might determine the type of splicing. Our study substantially extends the current knowledge of RNA processing by SLTS in S. mansoni, and provide basis for future studies on the trans-splicing mechanism in other eukaryotes. In contrast to conventional splicing in cis, trans-splicing connects exons of two diferent primary RNA molecules tran- scribed from a priori unrelated genomic loci. -
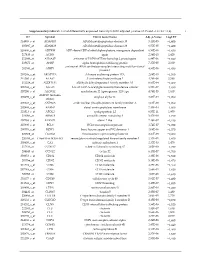
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

The Splicing Regulator PTBP1 Controls the Activity of the Transcription Factor
1 The splicing regulator PTBP1 controls the activity of the 2 transcription factor Pbx1 during neuronal differentiation. 3 4 Anthony J Linares1, Chia-Ho Lin2, Andrey Damianov2, Katrina L Adams1,3,4, Bennett G 5 Novitch3,4, Douglas L Black2,4 6 1Molecular Biology Institute Graduate Program, University of California, Los Angeles; 2Department of 7 Microbiology, Immunology and Molecular Genetics, University of California, Los Angeles; 3Department of 8 Neurobiology, David Geffen School of Medicine at UCLA; 4Eli and Edythe Broad Center of Regenerative 9 Medicine and Stem Cell Research, David Geffen School of Medicine at UCLA, Los Angeles, United States. 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 1 27 ABSTRACT 28 29 The RNA-binding proteins PTBP1 and PTBP2 control programs of alternative splicing 30 during neuronal development. PTBP2 was found to maintain embryonic splicing patterns 31 of many synaptic and cytoskeletal proteins during differentiation of neuronal progenitor 32 cells (NPCs) into early neurons. However, the role of the earlier PTBP1 program in 33 embryonic stem cells (ESCs) and NPCs was not clear. We show that PTBP1 controls a 34 program of neuronal gene expression that includes the transcription factor Pbx1. We 35 identify exons specifically regulated by PTBP1 and not PTBP2 as mouse ESCs 36 differentiate into NPCs. We find that PTBP1 represses Pbx1 exon 7 and the expression 37 of the neuronal Pbx1a isoform in ESCs. Using CRISPR-Cas9 to delete regulatory 38 elements for exon 7, we induce Pbx1a expression in ESCs, finding that this activates 39 transcription of neuronal genes. -

Significance of Polypyrimidine Tract–Binding Protein 1 Expression In
Published OnlineFirst April 22, 2015; DOI: 10.1158/1535-7163.MCT-14-0142 Cancer Biology and Signal Transduction Molecular Cancer Therapeutics Significance of Polypyrimidine Tract–Binding Protein 1 Expression in Colorectal Cancer Hidekazu Takahashi1, Junichi Nishimura1, Yoshinori Kagawa1,2, Yoshihiro Kano3, Yusuke Takahashi1,4, Xin Wu1, Masayuki Hiraki1, Atsushi Hamabe1, Masamitsu Konno3, Naotsugu Haraguchi1, Ichiro Takemasa1, Tsunekazu Mizushima1, Masaru Ishii2, Koshi Mimori4, Hideshi Ishii3, Yuichiro Doki1, Masaki Mori1, and Hirofumi Yamamoto1 Abstract Polypyrimidine tract–binding protein (PTBP1) is an RNA- observed that PTBP1 affects cell invasion, which was partially binding protein with various molecular functions related to correlated to CD44 splicing, and this correlation was also RNA metabolism and a major repressive regulator of alternative confirmed in clinical samples. PTBP1 expression also affected splicing, causing exon skipping in numerous alternatively anchorage-independent growth in colorectal cancer cell lines. spliced pre-mRNAs. Here, we have investigated the role of PTBP1 expression also affected cell proliferation. Using time- PTBP1 in colorectal cancer. PTBP1 expression levels were sig- lapse imaging analysis, PTBP1 was implicated in prolonged nificantly overexpressed in cancerous tissues compared with G2–M phase in HCT116 cells. As for the mechanism of pro- corresponding normal mucosal tissues. We also observed that longed G2–M phase in HCT116 siPTBP1 cells, Western blotting PTBP1 expression levels, c-MYC expression levels, and PKM2: revealed that PTBP1 expression level was correlated to PKM1 ratio were positively correlated in colorectal cancer CDK11p58 expression level, which was reported to play an specimens. Moreover, PTBP1 expression levels were positively important role on progression to complete mitosis. These correlated to poor prognosis and lymph node metastasis. -

A SARS-Cov-2 Protein Interaction Map Reveals Targets for Drug Repurposing
Article A SARS-CoV-2 protein interaction map reveals targets for drug repurposing https://doi.org/10.1038/s41586-020-2286-9 A list of authors and affiliations appears at the end of the paper Received: 23 March 2020 Accepted: 22 April 2020 A newly described coronavirus named severe acute respiratory syndrome Published online: 30 April 2020 coronavirus 2 (SARS-CoV-2), which is the causative agent of coronavirus disease 2019 (COVID-19), has infected over 2.3 million people, led to the death of more than Check for updates 160,000 individuals and caused worldwide social and economic disruption1,2. There are no antiviral drugs with proven clinical efcacy for the treatment of COVID-19, nor are there any vaccines that prevent infection with SARS-CoV-2, and eforts to develop drugs and vaccines are hampered by the limited knowledge of the molecular details of how SARS-CoV-2 infects cells. Here we cloned, tagged and expressed 26 of the 29 SARS-CoV-2 proteins in human cells and identifed the human proteins that physically associated with each of the SARS-CoV-2 proteins using afnity-purifcation mass spectrometry, identifying 332 high-confdence protein–protein interactions between SARS-CoV-2 and human proteins. Among these, we identify 66 druggable human proteins or host factors targeted by 69 compounds (of which, 29 drugs are approved by the US Food and Drug Administration, 12 are in clinical trials and 28 are preclinical compounds). We screened a subset of these in multiple viral assays and found two sets of pharmacological agents that displayed antiviral activity: inhibitors of mRNA translation and predicted regulators of the sigma-1 and sigma-2 receptors. -

Enriched Alternative Splicing in Islets of Diabetes-Susceptible Mice
International Journal of Molecular Sciences Article Enriched Alternative Splicing in Islets of Diabetes-Susceptible Mice Ilka Wilhelmi 1,2, Alexander Neumann 3 , Markus Jähnert 1,2 , Meriem Ouni 1,2,† and Annette Schürmann 1,2,4,5,*,† 1 Department of Experimental Diabetology, German Institute of Human Nutrition (DIfE), 14558 Potsdam, Germany; [email protected] (I.W.); [email protected] (M.J.); [email protected] (M.O.) 2 German Center for Diabetes Research (DZD), 85764 Munich, Germany 3 Omiqa Bioinformatics, 14195 Berlin, Germany; [email protected] 4 Institute of Nutritional Sciences, University of Potsdam, 14558 Nuthetal, Germany 5 Faculty of Health Sciences, Joint Faculty of the Brandenburg University of Technology Cottbus-Senftenberg, The Brandenburg Medical School Theodor Fontane and The University of Potsdam, 14469 Potsdam, Germany * Correspondence: [email protected] † These authors contributed equally to this work. Abstract: Dysfunctional islets of Langerhans are a hallmark of type 2 diabetes (T2D). We hypoth- esize that differences in islet gene expression alternative splicing which can contribute to altered protein function also participate in islet dysfunction. RNA sequencing (RNAseq) data from islets of obese diabetes-resistant and diabetes-susceptible mice were analyzed for alternative splicing and its putative genetic and epigenetic modulators. We focused on the expression levels of chromatin modifiers and SNPs in regulatory sequences. We identified alternative splicing events in islets of diabetes-susceptible mice amongst others in genes linked to insulin secretion, endocytosis or Citation: Wilhelmi, I.; Neumann, A.; ubiquitin-mediated proteolysis pathways. The expression pattern of 54 histones and chromatin Jähnert, M.; Ouni, M.; Schürmann, A. -

The Emerging Role of the RBM20 and PTBP1 Ribonucleoproteins in Heart Development and Cardiovascular Diseases
G C A T T A C G G C A T genes Review The Emerging Role of the RBM20 and PTBP1 Ribonucleoproteins in Heart Development and Cardiovascular Diseases Stefania Fochi, Pamela Lorenzi, Marilisa Galasso, Chiara Stefani , Elisabetta Trabetti, Donato Zipeto and Maria Grazia Romanelli * Department of Neurosciences, Biomedicine and Movement Sciences, Section of Biology and Genetics, University of Verona, 37134 Verona, Italy; [email protected] (S.F.); [email protected] (P.L.); [email protected] (M.G.); [email protected] (C.S.); [email protected] (E.T.); [email protected] (D.Z.) * Correspondence: [email protected]; Tel.: +39-045-802-7182 Received: 9 March 2020; Accepted: 6 April 2020; Published: 8 April 2020 Abstract: Alternative splicing is a regulatory mechanism essential for cell differentiation and tissue organization. More than 90% of human genes are regulated by alternative splicing events, which participate in cell fate determination. The general mechanisms of splicing events are well known, whereas only recently have deep-sequencing, high throughput analyses and animal models provided novel information on the network of functionally coordinated, tissue-specific, alternatively spliced exons. Heart development and cardiac tissue differentiation require thoroughly regulated splicing events. The ribonucleoprotein RBM20 is a key regulator of the alternative splicing events required for functional and structural heart properties, such as the expression of TTN isoforms. Recently, the polypyrimidine tract-binding protein PTBP1 has been demonstrated to participate with RBM20 in regulating splicing events. In this review, we summarize the updated knowledge relative to RBM20 and PTBP1 structure and molecular function; their role in alternative splicing mechanisms involved in the heart development and function; RBM20 mutations associated with idiopathic dilated cardiovascular disease (DCM); and the consequences of RBM20-altered expression or dysfunction. -

Using Positional Distribution to Identify Splicing Elements and Predict Pre-Mrna Processing Defects in Human Genes
Using positional distribution to identify splicing elements and predict pre-mRNA processing defects in human genes Kian Huat Lima, Luciana Ferrarisa, Madeleine E. Fillouxa, Benjamin J. Raphaelb,c, and William G. Fairbrothera,c,d,1 aDepartment of Molecular and Cellular Biology and Biochemistry, Brown University, 70 Ship Street, Providence, RI 02903; bDepartment of Computer Science, Brown University, Providence, RI 02912; cCenter for Computational Molecular Biology, 151 Waterman Street, Providence, RI 02912; and dCenter for Genomics and Proteomics, Brown University, 70 Ship Street, Providence, RI 02903 Edited by Phillip A. Sharp, MIT, Cambridge, MA, and approved May 13, 2011 (received for review February 11, 2011) We present an intuitive strategy for predicting the effect of molecular intervention. As physical methods for the detection of sequence variation on splicing. In contrast to transcriptional ele- alternative splicing require large panels of genotyped accessible ments, splicing elements appear to be strongly position dependent. tissue, these studies will probably continue to be limited to sam- We demonstrated that exonic binding of the normally intronic spli- ples harvested from human blood. An alternate approach is the cing factor, U2AF65, inhibits splicing. Reasoning that the positional prediction of causative variations from single-nucleo polymorph- distribution of a splicing element is a signature of its function, we ism (SNPs) that fall within splicing elements. The key to this developed a method for organizing all possible sequence motifs approach is being able to identify what the splicing elements into clusters based on the genomic profile of their positional dis- are and whether a variation is disruptive. tribution around splice sites.