Chylomicron Retention Disease)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
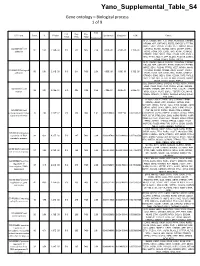
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

Cldn19 Clic2 Clmp Cln3
NewbornDx™ Advanced Sequencing Evaluation When time to diagnosis matters, the NewbornDx™ Advanced Sequencing Evaluation from Athena Diagnostics delivers rapid, 5- to 7-day results on a targeted 1,722-genes. A2ML1 ALAD ATM CAV1 CLDN19 CTNS DOCK7 ETFB FOXC2 GLUL HOXC13 JAK3 AAAS ALAS2 ATP1A2 CBL CLIC2 CTRC DOCK8 ETFDH FOXE1 GLYCTK HOXD13 JUP AARS2 ALDH18A1 ATP1A3 CBS CLMP CTSA DOK7 ETHE1 FOXE3 GM2A HPD KANK1 AASS ALDH1A2 ATP2B3 CC2D2A CLN3 CTSD DOLK EVC FOXF1 GMPPA HPGD K ANSL1 ABAT ALDH3A2 ATP5A1 CCDC103 CLN5 CTSK DPAGT1 EVC2 FOXG1 GMPPB HPRT1 KAT6B ABCA12 ALDH4A1 ATP5E CCDC114 CLN6 CUBN DPM1 EXOC4 FOXH1 GNA11 HPSE2 KCNA2 ABCA3 ALDH5A1 ATP6AP2 CCDC151 CLN8 CUL4B DPM2 EXOSC3 FOXI1 GNAI3 HRAS KCNB1 ABCA4 ALDH7A1 ATP6V0A2 CCDC22 CLP1 CUL7 DPM3 EXPH5 FOXL2 GNAO1 HSD17B10 KCND2 ABCB11 ALDOA ATP6V1B1 CCDC39 CLPB CXCR4 DPP6 EYA1 FOXP1 GNAS HSD17B4 KCNE1 ABCB4 ALDOB ATP7A CCDC40 CLPP CYB5R3 DPYD EZH2 FOXP2 GNE HSD3B2 KCNE2 ABCB6 ALG1 ATP8A2 CCDC65 CNNM2 CYC1 DPYS F10 FOXP3 GNMT HSD3B7 KCNH2 ABCB7 ALG11 ATP8B1 CCDC78 CNTN1 CYP11B1 DRC1 F11 FOXRED1 GNPAT HSPD1 KCNH5 ABCC2 ALG12 ATPAF2 CCDC8 CNTNAP1 CYP11B2 DSC2 F13A1 FRAS1 GNPTAB HSPG2 KCNJ10 ABCC8 ALG13 ATR CCDC88C CNTNAP2 CYP17A1 DSG1 F13B FREM1 GNPTG HUWE1 KCNJ11 ABCC9 ALG14 ATRX CCND2 COA5 CYP1B1 DSP F2 FREM2 GNS HYDIN KCNJ13 ABCD3 ALG2 AUH CCNO COG1 CYP24A1 DST F5 FRMD7 GORAB HYLS1 KCNJ2 ABCD4 ALG3 B3GALNT2 CCS COG4 CYP26C1 DSTYK F7 FTCD GP1BA IBA57 KCNJ5 ABHD5 ALG6 B3GAT3 CCT5 COG5 CYP27A1 DTNA F8 FTO GP1BB ICK KCNJ8 ACAD8 ALG8 B3GLCT CD151 COG6 CYP27B1 DUOX2 F9 FUCA1 GP6 ICOS KCNK3 ACAD9 ALG9 -

SAR1B Gene Secretion Associated Ras Related Gtpase 1B
SAR1B gene secretion associated Ras related GTPase 1B Normal Function The SAR1B gene provides instructions for making a protein that is produced in a variety of tissues. Most research involving the SAR1B protein has studied its role in the digestive tract. The SAR1B protein is needed for the transport of molecules called chylomicrons. During digestion, chylomicrons are formed within cells called enterocytes that line the small intestine and absorb nutrients. Chylomicrons are needed to absorb fat-soluble vitamins ( vitamins K, E, and D) and carry fats and cholesterol from the small intestine into the bloodstream. Within enterocytes, SAR1B proteins help transport immature chylomicrons from a cell structure called the endoplasmic reticulum to another cell structure called the Golgi apparatus. Immature chylomicrons are processed within the Golgi apparatus, resulting in mature chylomicrons. These mature chylomicrons are then released from enterocytes into the bloodstream so the body can use the fats and fat-soluble vitamins they carry. Sufficient levels of fats, cholesterol, and vitamins are necessary for normal growth and development. In other tissues, such as the heart and other muscles, the SAR1B protein is likely involved in transporting calcium within cells. Health Conditions Related to Genetic Changes Chylomicron retention disease More than 20 mutations in the SAR1B gene have been found to cause chylomicron retention disease. This is an inherited disorder that impairs the normal absorption of fats, cholesterol, and fat-soluble vitamins from food. Most of the mutations change one protein building block (amino acid) in the SAR1B protein. Other mutations lead to the production of an abnormally small version of the protein that cannot function properly. -

Molecular Analysis and Intestinal Expression of SAR1 Genes And
Georges et al. Orphanet Journal of Rare Diseases 2011, 6:1 http://www.ojrd.com/content/6/1/1 RESEARCH Open Access Molecular analysis and intestinal expression of SAR1 genes and proteins in Anderson’s disease (Chylomicron retention disease) Amandine Georges1, Jessica Bonneau2, Dominique Bonnefont-Rousselot3, Jacqueline Champigneulle4, Jean P Rabès2,8, Marianne Abifadel2, Thomas Aparicio5, Jean C Guenedet4,9, Eric Bruckert6, Catherine Boileau2,8, Alain Morali1, Mathilde Varret2, Lawrence P Aggerbeck7, Marie E Samson-Bouma2* Abstract Background: Anderson’s disease (AD) or chylomicron retention disease (CMRD) is a very rare hereditary lipid malabsorption syndrome. In order to discover novel mutations in the SAR1B gene and to evaluate the expression, as compared to healthy subjects, of the Sar1 gene and protein paralogues in the intestine, we investigated three previously undescribed individuals with the disease. Methods: The SAR1B, SAR1A and PCSK9 genes were sequenced. The expression of the SAR1B and SAR1A genes in intestinal biopsies of both normal individuals and patients was measured by RTqPCR. Immunohistochemistry using antibodies to recombinant Sar1 protein was used to evaluate the expression and localization of the Sar1 paralogues in the duodenal biopsies. Results: Two patients had a novel SAR1B mutation (p.Asp48ThrfsX17). The third patient, who had a previously described SAR1B mutation (p.Leu28ArgfsX7), also had a p.Leu21dup variant of the PCSK9 gene. The expression of the SAR1B gene in duodenal biopsies from an AD/CMRD patient was significantly decreased whereas the expression of the SAR1A gene was significantly increased, as compared to healthy individuals. The Sar1 proteins were present in decreased amounts in enterocytes in duodenal biopsies from the patients as compared to those from healthy subjects. -

How an Amino Acid Affects a Key Cell-Growth Regulator
News & views dynamics of actin — a structural protein of signalling compared with wild-type aged in the development of age-related mammalian the cell skeleton. Knockdown of eps-8 in adult worms is consistent with previous work show- disorders that involve protein aggregation, worms prevented the destabilization of actin ing that, in the mutant worms, deubiquitin- such as Parkinson’s disease and Alzheimer’s networks in muscles of the body wall and the ation is actively repressed and proteasome disease15. Therefore, understanding how the concomitant decrease in motility in old worms activity is increased8. Previous work also ubiquitination dynamics for different pro- (Fig. 1b). When eps-8 expression was reduced showed that the longevity of these animals teins change in different species with ageing specifically in the muscles or neuronal cells is, in part, supported by an increase in ubiq- could provide insight into potential strategies of adult worms, the worms lived longer than uitination9. for treating such disorders. normal. Concordantly, knockdown in adult- From a broader perspective, Koyuncu and hood of mig-2, one of the three C. elegans co-workers’ findings elegantly support the Bart P. Braeckman is in the Department of genes that encode RAC-like proteins, also antagonistic-pleiotropy hypothesis10 — an Biology, Ghent University, Ghent B-9000, extended lifespan. evolutionary explanation for why ageing is Belgium. In summary, Koyuncu et al. successfully link adaptive and therefore occurs. This hypoth- e-mail: [email protected] the age-related decrease in ubiquitination esis posits that genes that are beneficial for and subsequent increase in the abundance development or reproduction, but are detri- 1. -

Downregulation of Carnitine Acyl-Carnitine Translocase by Mirnas
Page 1 of 288 Diabetes 1 Downregulation of Carnitine acyl-carnitine translocase by miRNAs 132 and 212 amplifies glucose-stimulated insulin secretion Mufaddal S. Soni1, Mary E. Rabaglia1, Sushant Bhatnagar1, Jin Shang2, Olga Ilkayeva3, Randall Mynatt4, Yun-Ping Zhou2, Eric E. Schadt6, Nancy A.Thornberry2, Deborah M. Muoio5, Mark P. Keller1 and Alan D. Attie1 From the 1Department of Biochemistry, University of Wisconsin, Madison, Wisconsin; 2Department of Metabolic Disorders-Diabetes, Merck Research Laboratories, Rahway, New Jersey; 3Sarah W. Stedman Nutrition and Metabolism Center, Duke Institute of Molecular Physiology, 5Departments of Medicine and Pharmacology and Cancer Biology, Durham, North Carolina. 4Pennington Biomedical Research Center, Louisiana State University system, Baton Rouge, Louisiana; 6Institute for Genomics and Multiscale Biology, Mount Sinai School of Medicine, New York, New York. Corresponding author Alan D. Attie, 543A Biochemistry Addition, 433 Babcock Drive, Department of Biochemistry, University of Wisconsin-Madison, Madison, Wisconsin, (608) 262-1372 (Ph), (608) 263-9608 (fax), [email protected]. Running Title: Fatty acyl-carnitines enhance insulin secretion Abstract word count: 163 Main text Word count: 3960 Number of tables: 0 Number of figures: 5 Diabetes Publish Ahead of Print, published online June 26, 2014 Diabetes Page 2 of 288 2 ABSTRACT We previously demonstrated that micro-RNAs 132 and 212 are differentially upregulated in response to obesity in two mouse strains that differ in their susceptibility to obesity-induced diabetes. Here we show the overexpression of micro-RNAs 132 and 212 enhances insulin secretion (IS) in response to glucose and other secretagogues including non-fuel stimuli. We determined that carnitine acyl-carnitine translocase (CACT, Slc25a20) is a direct target of these miRNAs. -

Variations in Microrna-25 Expression Influence the Severity of Diabetic
BASIC RESEARCH www.jasn.org Variations in MicroRNA-25 Expression Influence the Severity of Diabetic Kidney Disease † † † Yunshuang Liu,* Hongzhi Li,* Jieting Liu,* Pengfei Han, Xuefeng Li, He Bai,* Chunlei Zhang,* Xuelian Sun,* Yanjie Teng,* Yufei Zhang,* Xiaohuan Yuan,* Yanhui Chu,* and Binghai Zhao* *Heilongjiang Key Laboratory of Anti-Fibrosis Biotherapy, Medical Research Center, Heilongjiang, People’s Republic of China; and †Clinical Laboratory of Hong Qi Hospital, Mudanjiang Medical University, Heilongjiang, People’s Republic of China ABSTRACT Diabetic nephropathy is characterized by persistent albuminuria, progressive decline in GFR, and second- ary hypertension. MicroRNAs are dysregulated in diabetic nephropathy, but identification of the specific microRNAs involved remains incomplete. Here, we show that the peripheral blood from patients with diabetes and the kidneys of animals with type 1 or 2 diabetes have low levels of microRNA-25 (miR-25) compared with those of their nondiabetic counterparts. Furthermore, treatment with high glucose decreased the expression of miR-25 in cultured kidney cells. In db/db mice, systemic administration of an miR-25 agomir repressed glomerular fibrosis and reduced high BP. Notably, knockdown of miR-25 in normal mice by systemic administration of an miR-25 antagomir resulted in increased proteinuria, extracellular matrix accumulation, podocyte foot process effacement, and hypertension with renin-angiotensin system activation. However, excessive miR-25 did not cause kidney dysfunction in wild-type mice. RNA sequencing showed the alteration of miR-25 target genes in antagomir-treated mice, including the Ras-related gene CDC42. In vitro,cotrans- fection with the miR-25 antagomir repressed luciferase activity from a reporter construct containing the CDC42 39 untranslated region. -

Perkinelmer Genomics to Request the Saliva Swab Collection Kit for Patients That Cannot Provide a Blood Sample As Whole Blood Is the Preferred Sample
Autism and Intellectual Disability TRIO Panel Test Code TR002 Test Summary This test analyzes 2429 genes that have been associated with Autism and Intellectual Disability and/or disorders associated with Autism and Intellectual Disability with the analysis being performed as a TRIO Turn-Around-Time (TAT)* 3 - 5 weeks Acceptable Sample Types Whole Blood (EDTA) (Preferred sample type) DNA, Isolated Dried Blood Spots Saliva Acceptable Billing Types Self (patient) Payment Institutional Billing Commercial Insurance Indications for Testing Comprehensive test for patients with intellectual disability or global developmental delays (Moeschler et al 2014 PMID: 25157020). Comprehensive test for individuals with multiple congenital anomalies (Miller et al. 2010 PMID 20466091). Patients with autism/autism spectrum disorders (ASDs). Suspected autosomal recessive condition due to close familial relations Previously negative karyotyping and/or chromosomal microarray results. Test Description This panel analyzes 2429 genes that have been associated with Autism and ID and/or disorders associated with Autism and ID. Both sequencing and deletion/duplication (CNV) analysis will be performed on the coding regions of all genes included (unless otherwise marked). All analysis is performed utilizing Next Generation Sequencing (NGS) technology. CNV analysis is designed to detect the majority of deletions and duplications of three exons or greater in size. Smaller CNV events may also be detected and reported, but additional follow-up testing is recommended if a smaller CNV is suspected. All variants are classified according to ACMG guidelines. Condition Description Autism Spectrum Disorder (ASD) refers to a group of developmental disabilities that are typically associated with challenges of varying severity in the areas of social interaction, communication, and repetitive/restricted behaviors. -
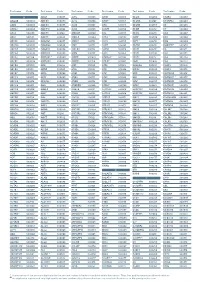
Aagab S00002 Aars S00003 Aars2 S00004 Aass S02483
Test name Code Test name Code Test name Code Test name Code Test name Code Test name Code A ADAR S00053 ALPL S00105 ARSB S00153 BCL10 S02266 C5AR2 S00263 AAGAB S00002 ADCK3 S00054 ALS2 S00106 ARSE * S00154 BCL11A S02167 C5ORF42 S00264 AARS S00003 ADCK4 S00055 ALX3 S00107 ARX S00155 BCL11B S02358 C6 S00265 AARS2 S00004 ADCY10 S02094 ALX4 S00108 ASAH1 S00156 BCOR S00212 C7 S00266 AASS S02483 ADCY3 S02184 AMACR S00109 ASL S00157 BCS1L S00213 C8A S00267 ABAT S02191 ADCY5 S02226 AMELX S02289 ASNS * S02508 BDNF S02509 C8B S00268 ABCA1 S00005 ADGRG1 S00057 AMER1 S00110 ASPA S00158 BDP1 * S00214 C8G S00269 ABCA12 S00006 ADGRG6 S02548 AMH S00111 ASPH S02425 BEAN1 S00215 C8ORF37 S00270 ABCA3 S00007 ADGRV1 S00058 AMHR2 S00112 ASPM S00159 BEST1 S00216 C9 S00271 ABCA4 S00008 ADIPOQ S00059 AMN S00113 ASS1 S00160 BFSP1 S02280 CA2 S00272 ABCA7 S02106 ADIPOR1 * S00060 AMPD1 S02670 ATAD3A * S02196 BFSP2 S00217 CA4 S02303 ABCB11 S00009 ADIPOR2 S00061 AMPD2 S02128 ATCAY S00162 BGN S02633 CA8 S00273 ABCB4 S00010 ADK S02595 AMT S00114 ATF6 S00163 BHLHA9 S00218 CABP2 S00274 ABCB6 S00011 ADNP S02320 ANG S00115 ATIC S02458 BICD2 S00220 CABP4 S00275 ABCB7 S00012 ADSL S00062 ANK1 S00116 ATL1 S00164 BIN1 S00221 CACNA1A S00276 ABCC2 S00013 AFF2 S00063 ANK2 S00117 ATL3 S00165 BLK S00222 CACNA1C * S00277 ABCC6 * S00014 AFG3L2 * S00064 ANKH S00118 ATM S00166 BLM S00223 CACNA1D S00278 ABCC8 S00015 AGA S00065 ANKRD11 * S02140 ATOH7 S02390 BLNK S02281 CACNA1F S00279 ABCC9 S00016 AGBL5 S02452 ANKS6 S00121 ATP13A2 S00168 BLOC1S3 S00224 CACNA1H S00280 ABCD1 * S00017 AGK * -

Supplementary Figure 1 Standardization of Gene Expression
Supplementary Figure 1 Standardization of gene expression Notes: (A) Standardization of GSE86544, (B) standardization of GSE103479, (C) standardization of GSE102238, (D) Standardization of GSE7055. The blue bar represents the data before normalization, and the red bar represents the data after normalization. Supplementary Figure 2 Correlation between module eigengenes and clinical traits especially PNI in GSE103479 and GSE102238 datasets. Notes: (A, B) Module-trait relationships in GSE103479 and GSE102238 datasets. The correlation coefficients and corresponding P-values in the brackets are contained in each cell. The table is color- coded by correlation between eigengenes and traits according to the color legend on the right side. The modules with the most significant differences are displayed in brackets. Abbreviations: PNI, perineural invasion. Supplementary Figure 3 The expression values of CCNB2 in pancreatic cancer (GSE102238) and colon cancer (GSE103479). Notes: (A, B) CCNB2 expression values were detected in GSE102238 and GSE103479. Abbreviations: CCNB2, cyclin B2 Supplementary Table 1 Results of top 20 pathway enrichment analysis of GSE7055 Term Category Description Count Log10(P) Genes GO:0000280 GO Biological nuclear division 33 -23.4 BIRC5,BUB1B,CCNB1,CCNE1,CDC20, Processes CKS2,KIF11,MAD2L1,MYBL2,SPAST, TOP2A,TTK,PRC1,PKMYT1,PTTG1,T RIP13,DLGAP5,TACC3,SMC2,SPAG5, UBE2C,ZWINT,TPX2,FBXO5,RACGA P1,NUSAP1,SPDL1,CDCA8,CEP55,ND C1,NSFL1C,KIF18B,ASPM GO:1902850 GO Biological microtubule 15 -12.89 BIRC5,CCNB1,CDC20,KIF11,MAD2L1 Processes -

Autocrine IFN Signaling Inducing Profibrotic Fibroblast Responses By
Downloaded from http://www.jimmunol.org/ by guest on September 23, 2021 Inducing is online at: average * The Journal of Immunology , 11 of which you can access for free at: 2013; 191:2956-2966; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication August 2013; doi: 10.4049/jimmunol.1300376 http://www.jimmunol.org/content/191/6/2956 A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Autocrine IFN Signaling Feng Fang, Kohtaro Ooka, Xiaoyong Sun, Ruchi Shah, Swati Bhattacharyya, Jun Wei and John Varga J Immunol cites 49 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130037 6.DC1 This article http://www.jimmunol.org/content/191/6/2956.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 23, 2021. The Journal of Immunology A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Inducing Autocrine IFN Signaling Feng Fang,* Kohtaro Ooka,* Xiaoyong Sun,† Ruchi Shah,* Swati Bhattacharyya,* Jun Wei,* and John Varga* Activation of TLR3 by exogenous microbial ligands or endogenous injury-associated ligands leads to production of type I IFN.