Alpha Actinin 4: an Intergral Component of Transcriptional
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

List of Genes Associated with Sudden Cardiac Death (Scdgseta) Gene
List of genes associated with sudden cardiac death (SCDgseta) mRNA expression in normal human heart Entrez_I Gene symbol Gene name Uniprot ID Uniprot name fromb D GTEx BioGPS SAGE c d e ATP-binding cassette subfamily B ABCB1 P08183 MDR1_HUMAN 5243 √ √ member 1 ATP-binding cassette subfamily C ABCC9 O60706 ABCC9_HUMAN 10060 √ √ member 9 ACE Angiotensin I–converting enzyme P12821 ACE_HUMAN 1636 √ √ ACE2 Angiotensin I–converting enzyme 2 Q9BYF1 ACE2_HUMAN 59272 √ √ Acetylcholinesterase (Cartwright ACHE P22303 ACES_HUMAN 43 √ √ blood group) ACTC1 Actin, alpha, cardiac muscle 1 P68032 ACTC_HUMAN 70 √ √ ACTN2 Actinin alpha 2 P35609 ACTN2_HUMAN 88 √ √ √ ACTN4 Actinin alpha 4 O43707 ACTN4_HUMAN 81 √ √ √ ADRA2B Adrenoceptor alpha 2B P18089 ADA2B_HUMAN 151 √ √ AGT Angiotensinogen P01019 ANGT_HUMAN 183 √ √ √ AGTR1 Angiotensin II receptor type 1 P30556 AGTR1_HUMAN 185 √ √ AGTR2 Angiotensin II receptor type 2 P50052 AGTR2_HUMAN 186 √ √ AKAP9 A-kinase anchoring protein 9 Q99996 AKAP9_HUMAN 10142 √ √ √ ANK2/ANKB/ANKYRI Ankyrin 2 Q01484 ANK2_HUMAN 287 √ √ √ N B ANKRD1 Ankyrin repeat domain 1 Q15327 ANKR1_HUMAN 27063 √ √ √ ANKRD9 Ankyrin repeat domain 9 Q96BM1 ANKR9_HUMAN 122416 √ √ ARHGAP24 Rho GTPase–activating protein 24 Q8N264 RHG24_HUMAN 83478 √ √ ATPase Na+/K+–transporting ATP1B1 P05026 AT1B1_HUMAN 481 √ √ √ subunit beta 1 ATPase sarcoplasmic/endoplasmic ATP2A2 P16615 AT2A2_HUMAN 488 √ √ √ reticulum Ca2+ transporting 2 AZIN1 Antizyme inhibitor 1 O14977 AZIN1_HUMAN 51582 √ √ √ UDP-GlcNAc: betaGal B3GNT7 beta-1,3-N-acetylglucosaminyltransfe Q8NFL0 -

Role of Nuclear Receptors in Central Nervous System Development and Associated Diseases
Role of Nuclear Receptors in Central Nervous System Development and Associated Diseases The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Olivares, Ana Maria, Oscar Andrés Moreno-Ramos, and Neena B. Haider. 2015. “Role of Nuclear Receptors in Central Nervous System Development and Associated Diseases.” Journal of Experimental Neuroscience 9 (Suppl 2): 93-121. doi:10.4137/JEN.S25480. http:// dx.doi.org/10.4137/JEN.S25480. Published Version doi:10.4137/JEN.S25480 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:27320246 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA Journal name: Journal of Experimental Neuroscience Journal type: Review Year: 2015 Volume: 9(S2) Role of Nuclear Receptors in Central Nervous System Running head verso: Olivares et al Development and Associated Diseases Running head recto: Nuclear receptors development and associated diseases Supplementary Issue: Molecular and Cellular Mechanisms of Neurodegeneration Ana Maria Olivares1, Oscar Andrés Moreno-Ramos2 and Neena B. Haider1 1Department of Ophthalmology, Schepens Eye Research Institute, Massachusetts Eye and Ear, Harvard Medical School, Boston, MA, USA. 2Departamento de Ciencias Biológicas, Facultad de Ciencias, Universidad de los Andes, Bogotá, Colombia. ABSTRACT: The nuclear hormone receptor (NHR) superfamily is composed of a wide range of receptors involved in a myriad of important biological processes, including development, growth, metabolism, and maintenance. -

Full-Text PDF (Final Published Version)
Alexander, S. P. H., Cidlowski, J. A., Kelly, E., Marrion, N. V., Peters, J. A., Faccenda, E., Harding, S. D., Pawson, A. J., Sharman, J. L., Southan, C., Davies, J. A., & CGTP Collaborators (2017). THE CONCISE GUIDE TO PHARMACOLOGY 2017/18: Nuclear hormone receptors. British Journal of Pharmacology, 174, S208-S224. https://doi.org/10.1111/bph.13880 Publisher's PDF, also known as Version of record License (if available): CC BY Link to published version (if available): 10.1111/bph.13880 Link to publication record in Explore Bristol Research PDF-document This is the final published version of the article (version of record). It first appeared online via Wiley at https://doi.org/10.1111/bph.13880 . Please refer to any applicable terms of use of the publisher. University of Bristol - Explore Bristol Research General rights This document is made available in accordance with publisher policies. Please cite only the published version using the reference above. Full terms of use are available: http://www.bristol.ac.uk/red/research-policy/pure/user-guides/ebr-terms/ S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2017/18: Nuclear hormone receptors. British Journal of Pharmacology (2017) 174, S208–S224 THE CONCISE GUIDE TO PHARMACOLOGY 2017/18: Nuclear hormone receptors Stephen PH Alexander1, John A Cidlowski2, Eamonn Kelly3, Neil V Marrion3, John A Peters4, Elena Faccenda5, Simon D Harding5,AdamJPawson5, Joanna L Sharman5, Christopher Southan5, Jamie A Davies5 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical -

Anti-Testicular Receptor 2 (TR2) (Rabbit) Antibody - 100-401-E45
Anti-Testicular Receptor 2 (TR2) (Rabbit) Antibody - 100-401-E45 Code: 100-401-E45 Size: 100 µL Product Description: Anti-Testicular Receptor 2 (TR2) (Rabbit) Antibody - 100-401-E45 Concentration: Titrated value sufficient to run approximately 10 mini blots. PhysicalState: Liquid Label Unconjugated Host Rabbit Gene Name NR2C1 Species Reactivity mouse, human, rat Storage Condition Store vial at -20° C prior to opening. This product is stable at 4° C as an undiluted liquid. For extended storage, aliquot contents and freeze at -20° C or below. Avoid cycles of freezing and thawing. Dilute only prior to immediate use. Synonyms Nuclear receptor subfamily 2 group C member 1, Orphan nuclear receptor TR2, mTR2 Application Note Anti-Testicular Receptor 2 (Rabbit) antibody is suitable for use in Western Blots. Anti-Testicular Receptor 2 antibodies are specific for the ~64 kDa TR2 protein in Western blots of testes and nuclear extracts from MEL cell lines. Researchers should determine optimal titers for applications that are not stated below. Background TR2 antibody detects Testicular receptor 2 (TR2), which is a member of the orphan nuclear receptor family. It is widely expressed at a low level throughout the adult testis. TR2 represses transcription and binds DNA directly interacting with HDAC3 and HDAC4 via DNA-binding domains. TR2 has also been implicated in regulation of estrogen receptor activity in mammary glands. In addition, TR2 has recently been shown to form a heterodimer with TR4 that can bind to the direct repeat 6 element of the hepatitis B virus (HBV) enhancer II region thus suppressing HBV gene expression. -

Alpha;-Actinin-4 Promotes Metastasis in Gastric Cancer
Laboratory Investigation (2017) 97, 1084–1094 © 2017 USCAP, Inc All rights reserved 0023-6837/17 α-Actinin-4 promotes metastasis in gastric cancer Xin Liu and Kent-Man Chu Metastasis increases the mortality rate of gastric cancer, which is the third leading cause of cancer-associated deaths worldwide. This study aims to identify the genes promoting metastasis of gastric cancer (GC). A human cell motility PCR array was used to analyze a pair of tumor and non-tumor tissue samples from a patient with stage IV GC (T3N3M1). Expression of the dysregulated genes was then evaluated in GC tissue samples (n = 10) and cell lines (n =6) via qPCR. Expression of α-actinin-4 (ACTN4) was validated in a larger sample size (n = 47) by qPCR, western blot and immunohistochemistry. Knockdown of ACTN4 with specific siRNAs was performed in GC cells, and adhesion assays, transwell invasion assays and migration assays were used to evaluate the function of these cells. Expression of potential targets of ACTN4 were then evaluated by qPCR. Thirty upregulated genes (greater than twofold) were revealed by the PCR array. We focused on ACTN4 because it was upregulated in 6 out of 10 pairs of tissue samples and 5 out of 6 GC cell lines. Further study indicated that ACTN4 was upregulated in 22/32 pairs of tissue samples at stage III & IV (P = 0.0069). Knockdown of ACTN4 in GC cells showed no significant effect on cell proliferation, but significantly increased cell-matrix adhesion, as well as reduced migration and invasion of AGS, MKN7 and NCI-N87 cells. -
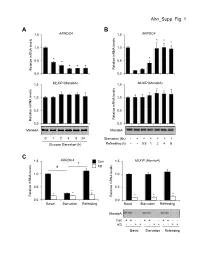
Ahn Supp. Fig. 1 AB 1.5 ARRDC4 1.5 ARRDC4 * * * 1.0 1.0
Ahn_Supp. Fig. 1 AB 1.5 ARRDC4 1.5 ARRDC4 * * * 1.0 1.0 * * 0.5 * 0.5 * * * Relative mRNA levels mRNA Relative Relative mRNA levels mRNA Relative 0.0 0.0 1.5 MLXIP (MondoA) 1.5 MLXIP (MondoA) 1.0 1.0 0.5 0.5 Relative mRNA levels mRNA Relative Relative mRNA levels mRNA Relative 0.0 0.0 MondoA MondoA 0124824 Starvation (6h) -++++++ Glucose Starvation (h) Refeeding (h) --0.51248 C 1.5 ARRDC4 1.5 MLXIP (MondoA) † Con # KD 1.0 1.0 0.5 0.5 * * * * Relative mRNA levels mRNA Relative Relative mRNA levels mRNA Relative * * 0.0 0.0 BasalStarvation Refeeding BasalStarvation Refeeding MondoA Con + + - - + + - - + + - - KD - - + + - - + + - - + + BasalStarvation Refeeding Supplemental Figure 1. Glucose-mediated regulation of ARRDC4 is dependent on MondoA in human skeletal myotubes. (A) (top) ARRDC4 and MLXIP (MondoA) mRNA levels were determined by qRT-PCR in human skeletal myotubes following deprivation of glucose at the indicated time (n=4). (bottom) Representative Western blot analysis of MondoA demonstrating the effect of glucose deprivation. *p<0.05 vs. 0h. (B) (top) ARRDC4 and MLXIP (MondoA) expression in human myotubes following a 6h glucose removal and refeeding at the times indicated (n=4). (bottom) Corresponding Western blot analysis. *p<0.05 vs Starvation 6h. (C) (top) Expression of ARRDC4 and MLXIP in human myotubes following deprivation and refeeding of glucose in the absence or presence of siRNA-mediated MondoA KD (n=4). (bottom) Corresponding Western blot analysis. *p<0.05 vs siControl. # p<0.05. § p<0.05. The data represents mean ± SD. All statistical significance determined by one-way ANOVA with Tukey multiple comparison post-hoc test. -

Proteomic Expression Profile in Human Temporomandibular Joint
diagnostics Article Proteomic Expression Profile in Human Temporomandibular Joint Dysfunction Andrea Duarte Doetzer 1,*, Roberto Hirochi Herai 1 , Marília Afonso Rabelo Buzalaf 2 and Paula Cristina Trevilatto 1 1 Graduate Program in Health Sciences, School of Medicine, Pontifícia Universidade Católica do Paraná (PUCPR), Curitiba 80215-901, Brazil; [email protected] (R.H.H.); [email protected] (P.C.T.) 2 Department of Biological Sciences, Bauru School of Dentistry, University of São Paulo, Bauru 17012-901, Brazil; [email protected] * Correspondence: [email protected]; Tel.: +55-41-991-864-747 Abstract: Temporomandibular joint dysfunction (TMD) is a multifactorial condition that impairs human’s health and quality of life. Its etiology is still a challenge due to its complex development and the great number of different conditions it comprises. One of the most common forms of TMD is anterior disc displacement without reduction (DDWoR) and other TMDs with distinct origins are condylar hyperplasia (CH) and mandibular dislocation (MD). Thus, the aim of this study is to identify the protein expression profile of synovial fluid and the temporomandibular joint disc of patients diagnosed with DDWoR, CH and MD. Synovial fluid and a fraction of the temporomandibular joint disc were collected from nine patients diagnosed with DDWoR (n = 3), CH (n = 4) and MD (n = 2). Samples were subjected to label-free nLC-MS/MS for proteomic data extraction, and then bioinformatics analysis were conducted for protein identification and functional annotation. The three Citation: Doetzer, A.D.; Herai, R.H.; TMD conditions showed different protein expression profiles, and novel proteins were identified Buzalaf, M.A.R.; Trevilatto, P.C. -

Alternative Splicing in the Nuclear Receptor Superfamily Expands Gene Function to Refine Endo-Xenobiotic Metabolism S
Supplemental material to this article can be found at: http://dmd.aspetjournals.org/content/suppl/2020/01/24/dmd.119.089102.DC1 1521-009X/48/4/272–287$35.00 https://doi.org/10.1124/dmd.119.089102 DRUG METABOLISM AND DISPOSITION Drug Metab Dispos 48:272–287, April 2020 Copyright ª 2020 by The American Society for Pharmacology and Experimental Therapeutics Minireview Alternative Splicing in the Nuclear Receptor Superfamily Expands Gene Function to Refine Endo-Xenobiotic Metabolism s Andrew J. Annalora, Craig B. Marcus, and Patrick L. Iversen Department of Environmental and Molecular Toxicology, Oregon State University, Corvallis, Oregon (A.J.A., C.B.M., P.L.I.) and United States Army Research Institute for Infectious Disease, Frederick, Maryland (P.L.I.) Received August 16, 2019; accepted December 31, 2019 ABSTRACT Downloaded from The human genome encodes 48 nuclear receptor (NR) genes, whose Exon inclusion options are differentially distributed across NR translated products transform chemical signals from endo- subfamilies, suggesting group-specific conservation of resilient func- xenobiotics into pleotropic RNA transcriptional profiles that refine tionalities. A deeper understanding of this transcriptional plasticity drug metabolism. This review describes the remarkable diversifica- expands our understanding of how chemical signals are refined and tion of the 48 human NR genes, which are potentially processed into mediated by NR genes. This expanded view of the NR transcriptome over 1000 distinct mRNA transcripts by alternative splicing (AS). The informs new models of chemical toxicity, disease diagnostics, and dmd.aspetjournals.org average human NR expresses ∼21 transcripts per gene and is precision-based approaches to personalized medicine. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

MAGE-A11 Is Activated Through TFCP2/ZEB1 Binding Sites De-Methylation As Well As Histone Modification and Facilitates ESCC Tumor Growth
www.impactjournals.com/oncotarget/ Oncotarget, 2018, Vol. 9, (No. 3), pp: 3365-3378 Research Paper MAGE-A11 is activated through TFCP2/ZEB1 binding sites de-methylation as well as histone modification and facilitates ESCC tumor growth Shina Liu1,*, Fei Liu1,*, Weina Huang1, Lina Gu1, Lingjiao Meng1, Yingchao Ju1,2, Yunyan Wu1, Juan Li1, Lihua Liu1 and Meixiang Sang1,3 1Research Center, the Fourth Hospital of Hebei Medical University, Shijiazhuang 050011, P. R. China 2Animal Center, the Fourth Hospital of Hebei Medical University, Shijiazhuang 050011, P. R. China 3Tumor Research Institute, the Fourth Hospital of Hebei Medical University, Shijiazhuang 050011, P. R. China *These authors contributed equally to this work Correspondence to: Meixiang Sang, email: [email protected] Keywords: MAGE-A11; ESCC; DNA methylation; histone acetylation; histone methylation Received: September 30, 2017 Accepted: November 15, 2017 Published: December 05, 2017 Copyright: Liu et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Recently, we have reported that the product of Melanoma Antigens Genes (MAGE) family member MAGE-A11 is an independent poor prognostic marker for esophageal squamous cell carcinoma (ESCC). However, the reason how MAGE-A11 is activated in ESCC progression still remains unclear. In the current study, we demonstrated that DNA methylation and the subsequent histone posttranslational modifications play crucial roles in the regulation of MAGE-A11 in ESCC progression. We found that the methylation rate of TFCP2/ZEB1 binding site on MAGE-A11 promoter in ESCC tissues and cells is higher than the normal esophageal epithelial tissues and cells. -

The Biological Role of Actinin-4 (ACTN4)
Honda Cell Biosci (2015) 5:41 DOI 10.1186/s13578-015-0031-0 REVIEW Open Access The biological role of actinin‑4 (ACTN4) in malignant phenotypes of cancer Kazufumi Honda1,2* Abstract Invasion and metastasis are malignant phenotypes in cancer that lead to patient death. Cell motility is involved in these processes. In 1998, we identified overexpression of the actin-bundling protein actinin-4 in several types of can- cer. Protein expression of actinin-4 is closely associated with the invasive phenotypes of cancers. Actinin-4 is predomi- nantly expressed in the cellular protrusions that stimulate the invasive phenotype in cancer cells and is essential for formation of cellular protrusions such as filopodia and lamellipodia. ACTN4 (gene name encoding actinin-4 protein) is located on human chromosome 19q. ACTN4 amplification is frequently observed in patients with carcinomas of the pancreas, ovary, lung, and salivary gland, and patients with ACTN4 amplifications have worse outcomes than patients without amplification. In addition, nuclear distribution of actinin-4 is frequently observed in small cell lung, breast, and ovarian cancer. Actinin-4, when expressed in cancer cell nuclei, functions as a transcriptional co-activator. In this review, we summarize recent developments regarding the biological roles of actinin-4 in cancer invasion. Keywords: Cancer invasion, Metastasis, Actinin-4 (ACTN4), Actin-bundling protein, Biomarker Background Alpha-actinin is an actin cross-linking protein that Despite successful complete resection at the primary belongs to the spectrin superfamily. Four isoforms of cancer site, poor outcomes are occasionally observed alpha-actinin have been identified: alpha-actinin-1 (gene in patients due to failure to control distant metastasis. -

Functions of Nuclear Actin‑Binding Proteins in Human Cancer (Review)
ONCOLOGY LETTERS 15: 2743-2748, 2018 Functions of nuclear actin‑binding proteins in human cancer (Review) XINYI YANG1,2 and YING LIN1,2 1Guangdong Provincial Key Laboratory of Malignant Tumor Epigenetics and Gene Regulation; 2Department of Gastroenterology and Hepatology, Sun Yat-sen Memorial Hospital, Sun Yat-sen University, Guangzhou, Guangdong 510120, P.R. China Received February 16, 2017; Accepted November 10, 2017 DOI: 10.3892/ol.2017.7658 Abstract. Nuclear actin-binding proteins (ABPs) perform that contain actin-binding domains that interact with actin. distinguishable functions compared with their cytoplasmic They can bind to actin monomers, actin polymers or both (2). counterparts in extensive activities of living cells. In addition Early studies have focused on the biological features and to the ability to regulate actin cytoskeleton dynamics, nuclear physiological mechanisms of cytoplasmic actin. Therefore, ABPs are associated with multiple nuclear biological processes, ABPs were considered to be distributed only in the cytoplasm including chromatin remodeling, gene transcriptional regula- and associated with the organization of actin cytoskeleton (3). tion, DNA damage response, nucleocytoplasmic trafficking In the cytoplasm, actin is associated with numerous cellular and nuclear structure maintenance. The nuclear translocation activities, including sustaining cellular morphology, deter- of ABPs is affected by numerous intracellular or extracellular mining cellular organelle distribution, mediating intracellular stimuli, which may lead to developmental malformation, transfer, endocytosis and exocytosis, cell division, cell migration tumor initiation, tumor progression and metastasis. Abnormal and adhesion (4-8). Meanwhile, ABPs regulate actin cytoskel- expression of certain ABPs have been reported in different etal structure by modulating actin filament cross‑linking into types of cancer.