Ohio Department of Health, Bureau of Infectious Diseases Disease Name Class A, Requires Immediate Phone Call to Local Health
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Distribution of Tick-Borne Diseases in China Xian-Bo Wu1, Ren-Hua Na2, Shan-Shan Wei2, Jin-Song Zhu3 and Hong-Juan Peng2*
Wu et al. Parasites & Vectors 2013, 6:119 http://www.parasitesandvectors.com/content/6/1/119 REVIEW Open Access Distribution of tick-borne diseases in China Xian-Bo Wu1, Ren-Hua Na2, Shan-Shan Wei2, Jin-Song Zhu3 and Hong-Juan Peng2* Abstract As an important contributor to vector-borne diseases in China, in recent years, tick-borne diseases have attracted much attention because of their increasing incidence and consequent significant harm to livestock and human health. The most commonly observed human tick-borne diseases in China include Lyme borreliosis (known as Lyme disease in China), tick-borne encephalitis (known as Forest encephalitis in China), Crimean-Congo hemorrhagic fever (known as Xinjiang hemorrhagic fever in China), Q-fever, tularemia and North-Asia tick-borne spotted fever. In recent years, some emerging tick-borne diseases, such as human monocytic ehrlichiosis, human granulocytic anaplasmosis, and a novel bunyavirus infection, have been reported frequently in China. Other tick-borne diseases that are not as frequently reported in China include Colorado fever, oriental spotted fever and piroplasmosis. Detailed information regarding the history, characteristics, and current epidemic status of these human tick-borne diseases in China will be reviewed in this paper. It is clear that greater efforts in government management and research are required for the prevention, control, diagnosis, and treatment of tick-borne diseases, as well as for the control of ticks, in order to decrease the tick-borne disease burden in China. Keywords: Ticks, Tick-borne diseases, Epidemic, China Review (Table 1) [2,4]. Continuous reports of emerging tick-borne Ticks can carry and transmit viruses, bacteria, rickettsia, disease cases in Shandong, Henan, Hebei, Anhui, and spirochetes, protozoans, Chlamydia, Mycoplasma,Bartonia other provinces demonstrate the rise of these diseases bodies, and nematodes [1,2]. -

Determination of the Effects That a Previously Uncharacterized Secreted Product from Klebsiella Pneumoniae Has on Citrobacter Fr
East Tennessee State University Digital Commons @ East Tennessee State University Undergraduate Honors Theses Student Works 5-2017 Determination of the effects that a previously uncharacterized secreted product from Klebsiella pneumoniae has on Citrobacter freundii and Enterobacter cloacae biofilms Cody M. Hastings Follow this and additional works at: https://dc.etsu.edu/honors Part of the Bacteria Commons, Bacteriology Commons, Biological Phenomena, Cell Phenomena, and Immunity Commons, Cell and Developmental Biology Commons, Medical Cell Biology Commons, Medical Microbiology Commons, Microbial Physiology Commons, and the Pathogenic Microbiology Commons Recommended Citation Hastings, Cody M., "Determination of the effects that a previously uncharacterized secreted product from Klebsiella pneumoniae has on Citrobacter freundii and Enterobacter cloacae biofilms" (2017). Undergraduate Honors Theses. Paper 419. https://dc.etsu.edu/ honors/419 This Honors Thesis - Withheld is brought to you for free and open access by the Student Works at Digital Commons @ East Tennessee State University. It has been accepted for inclusion in Undergraduate Honors Theses by an authorized administrator of Digital Commons @ East Tennessee State University. For more information, please contact [email protected]. Determination of the effects that a previously uncharacterized secreted product from Klebsiella pneumoniae has on Citrobacter freundii and Enterobacter cloacae biofilms By Cody Hastings An Undergraduate Thesis Submitted in Partial Fulfillment of the Requirements -

(Batch Learning Self-Organizing Maps), to the Microbiome Analysis of Ticks
Title A novel approach, based on BLSOMs (Batch Learning Self-Organizing Maps), to the microbiome analysis of ticks Nakao, Ryo; Abe, Takashi; Nijhof, Ard M; Yamamoto, Seigo; Jongejan, Frans; Ikemura, Toshimichi; Sugimoto, Author(s) Chihiro The ISME Journal, 7(5), 1003-1015 Citation https://doi.org/10.1038/ismej.2012.171 Issue Date 2013-03 Doc URL http://hdl.handle.net/2115/53167 Type article (author version) File Information ISME_Nakao.pdf Instructions for use Hokkaido University Collection of Scholarly and Academic Papers : HUSCAP A novel approach, based on BLSOMs (Batch Learning Self-Organizing Maps), to the microbiome analysis of ticks Ryo Nakao1,a, Takashi Abe2,3,a, Ard M. Nijhof4, Seigo Yamamoto5, Frans Jongejan6,7, Toshimichi Ikemura2, Chihiro Sugimoto1 1Division of Collaboration and Education, Research Center for Zoonosis Control, Hokkaido University, Kita-20, Nishi-10, Kita-ku, Sapporo, Hokkaido 001-0020, Japan 2Nagahama Institute of Bio-Science and Technology, Nagahama, Shiga 526-0829, Japan 3Graduate School of Science & Technology, Niigata University, 8050, Igarashi 2-no-cho, Nishi- ku, Niigata 950-2181, Japan 4Institute for Parasitology and Tropical Veterinary Medicine, Freie Universität Berlin, Königsweg 67, 14163 Berlin, Germany 5Miyazaki Prefectural Institute for Public Health and Environment, 2-3-2 Gakuen Kibanadai Nishi, Miyazaki 889-2155, Japan 6Utrecht Centre for Tick-borne Diseases (UCTD), Department of Infectious Diseases and Immunology, Faculty of Veterinary Medicine, Utrecht University, Yalelaan 1, 3584 CL Utrecht, The Netherlands 7Department of Veterinary Tropical Diseases, Faculty of Veterinary Science, University of Pretoria, Private Bag X04, 0110 Onderstepoort, South Africa aThese authors contributed equally to this work. Keywords: BLSOMs/emerging diseases/metagenomics/microbiomes/symbionts/ticks Running title: Tick microbiomes revealed by BLSOMs Subject category: Microbe-microbe and microbe-host interactions Abstract Ticks transmit a variety of viral, bacterial and protozoal pathogens, which are often zoonotic. -

Babela Massiliensis, a Representative of a Widespread Bacterial
Babela massiliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae Isabelle Pagnier, Natalya Yutin, Olivier Croce, Kira S Makarova, Yuri I Wolf, Samia Benamar, Didier Raoult, Eugene V. Koonin, Bernard La Scola To cite this version: Isabelle Pagnier, Natalya Yutin, Olivier Croce, Kira S Makarova, Yuri I Wolf, et al.. Babela mas- siliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae. Biology Direct, BioMed Central, 2015, 10 (13), 10.1186/s13062-015-0043-z. hal-01217089 HAL Id: hal-01217089 https://hal-amu.archives-ouvertes.fr/hal-01217089 Submitted on 19 Oct 2015 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Pagnier et al. Biology Direct (2015) 10:13 DOI 10.1186/s13062-015-0043-z RESEARCH Open Access Babela massiliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae Isabelle Pagnier1, Natalya Yutin2, Olivier Croce1, Kira S Makarova2, Yuri I Wolf2, Samia Benamar1, Didier Raoult1, Eugene V Koonin2 and Bernard La Scola1* Abstract Background: Only a small fraction of bacteria and archaea that are identifiable by metagenomics can be grown on standard media. -

Transition of Serum Cytokine Concentration in Rickettsia Japonica Infection
Article Transition of Serum Cytokine Concentration in Rickettsia japonica Infection Makoto Kondo, Yoshiaki Matsushima, Kento Mizutani, Shohei Iida, Koji Habe and Keiichi Yamanaka * Department of Dermatology, Mie University Graduate School of medicine, 2-174 Edobashi, Tsu, Mie 514-8507, Japan; [email protected] (M.K.); [email protected] (Y.M.); [email protected] (K.M.); [email protected] (S.I.); [email protected] (K.H.) * Correspondence: [email protected]; Tel.: +81-59-231-5025; Fax: +81-59-231-5206 Received: 31 October 2020; Accepted: 8 December 2020; Published: 11 December 2020 Abstract: (1) Background. Rickettsia japonica (R. japonica) infection induces severe inflammation, and the disappearance of eosinophil in the acute stage is one of the phenomena. (2) Materials and Methods. In the current study, we measured the serum concentrations of Th1, Th2, and Th17 cytokines in the acute and recovery stages. (3) Results. In the acute phase, IL-6 and IFN-γ levels were elevated and we speculated that they played a role as a defense mechanism against R. japonica. The high concentration of IFN-γ suppressed the differentiation of eosinophil and induced apoptosis of eosinophil, leading to the disappearance of eosinophil. On day 7, IL-6 and IFN-γ concentrations were decreased, and Th2 cytokines such as IL-5 and IL-9 were slightly increased. On day 14, eosinophil count recovered to the normal level. The transition of serum cytokine concentration in R. japonica infection was presented. (4) Conclusions. -
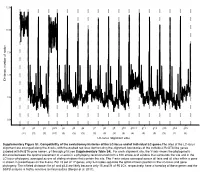
LC-Locus Alignment Sites Distance, Number of Nodes Supplementary
12.5 10.0 7.5 5.0 Distance, number of nodes 2.5 0.0 g1 g2 g3 g3.5 g4 g5 g6 g7 g8 g9 g10 g10.1 g11 g12 g13 g14 g15 (11) (3) (3) (10) (3) (3) (3) (3) (3) (3) (3) (4) (3) (3) (3) (2) (3) LC-locus alignment sites Supplementary Figure S1. Compatibility of the evolutionary histories of the LC-locus and of individual LC genes.The sites of the LC-locus alignment are arranged along the X-axis, with the dashed red lines demarcating the alignment boundaries of the individual RcGTA-like genes (labeled with RcGTA gene names, g1 through g15; see Supplementary Table S4). For each alignment site, the Y-axis shows the phylogenetic distance between the optimal placement of a taxon in a phylogeny reconstructed from a 100 amino-acid window that surrounds the site and in the LC-locus phylogeny, averaged across all sliding windows that contain the site. The Y-axis values averaged across all taxa and all sites within a gene is shown in parentheses on the X-axis. For 15 out of 17 genes, only 2-4 nodes separate the optimal taxon position in the LC-locus and gene phylogeny. The inflated distances for g1 and g3.5 are likely because only 15 and 21 of 95 LCs, respectively, have a homolog of these genes and the SSPB analysis is highly sensitive to missing data (Berger et al. 2011). a. Bacteria Unassigned Thermotogae Tenericutes Synergistetes Spirochaetes Proteobacteria ylum Planctomycetes h p Firmicutes Deferribacteres Cyanobacteria Chloroflexi Bacteroidetes Actinobacteria Acidobacteria 1(11,750) 2(1,750) 3(2,538) 4(168) 5(51) 6(54) 7(43) 8(32) 9(26) 10(40) 11(33) 12(198) 13(173) 14(101) 15(98) 16(43) 17(114) Number of rcc01682−rcc01698 homologs in a cluster b. -

(12) United States Patent (10) Patent No.: US 9,018,158 B2 Onsoyen Et Al
US0090181.58B2 (12) United States Patent (10) Patent No.: US 9,018,158 B2 Onsoyen et al. (45) Date of Patent: Apr. 28, 2015 (54) ALGINATE OLIGOMERS FOR USE IN 7,208,141 B2 * 4/2007 Montgomery .................. 424/45 OVERCOMING MULTIDRUG RESISTANCE 22:49 R: R388 al al W . aSOC ea. N BACTERA 7,671,102 B2 3/2010 Gaserod et al. 7,674,837 B2 3, 2010 G d et al. (75) Inventors: Edvar Onsoyen, Sandvika (NO); Rolf 7,758,856 B2 T/2010 it. Myrvold, Sandvika (NO); Arne Dessen, 7,776,839 B2 8/2010 Del Buono et al. Sandvika (NO); David Thomas, Cardiff 2006.8 R 38 8. Melist al. (GB); Timothy Rutland Walsh, Cardiff 2003/0022863 A1 1/2003 Stahlang et al. (GB) 2003/0224070 Al 12/2003 Sweazy et al. 2004/OO73964 A1 4/2004 Ellington et al. (73) Assignee: Algipharma AS, Sandvika (NO) 2004/0224922 A1 1 1/2004 King 2010.0068290 A1 3/2010 Ziegler et al. (*) Notice: Subject to any disclaimer, the term of this 2010/0305062 A1* 12/2010 Onsoyen et al. ................ 514/54 patent is extended or adjusted under 35 U.S.C. 154(b) by 184 days. FOREIGN PATENT DOCUMENTS DE 268865 A1 1, 1987 (21) Appl. No.: 13/376,164 EP O324720 A1 T, 1989 EP O 506,326 A2 9, 1992 (22) PCT Filed: Jun. 3, 2010 EP O590746 A1 4f1994 EP 1234584 A1 8, 2002 (86). PCT No.: PCT/GB2O1 O/OO1097 EP 1714660 A1 10, 2006 EP 1745705 A1 1, 2007 S371 (c)(1), FR T576 M 3/1968 (2), (4) Date: Jan. -

Genome Project Reveals a Putative Rickettsial Endosymbiont
GBE Bacterial DNA Sifted from the Trichoplax adhaerens (Animalia: Placozoa) Genome Project Reveals a Putative Rickettsial Endosymbiont Timothy Driscoll1,y, Joseph J. Gillespie1,2,*,y, Eric K. Nordberg1,AbduF.Azad2, and Bruno W. Sobral1,3 1Virginia Bioinformatics Institute at Virginia Polytechnic Institute and State University 2Department of Microbiology and Immunology, University of Maryland School of Medicine 3Present address: Nestle´ Institute of Health Sciences SA, Campus EPFL, Quartier de L’innovation, Lausanne, Switzerland *Corresponding author: E-mail: [email protected]. yThese authors contributed equally to this work. Accepted: March 1, 2013 Abstract Eukaryotic genome sequencing projects often yield bacterial DNA sequences, data typically considered as microbial contamination. However, these sequences may also indicate either symbiont genes or lateral gene transfer (LGT) to host genomes. These bacterial sequences can provide clues about eukaryote–microbe interactions. Here, we used the genome of the primitive animal Trichoplax adhaerens (Metazoa: Placozoa), which is known to harbor an uncharacterized Gram-negative endosymbiont, to search for the presence of bacterial DNA sequences. Bioinformatic and phylogenomic analyses of extracted data from the genome assembly (181 bacterial coding sequences [CDS]) and trace read archive (16S rDNA) revealed a dominant proteobacterial profile strongly skewed to Rickettsiales (Alphaproteobacteria) genomes. By way of phylogenetic analysis of 16S rDNA and 113 proteins conserved across proteobacterial genomes, as well as identification of 27 rickettsial signature genes, we propose a Rickettsiales endosymbiont of T. adhaerens (RETA). The majority (93%) of the identified bacterial CDS belongs to small scaffolds containing prokaryotic-like genes; however, 12 CDS were identified on large scaffolds comprised of eukaryotic-like genes, suggesting that T. -

Klebsiella Ornithinolytica
international Journal of Systematic Bacteriology (1 999), 49, 1695-1 700 Printed in Great Britain Phylogenetic evidence for reclassification of Calymmatobacterium granulomatis as Klebsiella granulomatis comb. nov. Jenny 5. Carter,’l2 Francis J. B~wden,~Ivan Ba~tian,~Garry M. Myers,’ K. S. Sriprakash’ and David J. Kemp’ Author for correspondence : David J. Kemp. Tel : + 6 18 8922 84 12. Fax : + 6 18 8927 5 187 e-mail : [email protected] 1 Menzies School of Health By sequencing a total of 2089 bp of the 16s rRNA and phoE genes it was Research, Darwin, demonstratedthat Calymmatobacterium grandomatis (the causative Austra Iia organism of donovanosis) shows a high level of identity with Klebsiella * Centre for Indigenous species pathogenic to humans (Klebsiellapneumoniae, Klebsiella Natural and Cultural Resource Management, rhinoscleromatis). It is proposed that C. grandomatis should be reclassified as Faculty of Aboriginal and Klebsiella granulomatis comb. nov. An emended description of the genus Torres Strait Islander Klebsiella is given. Studies, Northern Territory University, Darwin, Australia 3 Institute of Medical and Keywords : Calymmatobacteriurn, Klebsiella, sequence data, phylogenetic inferences Veterinary Science, Adelaide, Australia 4 AIDS/STD Unit, Royal Darwin Hospital, Darwin, Australia Calymmatobacterium granulomatis is the presumed ganism (Richens, 1991) have prevented further char- causative agent of donovanosis, an important cause of acterization of this relationship. genital ulceration that occurs in small endemic foci in all continents except Europe and Antarctica. The name Non-cultivable pathogenic eubacteria have been C. granulomatis was originally given to the pleo- identified by PCR using primers targeting conserved morphic bacterium cultured from donovanosis lesions genes (Fredricks & Relman, 1996). -

Expansion of Tick-Borne Rickettsioses in the World
microorganisms Review Expansion of Tick-Borne Rickettsioses in the World Mariusz Piotrowski * and Anna Rymaszewska Institute of Biology, University of Szczecin, 70-453 Szczecin, Poland; [email protected] * Correspondence: [email protected] Received: 24 September 2020; Accepted: 25 November 2020; Published: 30 November 2020 Abstract: Tick-borne rickettsioses are caused by obligate intracellular bacteria belonging to the spotted fever group of the genus Rickettsia. These infections are among the oldest known diseases transmitted by vectors. In the last three decades there has been a rapid increase in the recognition of this disease complex. This unusual expansion of information was mainly caused by the development of molecular diagnostic techniques that have facilitated the identification of new and previously recognized rickettsiae. A lot of currently known bacteria of the genus Rickettsia have been considered nonpathogenic for years, and moreover, many new species have been identified with unknown pathogenicity. The genus Rickettsia is distributed all over the world. Many Rickettsia species are present on several continents. The geographical distribution of rickettsiae is related to their vectors. New cases of rickettsioses and new locations, where the presence of these bacteria is recognized, are still being identified. The variety and rapid evolution of the distribution and density of ticks and diseases which they transmit shows us the scale of the problem. This review article presents a comparison of the current understanding of the geographic distribution of pathogenic Rickettsia species to that of the beginning of the century. Keywords: Tick-borne rickettsioses; Tick-borne diseases; Rickettsiales 1. Introduction Tick-borne rickettsioses are caused by obligate intracellular Gram-negative bacteria belonging to the spotted fever group (SFG) of the genus Rickettsia. -

Gene Gain and Loss Events in Rickettsia and Orientia Species Kalliopi Georgiades1,2, Vicky Merhej1, Khalid El Karkouri1, Didier Raoult1, Pierre Pontarotti2*
Georgiades et al. Biology Direct 2011, 6:6 http://www.biology-direct.com/content/6/1/6 RESEARCH Open Access Gene gain and loss events in Rickettsia and Orientia species Kalliopi Georgiades1,2, Vicky Merhej1, Khalid El Karkouri1, Didier Raoult1, Pierre Pontarotti2* Abstract Background: Genome degradation is an ongoing process in all members of the Rickettsiales order, which makes these bacterial species an excellent model for studying reductive evolution through interspecies variation in genome size and gene content. In this study, we evaluated the degree to which gene loss shaped the content of some Rickettsiales genomes. We shed light on the role played by horizontal gene transfers in the genome evolution of Rickettsiales. Results: Our phylogenomic tree, based on whole-genome content, presented a topology distinct from that of the whole core gene concatenated phylogenetic tree, suggesting that the gene repertoires involved have different evolutionary histories. Indeed, we present evidence for 3 possible horizontal gene transfer events from various organisms to Orientia and 6 to Rickettsia spp., while we also identified 3 possible horizontal gene transfer events from Rickettsia and Orientia to other bacteria. We found 17 putative genes in Rickettsia spp. that are probably the result of de novo gene creation; 2 of these genes appear to be functional. On the basis of these results, we were able to reconstruct the gene repertoires of “proto-Rickettsiales” and “proto-Rickettsiaceae”, which correspond to the ancestors of Rickettsiales and Rickettsiaceae, respectively. Finally, we found that 2,135 genes were lost during the evolution of the Rickettsiaceae to an intracellular lifestyle. Conclusions: Our phylogenetic analysis allowed us to track the gene gain and loss events occurring in bacterial genomes during their evolution from a free-living to an intracellular lifestyle. -

Use of the Diagnostic Bacteriology Laboratory: a Practical Review for the Clinician
148 Postgrad Med J 2001;77:148–156 REVIEWS Postgrad Med J: first published as 10.1136/pmj.77.905.148 on 1 March 2001. Downloaded from Use of the diagnostic bacteriology laboratory: a practical review for the clinician W J Steinbach, A K Shetty Lucile Salter Packard Children’s Hospital at EVective utilisation and understanding of the Stanford, Stanford Box 1: Gram stain technique University School of clinical bacteriology laboratory can greatly aid Medicine, 725 Welch in the diagnosis of infectious diseases. Al- (1) Air dry specimen and fix with Road, Palo Alto, though described more than a century ago, the methanol or heat. California, USA 94304, Gram stain remains the most frequently used (2) Add crystal violet stain. USA rapid diagnostic test, and in conjunction with W J Steinbach various biochemical tests is the cornerstone of (3) Rinse with water to wash unbound A K Shetty the clinical laboratory. First described by Dan- dye, add mordant (for example, iodine: 12 potassium iodide). Correspondence to: ish pathologist Christian Gram in 1884 and Dr Steinbach later slightly modified, the Gram stain easily (4) After waiting 30–60 seconds, rinse with [email protected] divides bacteria into two groups, Gram positive water. Submitted 27 March 2000 and Gram negative, on the basis of their cell (5) Add decolorising solvent (ethanol or Accepted 5 June 2000 wall and cell membrane permeability to acetone) to remove unbound dye. Growth on artificial medium Obligate intracellular (6) Counterstain with safranin. Chlamydia Legionella Gram positive bacteria stain blue Coxiella Ehrlichia Rickettsia (retained crystal violet).