Comprehensive Analysis of the Expression of Sodium/Potassium-Atpase Α Subunits and Prognosis of Ovarian Serous Cystadenocarcinoma
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Information to Mammadova-Bach Et Al., “Laminin Α1 Orchestrates VEGFA Functions in the Ecosystem of Colorectal Carcinogenesis”
Supplemental information to Mammadova-Bach et al., “Laminin α1 orchestrates VEGFA functions in the ecosystem of colorectal carcinogenesis” Supplemental material and methods Cloning of the villin-LMα1 vector The plasmid pBS-villin-promoter containing the 3.5 Kb of the murine villin promoter, the first non coding exon, 5.5 kb of the first intron and 15 nucleotides of the second villin exon, was generated by S. Robine (Institut Curie, Paris, France). The EcoRI site in the multi cloning site was destroyed by fill in ligation with T4 polymerase according to the manufacturer`s instructions (New England Biolabs, Ozyme, Saint Quentin en Yvelines, France). Site directed mutagenesis (GeneEditor in vitro Site-Directed Mutagenesis system, Promega, Charbonnières-les-Bains, France) was then used to introduce a BsiWI site before the start codon of the villin coding sequence using the 5’ phosphorylated primer: 5’CCTTCTCCTCTAGGCTCGCGTACGATGACGTCGGACTTGCGG3’. A double strand annealed oligonucleotide, 5’GGCCGGACGCGTGAATTCGTCGACGC3’ and 5’GGCCGCGTCGACGAATTCACGC GTCC3’ containing restriction site for MluI, EcoRI and SalI were inserted in the NotI site (present in the multi cloning site), generating the plasmid pBS-villin-promoter-MES. The SV40 polyA region of the pEGFP plasmid (Clontech, Ozyme, Saint Quentin Yvelines, France) was amplified by PCR using primers 5’GGCGCCTCTAGATCATAATCAGCCATA3’ and 5’GGCGCCCTTAAGATACATTGATGAGTT3’ before subcloning into the pGEMTeasy vector (Promega, Charbonnières-les-Bains, France). After EcoRI digestion, the SV40 polyA fragment was purified with the NucleoSpin Extract II kit (Machery-Nagel, Hoerdt, France) and then subcloned into the EcoRI site of the plasmid pBS-villin-promoter-MES. Site directed mutagenesis was used to introduce a BsiWI site (5’ phosphorylated AGCGCAGGGAGCGGCGGCCGTACGATGCGCGGCAGCGGCACG3’) before the initiation codon and a MluI site (5’ phosphorylated 1 CCCGGGCCTGAGCCCTAAACGCGTGCCAGCCTCTGCCCTTGG3’) after the stop codon in the full length cDNA coding for the mouse LMα1 in the pCIS vector (kindly provided by P. -

Sodium Pumps Mediate Activity-Dependent Changes in Mammalian Motor Networks
906 • The Journal of Neuroscience, January 25, 2017 • 37(4):906–921 Systems/Circuits Sodium Pumps Mediate Activity-Dependent Changes in Mammalian Motor Networks X Laurence D. Picton, XFilipe Nascimento, XMatthew J. Broadhead, XKeith T. Sillar, and XGareth B. Miles School of Psychology and Neuroscience, University of St Andrews, St Andrews KY16 9JP, United Kingdom Ubiquitously expressed sodium pumps are best known for maintaining the ionic gradients and resting membrane potential required for generating action potentials. However, activity- and state-dependent changes in pump activity can also influence neuronal firing and regulate rhythmic network output. Here we demonstrate that changes in sodium pump activity regulate locomotor networks in the spinal cord of neonatal mice. The sodium pump inhibitor, ouabain, increased the frequency and decreased the amplitude of drug-induced locomotor bursting, effects that were dependent on the presence of the neuromodulator dopamine. Conversely, activating the pump with the sodium ionophore monensin decreased burst frequency. When more “natural” locomotor output was evoked using dorsal-root stimulation, ouabain increased burst frequency and extended locomotor episode duration, whereas monensin slowed and shortened episodes. Decreasing the time between dorsal-root stimulation, and therefore interepisode interval, also shortened and slowed activity, suggesting that pump activity encodes information about past network output and contributes to feedforward control of subsequent locomotor bouts. Using whole-cell patch-clamp recordings from spinal motoneurons and interneurons, we describe a long-duration (ϳ60 s), activity-dependent, TTX- and ouabain-sensitive, hyperpolarization (ϳ5 mV), which is mediated by spike-dependent increases in pump activity. The duration of this dynamic pump potential is enhanced by dopamine. -

A SARS-Cov-2-Human Protein-Protein Interaction Map Reveals Drug Targets and Potential Drug-Repurposing
A SARS-CoV-2-Human Protein-Protein Interaction Map Reveals Drug Targets and Potential Drug-Repurposing Supplementary Information Supplementary Discussion All SARS-CoV-2 protein and gene functions described in the subnetwork appendices, including the text below and the text found in the individual bait subnetworks, are based on the functions of homologous genes from other coronavirus species. These are mainly from SARS-CoV and MERS-CoV, but when available and applicable other related viruses were used to provide insight into function. The SARS-CoV-2 proteins and genes listed here were designed and researched based on the gene alignments provided by Chan et. al. 1 2020 . Though we are reasonably sure the genes here are well annotated, we want to note that not every protein has been verified to be expressed or functional during SARS-CoV-2 infections, either in vitro or in vivo. In an effort to be as comprehensive and transparent as possible, we are reporting the sub-networks of these functionally unverified proteins along with the other SARS-CoV-2 proteins. In such cases, we have made notes within the text below, and on the corresponding subnetwork figures, and would advise that more caution be taken when examining these proteins and their molecular interactions. Due to practical limits in our sample preparation and data collection process, we were unable to generate data for proteins corresponding to Nsp3, Orf7b, and Nsp16. Therefore these three genes have been left out of the following literature review of the SARS-CoV-2 proteins and the protein-protein interactions (PPIs) identified in this study. -

Genetics of Migraine: Insights Into the Molecular Basis of Migraine Disorders
This may be the author’s version of a work that was submitted/accepted for publication in the following source: Sutherland, Heidi& Griffiths, Lyn (2017) Genetics of migraine: Insights into the molecular basis of migraine disor- ders. Headache, 57(4), pp. 537-569. This file was downloaded from: https://eprints.qut.edu.au/105633/ c Consult author(s) regarding copyright matters This work is covered by copyright. Unless the document is being made available under a Creative Commons Licence, you must assume that re-use is limited to personal use and that permission from the copyright owner must be obtained for all other uses. If the docu- ment is available under a Creative Commons License (or other specified license) then refer to the Licence for details of permitted re-use. It is a condition of access that users recog- nise and abide by the legal requirements associated with these rights. If you believe that this work infringes copyright please provide details by email to [email protected] Notice: Please note that this document may not be the Version of Record (i.e. published version) of the work. Author manuscript versions (as Sub- mitted for peer review or as Accepted for publication after peer review) can be identified by an absence of publisher branding and/or typeset appear- ance. If there is any doubt, please refer to the published source. https://doi.org/10.1111/head.13053 Genetics of Migraine: insights into the molecular basis of migraine disorders Heidi G. Sutherland, PhD and Lyn R. Griffiths, PhD Genomics Research Centre, Institute of Health and Biomedical Innovation, QUT, Musk Ave, Kelvin Grove, QLD 4059, Australia The authors declare no conflicts of interest. -

Transcriptomic Uniqueness and Commonality of the Ion Channels and Transporters in the Four Heart Chambers Sanda Iacobas1, Bogdan Amuzescu2 & Dumitru A
www.nature.com/scientificreports OPEN Transcriptomic uniqueness and commonality of the ion channels and transporters in the four heart chambers Sanda Iacobas1, Bogdan Amuzescu2 & Dumitru A. Iacobas3,4* Myocardium transcriptomes of left and right atria and ventricles from four adult male C57Bl/6j mice were profled with Agilent microarrays to identify the diferences responsible for the distinct functional roles of the four heart chambers. Female mice were not investigated owing to their transcriptome dependence on the estrous cycle phase. Out of the quantifed 16,886 unigenes, 15.76% on the left side and 16.5% on the right side exhibited diferential expression between the atrium and the ventricle, while 5.8% of genes were diferently expressed between the two atria and only 1.2% between the two ventricles. The study revealed also chamber diferences in gene expression control and coordination. We analyzed ion channels and transporters, and genes within the cardiac muscle contraction, oxidative phosphorylation, glycolysis/gluconeogenesis, calcium and adrenergic signaling pathways. Interestingly, while expression of Ank2 oscillates in phase with all 27 quantifed binding partners in the left ventricle, the percentage of in-phase oscillating partners of Ank2 is 15% and 37% in the left and right atria and 74% in the right ventricle. The analysis indicated high interventricular synchrony of the ion channels expressions and the substantially lower synchrony between the two atria and between the atrium and the ventricle from the same side. Starting with crocodilians, the heart pumps the blood through the pulmonary circulation and the systemic cir- culation by the coordinated rhythmic contractions of its upper lef and right atria (LA, RA) and lower lef and right ventricles (LV, RV). -

MALE Protein Name Accession Number Molecular Weight CP1 CP2 H1 H2 PDAC1 PDAC2 CP Mean H Mean PDAC Mean T-Test PDAC Vs. H T-Test
MALE t-test t-test Accession Molecular H PDAC PDAC vs. PDAC vs. Protein Name Number Weight CP1 CP2 H1 H2 PDAC1 PDAC2 CP Mean Mean Mean H CP PDAC/H PDAC/CP - 22 kDa protein IPI00219910 22 kDa 7 5 4 8 1 0 6 6 1 0.1126 0.0456 0.1 0.1 - Cold agglutinin FS-1 L-chain (Fragment) IPI00827773 12 kDa 32 39 34 26 53 57 36 30 55 0.0309 0.0388 1.8 1.5 - HRV Fab 027-VL (Fragment) IPI00827643 12 kDa 4 6 0 0 0 0 5 0 0 - 0.0574 - 0.0 - REV25-2 (Fragment) IPI00816794 15 kDa 8 12 5 7 8 9 10 6 8 0.2225 0.3844 1.3 0.8 A1BG Alpha-1B-glycoprotein precursor IPI00022895 54 kDa 115 109 106 112 111 100 112 109 105 0.6497 0.4138 1.0 0.9 A2M Alpha-2-macroglobulin precursor IPI00478003 163 kDa 62 63 86 72 14 18 63 79 16 0.0120 0.0019 0.2 0.3 ABCB1 Multidrug resistance protein 1 IPI00027481 141 kDa 41 46 23 26 52 64 43 25 58 0.0355 0.1660 2.4 1.3 ABHD14B Isoform 1 of Abhydrolase domain-containing proteinIPI00063827 14B 22 kDa 19 15 19 17 15 9 17 18 12 0.2502 0.3306 0.7 0.7 ABP1 Isoform 1 of Amiloride-sensitive amine oxidase [copper-containing]IPI00020982 precursor85 kDa 1 5 8 8 0 0 3 8 0 0.0001 0.2445 0.0 0.0 ACAN aggrecan isoform 2 precursor IPI00027377 250 kDa 38 30 17 28 34 24 34 22 29 0.4877 0.5109 1.3 0.8 ACE Isoform Somatic-1 of Angiotensin-converting enzyme, somaticIPI00437751 isoform precursor150 kDa 48 34 67 56 28 38 41 61 33 0.0600 0.4301 0.5 0.8 ACE2 Isoform 1 of Angiotensin-converting enzyme 2 precursorIPI00465187 92 kDa 11 16 20 30 4 5 13 25 5 0.0557 0.0847 0.2 0.4 ACO1 Cytoplasmic aconitate hydratase IPI00008485 98 kDa 2 2 0 0 0 0 2 0 0 - 0.0081 - 0.0 -

Anti-ATP1A4 / Na+ K+ Atpase Alpha 4 Antibody (ARG58367)
Product datasheet [email protected] ARG58367 Package: 100 μl anti-ATP1A4 / Na+ K+ ATPase alpha 4 antibody Store at: -20°C Summary Product Description Rabbit Polyclonal antibody recognizes ATP1A4 / Na+ K+ ATPase alpha 4 Tested Reactivity Ms, Rat Tested Application WB Host Rabbit Clonality Polyclonal Isotype IgG Target Name ATP1A4 / Na+ K+ ATPase alpha 4 Antigen Species Human Immunogen Recombinant fusion protein corresponding to aa. 1-90 of Human ATP1A4 (NP_653300.2). Conjugation Un-conjugated Alternate Names Sodium pump subunit alpha-4; Sodium/potassium-transporting ATPase subunit alpha-4; ATP1AL2; EC 3.6.3.9; ATP1A1; Na+ K+ ATPase alpha 4; Na K ATPase alpha 4; sodium potassium ATPase alpha 4; ATPase Na+ K+ alpha 4; ATPase Na K alpha 4; ATPase sodium potassium alpha 4 Application Instructions Application table Application Dilution WB 1:500 - 1:2000 Application Note * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Positive Control Mouse testis Calculated Mw 114 kDa Observed Size 100 kDa Properties Form Liquid Purification Affinity purified. Buffer PBS (pH 7.3), 0.02% Sodium azide and 50% Glycerol. Preservative 0.02% Sodium azide Stabilizer 50% Glycerol Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. www.arigobio.com 1/2 Note For laboratory research only, not for drug, diagnostic or other use. -

Supplementary Information for the Skeletal Muscle Molecular Clock Regulates Sarcomere Length Through Titin Splicing
Supplementary Information for The skeletal muscle molecular clock regulates sarcomere length through titin splicing Lance A. Riley, Xiping Zhang, Joseph M. Mijares, David W. Hammers, , Hailey R. Olafson, Ping Du, Collin M. Douglas, Siegfried Labeit, Eric T. Wang, and Karyn A. Esser Karyn A. Esser Email: [email protected] This PDF file includes: Tables S1 to S7 1 Table S1. Antibodies and concentrations used in this study. Antibody Source Concentration Rb anti-sarcomeric α-actinin (EP2529Y) Abcam 1:1000 Rb anti-titin N2A Myomedix 1:250 Rb anti-titin Z1Z2 Myomedix 1:100 Rb anti-RBM20 Myomedix 1:500 Ms anti-γ-tubulin (T6557) Sigma 1:1000 Rt anti-HA High Affinity (11867423001) Roche 1:1000 Gt anti-rabbit Alexa Fluor 488 Thermo 1:500 Gt anti-rabbit Alexa Fluor 405 Thermo 1:500 Gt anti-rabbit Alexa Fluor 647 Thermo 1:500 Gt anti-rabbit IgG (H+L) HRP Sigma 1:10000 Gt anti-mouse IgG (H+L) HRP Sigma 1:10000 2 Table S2. RNA Integrity Values and number of reads sequenced for each sample used for RNAseq analysis. Sample Name RIN Read Number (Millions) M10 8.6 61.4 M11 8.3 58.6 M13 8.4 77.5 M15 8.9 69.3 M16 8.8 45.6 M17 8.7 59.8 3 Table S3. Primers used for plasmid generation and qRT-PCR. Sequence (5’ – 3’) Name GGGAGATCTTTAACAACATAGGAGCTGTGATTGGCTGT U7-BglII F AAACTGCAGCACAACGCGTTTCCTAGGAAACCA U7-PstI R GCTCTTTTAGAATTTTTGGAGCAGGTTTTCTGAC SDM U7smOpt F GTCAGAAAACCTGCTGGTTAAATTCTAAAAGAGC SDM U7smOpt R TTAGGGTGGGTGGATACGCCTCTGCAAAAGAATTTTTGGAGCAGGTTTTCTG Ttn-51-AS F TATCCACCCACCCTAAGTCCCTATCATAGCGGAAGTGCGTCTGTAG Ttn-51-AS R TAGGGTGCAAGGTACTCCTTAGAGTGAAAGAATTTTTGGAGCAGGTTT Ttn-89-AS F GTACCTTGCACCCTAAGTCCCTATCATAGCGGAAGTGCGTCTGTAG Ttn-89-AS R TGCATGCCCAGAAATGCCTGCT Rbm20 qPCR F AAAGGCCCTCGTTGGAATGGCT Rbm20 qPCR R CGAGTCCAGCGAGAGAAGG Rpl26 F GCAGTCTTTAATGAAAGCCGTG Rpl26 R CTAGGACGGAATCTGCTGTG Rbm20 Intron 1 F AACAGGGTGTCTGTCTGTCT Rbm20 Intron 1 R 4 Table S4. -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942 -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -
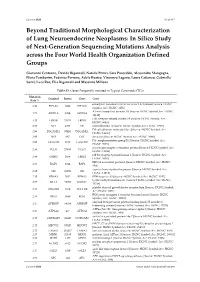
Beyond Traditional Morphological Characterization of Lung
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -

Atp1a4 (NM 013734) Mouse Tagged ORF Clone – MR220827L4 | Origene
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for MR220827L4 Atp1a4 (NM_013734) Mouse Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: Atp1a4 (NM_013734) Mouse Tagged ORF Clone Tag: mGFP Symbol: Atp1a4 Vector: pLenti-C-mGFP-P2A-Puro (PS100093) E. coli Selection: Chloramphenicol (34 ug/mL) Cell Selection: Puromycin ORF Nucleotide The ORF insert of this clone is exactly the same as(MR220827). Sequence: Restriction Sites: SgfI-MluI Cloning Scheme: ACCN: NM_013734 ORF Size: 3096 bp This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 Atp1a4 (NM_013734) Mouse Tagged ORF Clone – MR220827L4 OTI Disclaimer: Due to the inherent nature of this plasmid, standard methods to replicate additional amounts of DNA in E. coli are highly likely to result in mutations and/or rearrangements. Therefore, OriGene does not guarantee the capability to replicate this plasmid DNA. Additional amounts of DNA can be purchased from OriGene with batch-specific, full-sequence verification at a reduced cost. Please contact our customer care team at [email protected] or by calling 301.340.3188 option 3 for pricing and delivery. The molecular sequence of this clone aligns with the gene accession number as a point of reference only. However, individual transcript sequences of the same gene can differ through naturally occurring variations (e.g.