Using Core Genome Alignments to Assign Bacterial Species 2
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

(Batch Learning Self-Organizing Maps), to the Microbiome Analysis of Ticks
Title A novel approach, based on BLSOMs (Batch Learning Self-Organizing Maps), to the microbiome analysis of ticks Nakao, Ryo; Abe, Takashi; Nijhof, Ard M; Yamamoto, Seigo; Jongejan, Frans; Ikemura, Toshimichi; Sugimoto, Author(s) Chihiro The ISME Journal, 7(5), 1003-1015 Citation https://doi.org/10.1038/ismej.2012.171 Issue Date 2013-03 Doc URL http://hdl.handle.net/2115/53167 Type article (author version) File Information ISME_Nakao.pdf Instructions for use Hokkaido University Collection of Scholarly and Academic Papers : HUSCAP A novel approach, based on BLSOMs (Batch Learning Self-Organizing Maps), to the microbiome analysis of ticks Ryo Nakao1,a, Takashi Abe2,3,a, Ard M. Nijhof4, Seigo Yamamoto5, Frans Jongejan6,7, Toshimichi Ikemura2, Chihiro Sugimoto1 1Division of Collaboration and Education, Research Center for Zoonosis Control, Hokkaido University, Kita-20, Nishi-10, Kita-ku, Sapporo, Hokkaido 001-0020, Japan 2Nagahama Institute of Bio-Science and Technology, Nagahama, Shiga 526-0829, Japan 3Graduate School of Science & Technology, Niigata University, 8050, Igarashi 2-no-cho, Nishi- ku, Niigata 950-2181, Japan 4Institute for Parasitology and Tropical Veterinary Medicine, Freie Universität Berlin, Königsweg 67, 14163 Berlin, Germany 5Miyazaki Prefectural Institute for Public Health and Environment, 2-3-2 Gakuen Kibanadai Nishi, Miyazaki 889-2155, Japan 6Utrecht Centre for Tick-borne Diseases (UCTD), Department of Infectious Diseases and Immunology, Faculty of Veterinary Medicine, Utrecht University, Yalelaan 1, 3584 CL Utrecht, The Netherlands 7Department of Veterinary Tropical Diseases, Faculty of Veterinary Science, University of Pretoria, Private Bag X04, 0110 Onderstepoort, South Africa aThese authors contributed equally to this work. Keywords: BLSOMs/emerging diseases/metagenomics/microbiomes/symbionts/ticks Running title: Tick microbiomes revealed by BLSOMs Subject category: Microbe-microbe and microbe-host interactions Abstract Ticks transmit a variety of viral, bacterial and protozoal pathogens, which are often zoonotic. -

Babela Massiliensis, a Representative of a Widespread Bacterial
Babela massiliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae Isabelle Pagnier, Natalya Yutin, Olivier Croce, Kira S Makarova, Yuri I Wolf, Samia Benamar, Didier Raoult, Eugene V. Koonin, Bernard La Scola To cite this version: Isabelle Pagnier, Natalya Yutin, Olivier Croce, Kira S Makarova, Yuri I Wolf, et al.. Babela mas- siliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae. Biology Direct, BioMed Central, 2015, 10 (13), 10.1186/s13062-015-0043-z. hal-01217089 HAL Id: hal-01217089 https://hal-amu.archives-ouvertes.fr/hal-01217089 Submitted on 19 Oct 2015 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Pagnier et al. Biology Direct (2015) 10:13 DOI 10.1186/s13062-015-0043-z RESEARCH Open Access Babela massiliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae Isabelle Pagnier1, Natalya Yutin2, Olivier Croce1, Kira S Makarova2, Yuri I Wolf2, Samia Benamar1, Didier Raoult1, Eugene V Koonin2 and Bernard La Scola1* Abstract Background: Only a small fraction of bacteria and archaea that are identifiable by metagenomics can be grown on standard media. -

Characterization of the Interaction Between R. Conorii and Human
Louisiana State University LSU Digital Commons LSU Doctoral Dissertations Graduate School 4-5-2018 Characterization of the Interaction Between R. Conorii and Human Host Vitronectin in Rickettsial Pathogenesis Abigail Inez Fish Louisiana State University and Agricultural and Mechanical College, [email protected] Follow this and additional works at: https://digitalcommons.lsu.edu/gradschool_dissertations Part of the Bacteria Commons, Bacteriology Commons, Biology Commons, Immunology of Infectious Disease Commons, and the Pathogenic Microbiology Commons Recommended Citation Fish, Abigail Inez, "Characterization of the Interaction Between R. Conorii and Human Host Vitronectin in Rickettsial Pathogenesis" (2018). LSU Doctoral Dissertations. 4566. https://digitalcommons.lsu.edu/gradschool_dissertations/4566 This Dissertation is brought to you for free and open access by the Graduate School at LSU Digital Commons. It has been accepted for inclusion in LSU Doctoral Dissertations by an authorized graduate school editor of LSU Digital Commons. For more information, please [email protected]. CHARACTERIZATION OF THE INTERACTION BETWEEN R. CONORII AND HUMAN HOST VITRONECTIN IN RICKETTSIAL PATHOGENESIS A Dissertation Submitted to the Graduate Faculty of the Louisiana State University and Agricultural and Mechanical College in partial fulfillment of the requirements for the degree of Doctor of Philosophy in The Interdepartmental Program in Biomedical and Veterinary Medical Sciences Through the Department of Pathobiological Sciences by Abigail Inez -

“Candidatus Deianiraea Vastatrix” with the Ciliate Paramecium Suggests
bioRxiv preprint doi: https://doi.org/10.1101/479196; this version posted November 27, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The extracellular association of the bacterium “Candidatus Deianiraea vastatrix” with the ciliate Paramecium suggests an alternative scenario for the evolution of Rickettsiales 5 Castelli M.1, Sabaneyeva E.2, Lanzoni O.3, Lebedeva N.4, Floriano A.M.5, Gaiarsa S.5,6, Benken K.7, Modeo L. 3, Bandi C.1, Potekhin A.8, Sassera D.5*, Petroni G.3* 1. Centro Romeo ed Enrica Invernizzi Ricerca Pediatrica, Dipartimento di Bioscienze, Università 10 degli studi di Milano, Milan, Italy 2. Department of Cytology and Histology, Faculty of Biology, Saint Petersburg State University, Saint-Petersburg, Russia 3. Dipartimento di Biologia, Università di Pisa, Pisa, Italy 4 Centre of Core Facilities “Culture Collections of Microorganisms”, Saint Petersburg State 15 University, Saint Petersburg, Russia 5. Dipartimento di Biologia e Biotecnologie, Università degli studi di Pavia, Pavia, Italy 6. UOC Microbiologia e Virologia, Fondazione IRCCS Policlinico San Matteo, Pavia, Italy 7. Core Facility Center for Microscopy and Microanalysis, Saint Petersburg State University, Saint- Petersburg, Russia 20 8. Department of Microbiology, Faculty of Biology, Saint Petersburg State University, Saint- Petersburg, Russia * Corresponding authors, contacts: [email protected] ; [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/479196; this version posted November 27, 2018. -

Evolutionary Origin of Insect–Wolbachia Nutritional Mutualism
Evolutionary origin of insect–Wolbachia nutritional mutualism Naruo Nikoha,1, Takahiro Hosokawab,1, Minoru Moriyamab,1, Kenshiro Oshimac, Masahira Hattoric, and Takema Fukatsub,2 aDepartment of Liberal Arts, The Open University of Japan, Chiba 261-8586, Japan; bBioproduction Research Institute, National Institute of Advanced Industrial Science and Technology, Tsukuba 305-8566, Japan; and cCenter for Omics and Bioinformatics, Graduate School of Frontier Sciences, University of Tokyo, Kashiwa 277-8561, Japan Edited by Nancy A. Moran, University of Texas at Austin, Austin, TX, and approved June 3, 2014 (received for review May 20, 2014) Obligate insect–bacterium nutritional mutualism is among the insects, generally conferring negative fitness consequences to most sophisticated forms of symbiosis, wherein the host and the their hosts and often causing hosts’ reproductive aberrations to symbiont are integrated into a coherent biological entity and un- enhance their own transmission in a selfish manner (7, 8). Re- able to survive without the partnership. Originally, however, such cently, however, a Wolbachia strain associated with the bedbug obligate symbiotic bacteria must have been derived from free-living Cimex lectularius,designatedaswCle, was shown to be es- bacteria. How highly specialized obligate mutualisms have arisen sential for normal growth and reproduction of the blood- from less specialized associations is of interest. Here we address this sucking insect host via provisioning of B vitamins (9). Hence, it –Wolbachia evolutionary -

Ehrlichiosis and Anaplasmosis Are Tick-Borne Diseases Caused by Obligate Anaplasmosis: Intracellular Bacteria in the Genera Ehrlichia and Anaplasma
Ehrlichiosis and Importance Ehrlichiosis and anaplasmosis are tick-borne diseases caused by obligate Anaplasmosis: intracellular bacteria in the genera Ehrlichia and Anaplasma. These organisms are widespread in nature; the reservoir hosts include numerous wild animals, as well as Zoonotic Species some domesticated species. For many years, Ehrlichia and Anaplasma species have been known to cause illness in pets and livestock. The consequences of exposure vary Canine Monocytic Ehrlichiosis, from asymptomatic infections to severe, potentially fatal illness. Some organisms Canine Hemorrhagic Fever, have also been recognized as human pathogens since the 1980s and 1990s. Tropical Canine Pancytopenia, Etiology Tracker Dog Disease, Ehrlichiosis and anaplasmosis are caused by members of the genera Ehrlichia Canine Tick Typhus, and Anaplasma, respectively. Both genera contain small, pleomorphic, Gram negative, Nairobi Bleeding Disorder, obligate intracellular organisms, and belong to the family Anaplasmataceae, order Canine Granulocytic Ehrlichiosis, Rickettsiales. They are classified as α-proteobacteria. A number of Ehrlichia and Canine Granulocytic Anaplasmosis, Anaplasma species affect animals. A limited number of these organisms have also Equine Granulocytic Ehrlichiosis, been identified in people. Equine Granulocytic Anaplasmosis, Recent changes in taxonomy can make the nomenclature of the Anaplasmataceae Tick-borne Fever, and their diseases somewhat confusing. At one time, ehrlichiosis was a group of Pasture Fever, diseases caused by organisms that mostly replicated in membrane-bound cytoplasmic Human Monocytic Ehrlichiosis, vacuoles of leukocytes, and belonged to the genus Ehrlichia, tribe Ehrlichieae and Human Granulocytic Anaplasmosis, family Rickettsiaceae. The names of the diseases were often based on the host Human Granulocytic Ehrlichiosis, species, together with type of leukocyte most often infected. -

Fleas and Flea-Borne Diseases
International Journal of Infectious Diseases 14 (2010) e667–e676 Contents lists available at ScienceDirect International Journal of Infectious Diseases journal homepage: www.elsevier.com/locate/ijid Review Fleas and flea-borne diseases Idir Bitam a, Katharina Dittmar b, Philippe Parola a, Michael F. Whiting c, Didier Raoult a,* a Unite´ de Recherche en Maladies Infectieuses Tropicales Emergentes, CNRS-IRD UMR 6236, Faculte´ de Me´decine, Universite´ de la Me´diterrane´e, 27 Bd Jean Moulin, 13385 Marseille Cedex 5, France b Department of Biological Sciences, SUNY at Buffalo, Buffalo, NY, USA c Department of Biology, Brigham Young University, Provo, Utah, USA ARTICLE INFO SUMMARY Article history: Flea-borne infections are emerging or re-emerging throughout the world, and their incidence is on the Received 3 February 2009 rise. Furthermore, their distribution and that of their vectors is shifting and expanding. This publication Received in revised form 2 June 2009 reviews general flea biology and the distribution of the flea-borne diseases of public health importance Accepted 4 November 2009 throughout the world, their principal flea vectors, and the extent of their public health burden. Such an Corresponding Editor: William Cameron, overall review is necessary to understand the importance of this group of infections and the resources Ottawa, Canada that must be allocated to their control by public health authorities to ensure their timely diagnosis and treatment. Keywords: ß 2010 International Society for Infectious Diseases. Published by Elsevier Ltd. All rights reserved. Flea Siphonaptera Plague Yersinia pestis Rickettsia Bartonella Introduction to 16 families and 238 genera have been described, but only a minority is synanthropic, that is they live in close association with The past decades have seen a dramatic change in the geographic humans (Table 1).4,5 and host ranges of many vector-borne pathogens, and their diseases. -

Case Report: Coinfection with Rickettsia Monacensis and Orientia Tsutsugamushi
Am. J. Trop. Med. Hyg., 101(2), 2019, pp. 332–335 doi:10.4269/ajtmh.18-0631 Copyright © 2019 by The American Society of Tropical Medicine and Hygiene Case Report: Coinfection with Rickettsia monacensis and Orientia tsutsugamushi Seok Won Kim,1† Choon-Mee Kim,2† Dong-Min Kim,3* and Na Ra Yun3 1Department of Neurosurgery, College of Medicine, Chosun University, Gwangju, Republic of Korea; 2Premedical Science, College of Medicine, Chosun University, Gwangju, Republic of Korea; 3Department of Internal Medicine, College of Medicine, Chosun University, Gwangju, Republic of Korea Abstract. Rickettsia monacensis and Orientia tsutsugamushi are bacteria of the family Rickettsiaceae, which causes fever, rash, and eschar formation; outdoor activities are a risk factor for Rickettsiaceae infection. A 75-year-old woman presented with fever, rash, and eschar and was confirmed as being scrub typhus based on a nested-polymerase chain reaction (N-PCR) test for a 56-kDa gene of O. tsutsugamushi; the genome was identified as the Boryong genotype. In addition, a pan-Rickettsia real-time PCR test was positive and a N-PCR test using a Rickettsia-specific partial outer membrane protein A (rOmpA) confirmed R. monacensis. This is the first case wherein a patient suspected of having scrub typhus owing to the presence of rash and eschar was also found to be coinfected with O. tsutsugamushi and R. monacensis based on molecular testing. INTRODUCTION leukocyte count, 7,200/mm3; hemoglobin, 11.6 g/dL; platelet count, 232,000/mm3; and erythrocyte sedimentation rate, 31 Rickettsia monacensis is a pathogen that causes spotted mm/hours. C-reactive protein and procalcitonin levels were fever group rickettsial infection; the main symptoms of in- elevated at 9.26 mg/dL and 0.836 ng/mL (0–0.5 ng/mL), re- fection include fever, headache, and myalgia, as well as es- 1 spectively. -

Rickettsia Felis: Molecular Characterization of a New Member of the Spotted Fever Group
International Journal of Systematic and Evolutionary Microbiology (2001), 51, 339–347 Printed in Great Britain Rickettsia felis: molecular characterization of a new member of the spotted fever group Donald H. Bouyer,1 John Stenos,2 Patricia Crocquet-Valdes,1 Cecilia G. Moron,1 Vsevolod L. Popov,1 Jorge E. Zavala-Velazquez,3 Lane D. Foil,4 Diane R. Stothard,5 Abdu F. Azad6 and David H. Walker1 Author for correspondence: David H. Walker. Tel: j1 409 772 2856. Fax: j1 409 772 2500. e-mail: dwalker!utmb.edu 1 Department of Pathology, In this report, placement of Rickettsia felis in the spotted fever group (SFG) WHO Collaborating Center rather than the typhus group (TG) of Rickettsia is proposed. The organism, for Tropical Diseases, University of Texas Medical which was first observed in cat fleas (Ctenocephalides felis) by electron Branch, 301 University microscopy, has not yet been reported to have been cultivated reproducibly, Blvd, Galveston, TX thereby limiting the standard rickettsial typing by serological means. To 77555-0609, USA overcome this challenge, several genes were selected as targets to be utilized 2 Australian Rickettsial for the classification of R. felis. DNA from cat fleas naturally infected with R. Reference Laboratory, Douglas Hocking Medical felis was amplified by PCR utilizing primer sets specific for the 190 kDa surface Institute, Geelong antigen (rOmpA) and 17 kDa antigen genes. The entire 5513 bp rompA gene Hospital, Geelong, was sequenced, characterized and found to have several unique features when Australia compared to the rompA genes of other Rickettsia. Phylogenetic analysis of the 3 Department of Tropical partial sequence of the 17 kDa antigen gene indicated that R. -
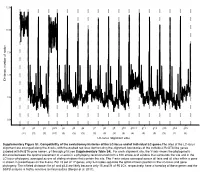
LC-Locus Alignment Sites Distance, Number of Nodes Supplementary
12.5 10.0 7.5 5.0 Distance, number of nodes 2.5 0.0 g1 g2 g3 g3.5 g4 g5 g6 g7 g8 g9 g10 g10.1 g11 g12 g13 g14 g15 (11) (3) (3) (10) (3) (3) (3) (3) (3) (3) (3) (4) (3) (3) (3) (2) (3) LC-locus alignment sites Supplementary Figure S1. Compatibility of the evolutionary histories of the LC-locus and of individual LC genes.The sites of the LC-locus alignment are arranged along the X-axis, with the dashed red lines demarcating the alignment boundaries of the individual RcGTA-like genes (labeled with RcGTA gene names, g1 through g15; see Supplementary Table S4). For each alignment site, the Y-axis shows the phylogenetic distance between the optimal placement of a taxon in a phylogeny reconstructed from a 100 amino-acid window that surrounds the site and in the LC-locus phylogeny, averaged across all sliding windows that contain the site. The Y-axis values averaged across all taxa and all sites within a gene is shown in parentheses on the X-axis. For 15 out of 17 genes, only 2-4 nodes separate the optimal taxon position in the LC-locus and gene phylogeny. The inflated distances for g1 and g3.5 are likely because only 15 and 21 of 95 LCs, respectively, have a homolog of these genes and the SSPB analysis is highly sensitive to missing data (Berger et al. 2011). a. Bacteria Unassigned Thermotogae Tenericutes Synergistetes Spirochaetes Proteobacteria ylum Planctomycetes h p Firmicutes Deferribacteres Cyanobacteria Chloroflexi Bacteroidetes Actinobacteria Acidobacteria 1(11,750) 2(1,750) 3(2,538) 4(168) 5(51) 6(54) 7(43) 8(32) 9(26) 10(40) 11(33) 12(198) 13(173) 14(101) 15(98) 16(43) 17(114) Number of rcc01682−rcc01698 homologs in a cluster b. -

Genome Project Reveals a Putative Rickettsial Endosymbiont
GBE Bacterial DNA Sifted from the Trichoplax adhaerens (Animalia: Placozoa) Genome Project Reveals a Putative Rickettsial Endosymbiont Timothy Driscoll1,y, Joseph J. Gillespie1,2,*,y, Eric K. Nordberg1,AbduF.Azad2, and Bruno W. Sobral1,3 1Virginia Bioinformatics Institute at Virginia Polytechnic Institute and State University 2Department of Microbiology and Immunology, University of Maryland School of Medicine 3Present address: Nestle´ Institute of Health Sciences SA, Campus EPFL, Quartier de L’innovation, Lausanne, Switzerland *Corresponding author: E-mail: [email protected]. yThese authors contributed equally to this work. Accepted: March 1, 2013 Abstract Eukaryotic genome sequencing projects often yield bacterial DNA sequences, data typically considered as microbial contamination. However, these sequences may also indicate either symbiont genes or lateral gene transfer (LGT) to host genomes. These bacterial sequences can provide clues about eukaryote–microbe interactions. Here, we used the genome of the primitive animal Trichoplax adhaerens (Metazoa: Placozoa), which is known to harbor an uncharacterized Gram-negative endosymbiont, to search for the presence of bacterial DNA sequences. Bioinformatic and phylogenomic analyses of extracted data from the genome assembly (181 bacterial coding sequences [CDS]) and trace read archive (16S rDNA) revealed a dominant proteobacterial profile strongly skewed to Rickettsiales (Alphaproteobacteria) genomes. By way of phylogenetic analysis of 16S rDNA and 113 proteins conserved across proteobacterial genomes, as well as identification of 27 rickettsial signature genes, we propose a Rickettsiales endosymbiont of T. adhaerens (RETA). The majority (93%) of the identified bacterial CDS belongs to small scaffolds containing prokaryotic-like genes; however, 12 CDS were identified on large scaffolds comprised of eukaryotic-like genes, suggesting that T. -

Ohio Department of Health, Bureau of Infectious Diseases Disease Name Class A, Requires Immediate Phone Call to Local Health
Ohio Department of Health, Bureau of Infectious Diseases Reporting specifics for select diseases reportable by ELR Class A, requires immediate phone Susceptibilities specimen type Reportable test name (can change if Disease Name other specifics+ call to local health required* specifics~ state/federal case definition or department reporting requirements change) Culture independent diagnostic tests' (CIDT), like BioFire panel or BD MAX, E. histolytica Stain specimen = stool, bile results should be sent as E. histolytica DNA fluid, duodenal fluid, 260373001^DETECTED^SCT with E. histolytica Antigen Amebiasis (Entamoeba histolytica) No No tissue large intestine, disease/organism-specific DNA LOINC E. histolytica Antibody tissue small intestine codes OR a generic CIDT-LOINC code E. histolytica IgM with organism-specific DNA SNOMED E. histolytica IgG codes E. histolytica Total Antibody Ova and Parasite Anthrax Antibody Anthrax Antigen Anthrax EITB Acute Anthrax EITB Convalescent Anthrax Yes No Culture ELISA PCR Stain/microscopy Stain/spore ID Eastern Equine Encephalitis virus Antibody Eastern Equine Encephalitis virus IgG Antibody Eastern Equine Encephalitis virus IgM Arboviral neuroinvasive and non- Eastern Equine Encephalitis virus RNA neuroinvasive disease: Eastern equine California serogroup virus Antibody encephalitis virus disease; LaCrosse Equivocal results are accepted for all California serogroup virus IgG Antibody virus disease (other California arborviral diseases; California serogroup virus IgM Antibody specimen = blood, serum, serogroup