Fgf/Ets Signalling in Xenopus Ectoderm Initiates Neural Induction and Patterning in An
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genome-Wide DNA Methylation Analysis of KRAS Mutant Cell Lines Ben Yi Tew1,5, Joel K
www.nature.com/scientificreports OPEN Genome-wide DNA methylation analysis of KRAS mutant cell lines Ben Yi Tew1,5, Joel K. Durand2,5, Kirsten L. Bryant2, Tikvah K. Hayes2, Sen Peng3, Nhan L. Tran4, Gerald C. Gooden1, David N. Buckley1, Channing J. Der2, Albert S. Baldwin2 ✉ & Bodour Salhia1 ✉ Oncogenic RAS mutations are associated with DNA methylation changes that alter gene expression to drive cancer. Recent studies suggest that DNA methylation changes may be stochastic in nature, while other groups propose distinct signaling pathways responsible for aberrant methylation. Better understanding of DNA methylation events associated with oncogenic KRAS expression could enhance therapeutic approaches. Here we analyzed the basal CpG methylation of 11 KRAS-mutant and dependent pancreatic cancer cell lines and observed strikingly similar methylation patterns. KRAS knockdown resulted in unique methylation changes with limited overlap between each cell line. In KRAS-mutant Pa16C pancreatic cancer cells, while KRAS knockdown resulted in over 8,000 diferentially methylated (DM) CpGs, treatment with the ERK1/2-selective inhibitor SCH772984 showed less than 40 DM CpGs, suggesting that ERK is not a broadly active driver of KRAS-associated DNA methylation. KRAS G12V overexpression in an isogenic lung model reveals >50,600 DM CpGs compared to non-transformed controls. In lung and pancreatic cells, gene ontology analyses of DM promoters show an enrichment for genes involved in diferentiation and development. Taken all together, KRAS-mediated DNA methylation are stochastic and independent of canonical downstream efector signaling. These epigenetically altered genes associated with KRAS expression could represent potential therapeutic targets in KRAS-driven cancer. Activating KRAS mutations can be found in nearly 25 percent of all cancers1. -

Accompanies CD8 T Cell Effector Function Global DNA Methylation
Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer, Benjamin G. Barwick, Benjamin A. Youngblood, Rafi Ahmed and Jeremy M. Boss This information is current as of October 1, 2021. J Immunol 2013; 191:3419-3429; Prepublished online 16 August 2013; doi: 10.4049/jimmunol.1301395 http://www.jimmunol.org/content/191/6/3419 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130139 Material 5.DC1 References This article cites 81 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/191/6/3419.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer,* Benjamin G. Barwick,* Benjamin A. Youngblood,*,† Rafi Ahmed,*,† and Jeremy M. -

Identification of Potential Key Genes and Pathway Linked with Sporadic Creutzfeldt-Jakob Disease Based on Integrated Bioinformatics Analyses
medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Identification of potential key genes and pathway linked with sporadic Creutzfeldt-Jakob disease based on integrated bioinformatics analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Abstract Sporadic Creutzfeldt-Jakob disease (sCJD) is neurodegenerative disease also called prion disease linked with poor prognosis. The aim of the current study was to illuminate the underlying molecular mechanisms of sCJD. The mRNA microarray dataset GSE124571 was downloaded from the Gene Expression Omnibus database. Differentially expressed genes (DEGs) were screened. -

Single Cell Regulatory Landscape of the Mouse Kidney Highlights Cellular Differentiation Programs and Disease Targets
ARTICLE https://doi.org/10.1038/s41467-021-22266-1 OPEN Single cell regulatory landscape of the mouse kidney highlights cellular differentiation programs and disease targets Zhen Miao 1,2,3,8, Michael S. Balzer 1,2,8, Ziyuan Ma 1,2,8, Hongbo Liu1,2, Junnan Wu 1,2, Rojesh Shrestha 1,2, Tamas Aranyi1,2, Amy Kwan4, Ayano Kondo 4, Marco Pontoglio 5, Junhyong Kim6, ✉ Mingyao Li 7, Klaus H. Kaestner2,4 & Katalin Susztak 1,2,4 1234567890():,; Determining the epigenetic program that generates unique cell types in the kidney is critical for understanding cell-type heterogeneity during tissue homeostasis and injury response. Here, we profile open chromatin and gene expression in developing and adult mouse kidneys at single cell resolution. We show critical reliance of gene expression on distal regulatory elements (enhancers). We reveal key cell type-specific transcription factors and major gene- regulatory circuits for kidney cells. Dynamic chromatin and expression changes during nephron progenitor differentiation demonstrates that podocyte commitment occurs early and is associated with sustained Foxl1 expression. Renal tubule cells follow a more complex differentiation, where Hfn4a is associated with proximal and Tfap2b with distal fate. Mapping single nucleotide variants associated with human kidney disease implicates critical cell types, developmental stages, genes, and regulatory mechanisms. The single cell multi-omics atlas reveals key chromatin remodeling events and gene expression dynamics associated with kidney development. 1 Renal, Electrolyte, and Hypertension Division, Department of Medicine, University of Pennsylvania, Perelman School of Medicine, Philadelphia, PA, USA. 2 Institute for Diabetes, Obesity, and Metabolism, University of Pennsylvania, Perelman School of Medicine, Philadelphia, PA, USA. -

Evidence for a Role of Developmental Genes in the Origin of Obesity and Body Fat Distribution
Evidence for a role of developmental genes in the origin of obesity and body fat distribution Stephane Gesta*, Matthias Blu¨ her†, Yuji Yamamoto*, Andrew W. Norris*, Janin Berndt†, Susan Kralisch†, Jeremie Boucher*, Choy Lewis*, and C. Ronald Kahn*‡ *Joslin Diabetes Center and Harvard Medical School, Boston, MA 02215; and †Department of Internal Medicine III, University of Leipzig, 04103 Leipzig, Germany Contributed by C. Ronald Kahn, March 9, 2006 Obesity, especially central obesity, is a hereditable trait associated In the present study, we have explored the hypothesis that with a high risk for development of diabetes and metabolic patterns of fat distribution and, perhaps, to some degree, obesity disorders. Combined gene expression analysis of adipocyte- and itself may have a developmental genetic origin. Indeed, we find preadipocyte-containing fractions from intraabdominal and sub- major differences in expression of multiple genes involved in cutaneous adipose tissue of mice revealed coordinated depot- embryonic development and pattern specification between adipo- specific differences in expression of multiple genes involved in cytes taken from intraabdominal and s.c. depots in rodents and embryonic development and pattern specification. These differ- humans. We also demonstrate similar differences in the stromo- ences were intrinsic and persisted during in vitro culture and vascular fraction (SVF)-containing preadipocytes and that these differentiation. Similar depot-specific differences in expression of differences persist in culture. Most importantly, we demonstrate developmental genes were observed in human subcutaneous ver- that some of these developmental genes exhibit changes in expres- sus visceral adipose tissue. Furthermore, in humans, several genes sion that are closely correlated with the level of obesity and the exhibited changes in expression that correlated closely with body pattern of fat distribution. -

Supplementary Material Sumoylation Regulates the Chromatin
Supplementary material SUMOylation Regulates the Chromatin Occupancy and Anti-Proliferative Gene Programs of Glucocorticoid Receptor Ville Paakinaho, Sanna Kaikkonen, Harri Makkonen, Vladimir Benes, and Jorma J. Palvimo A FRT wtGR GR3KR clone wt-1 wt-2 wt-3 3KR-1 3KR-2 3KR-3 - - - - - - - dex + + + + + + + kDa a-GR - 100 a-GAPDH - 35 A549 + - - HEK293-wtGR - + - - - HEK293-GR3KR + kDa a-GR - 100 a-GAPDH - 35 B wtGR GR3KR 43°C - - + + - - + + dex - + - + - + - + kDa - 170 a-GR - 130 - 100 a-GAPDH - 35 - 170 a-SUMO-1 - 130 R - 100 G - a : P I a-SUMO-2/3 - 170 - 130 - 100 Figure S1. Expression and SUMOylation of GR in stable isogenic HEK293 cells. (A) Upper panel, immunoblot analysis of three HEK293 cell clones expressing either wtGR (wt-1-3) or GR3KR (3KR-1-3). The clones and background HEK293 (FRT) cells were exposed to vehicle or dex for 17 h, and cell samples were immunoblotting with anti-GR (Santa Cruz, sc-1003) and anti- GAPDH (Santa Cruz, sc-25778) antibodies. Lower panel, comparison of the GR level in A549 cells (from ATCC) with that of the selected clones from isogenic HEK293 cells (wt-3 and 3KR-3) by immunoblotting. (B) To analyse GR SUMOylation, wtGR- and GR3KR-expressing cells were grown with of without dex for 2 h and exposed for 30 min to 43°C. Immunoprecipitation (IP) with anti- GR antibody and immunoblotting with anti-SUMO-1 (Invitrogen, 33-2400) or anti-SUMO-2/3 (MBL M114-3) antibody were performed essentially as described (Rytinki et al. 2011, Methods Mol. Biol.). A total dex- sensitivity induced for dex expression ADARB1 -

Aberrant HOXC Expression Accompanies the Malignant Phenotype in Human Prostate1
[CANCER RESEARCH 63, 5879–5888, September 15, 2003] Aberrant HOXC Expression Accompanies the Malignant Phenotype in Human Prostate1 Gary J. Miller,2 Heidi L. Miller, Adrie van Bokhoven, James R. Lambert, Priya N. Werahera, Osvaldo Schirripa,3 M. Scott Lucia, and Steven K. Nordeen4 Department of Pathology, University of Colorado Health Sciences Center, Denver, Colorado 80262 ABSTRACT breast (13, 14), and renal (15) carcinomas; melanomas (16); and squamous carcinomas of the skin (17). Because the genes implicated Dysregulation of HOX gene expression has been implicated as a factor show little consensus, the dysregulation may be a tissue-specific in malignancies for a number of years. However, no consensus has perturbation of the existing HOX expression pattern rather than a emerged regarding specific causative genes. Using a degenerate reverse transcription-PCR technique, we show up-regulation of genes from the single causative gene. Tissue-specific expression patterns have been HOXC cluster in malignant prostate cell lines and lymph node metastases. reported in kidney and colon, by Northern blot analysis (12, 15). When relative expression levels of the four HOX clusters were examined, Primary tumors in both kidney and colon showed variations in spe- lymph node metastases and cell lines derived from lymph node metastases cific HOX gene expression from the corresponding normal tissue, but exhibited very similar patterns, patterns distinct from those in benign cells overall expression patterns for individual tumors were not reported. or malignant cell lines derived from other tumor sites. Specific reverse Only primary kidney tumors were examined (15), but liver metastases transcription-PCR for HOXC4, HOXC5, HOXC6, and HOXC8 confirmed from colon tumors reportedly displayed expression of specific HOX overexpression of these genes in malignant cell lines and lymph node genes similar to that seen in either primary colon tumors or normal metastases. -

The Notch Target Hes1 Directly Modulates Gli1 Expression and Hedgehog Signaling: a Potential Mechanism of Therapeutic Resistance
Clinical Cancer Cancer Therapy: Preclinical Research The Notch Target Hes1 Directly Modulates Gli1 Expression and Hedgehog Signaling: A Potential Mechanism of Therapeutic Resistance Karisa C. Schreck1,2,5, Pete Taylor6, Luigi Marchionni4, Vidya Gopalakrishnan6, Eli E. Bar2, Nicholas Gaiano1,3,5, and Charles G. Eberhart2,4 Abstract Purpose: Multiple developmental pathways including Notch, Hedgehog, and Wnt are active in malignant brain tumors such as medulloblastoma and glioblastoma (GBM). This raises the possibility that tumors might compensate for therapy directed against one pathway by upregulating a different one. We investigated whether brain tumors show resistance to therapies against Notch, and whether targeting multiple pathways simultaneously would kill brain tumor cells more effectively than monotherapy. Experimental Design: We used GBM neurosphere lines to investigate the effects of a gamma-secretase inhibitor (MRK-003) on tumor growth, and chromatin immunoprecipitation to study the regulation of other genes by Notch targets. We also evaluated the effect of combined therapy with a Hedgehog inhibitor (cyclopamine) in GBM and medulloblastoma lines, and in primary human GBM cultures. Results: GBM cells are at least partially resistant to long-term MRK-003 treatment, despite ongoing Notch pathway suppression, and show concomitant upregulation of Wnt and Hedgehog activity. The Notch target Hes1, a repressive transcription factor, bound the Gli1 first intron, and may inhibit its expression. Similar results were observed in a melanoma-derived cell line. Targeting Notch and Hedgehog simultaneously induced apoptosis, decreased cell growth, and inhibited colony-forming ability more dramatically than monotherapy. Low-passage neurospheres isolated from freshly resected human GBMs were also highly susceptible to coinhibition of the two pathways, indicating that targeting multiple developmental pathways can be more effective than monotherapy at eliminating GBM-derived cells. -

A PHOSPHORYLATION SITE in the FTZ HOMEODOMAIN IS REQUIRED for Segrnntation in DROSOPHILA
A PHOSPHORYLATION SITE IN THE FTZ HOMEODOMAIN IS REQUIRED FOR SEGrnNTATION IN DROSOPHILA Jianli Dong A thesis submitted in conformity with the requirements r for the degree of Doctor of Philosophy Graduate Department of Molecular and Medical Genetics University of Toronto O Copyright by Jianli Dong 1999 National Library Biblioth&que nationale du Canada Acquisitions and Acquisitions et Bibliographic Services services bibliographiques 395 Wellington Street 395, rue Wellington Ottawa ON KIA ON4 OttawaON KlAON4 Canada Canada YOW m~ vam dnrrcnr00 Our Me Nolre dfdf~ll~~ The author has granted a non- L'auteur a accorde melicence non exclusive licence allowing the exclusive pennettant &la National Library of Canada to Bibliotheque nationale du Canada de reproduce, loan, distribute or sell reproduire, prster, distribuer ou copies of this thesis in microform, vendre des copies de cette these sous paper or electronic formats. la forme de microfiche/film, de reproduction sur papier ou sur format Bectronique. The author retains ownership of the L'auteur conserve la propriete du copyright in this thesis. Neither the droit d'auteur qui protege cette these. thesis nor substantial extracts fiom it Ni la these ni des extraits substantiels may be printed or otherwise de celle-ci ne doivent Stre imphes reproduced without the author's ou autrement reproduits sans son permission. autorisation. A Phosphorylation site in the Ftz homeodomain is required for segmentation in Drosophila Doctor of Philosophy, 1999 Jianli Dong Department of Molecular and Medical Genetics University of Toronto Abstract The Drosophila homeodomain-containing protein Fushi tarazu (Ftz) is expressed sequentially in the embryo, first in alternate segments, then in specific neuroblasts and neurons in the central nervous system and finally in parts of the gut. -

Robles JTO Supplemental Digital Content 1
Supplementary Materials An Integrated Prognostic Classifier for Stage I Lung Adenocarcinoma based on mRNA, microRNA and DNA Methylation Biomarkers Ana I. Robles1, Eri Arai2, Ewy A. Mathé1, Hirokazu Okayama1, Aaron Schetter1, Derek Brown1, David Petersen3, Elise D. Bowman1, Rintaro Noro1, Judith A. Welsh1, Daniel C. Edelman3, Holly S. Stevenson3, Yonghong Wang3, Naoto Tsuchiya4, Takashi Kohno4, Vidar Skaug5, Steen Mollerup5, Aage Haugen5, Paul S. Meltzer3, Jun Yokota6, Yae Kanai2 and Curtis C. Harris1 Affiliations: 1Laboratory of Human Carcinogenesis, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 2Division of Molecular Pathology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 3Genetics Branch, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 4Division of Genome Biology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 5Department of Chemical and Biological Working Environment, National Institute of Occupational Health, NO-0033 Oslo, Norway. 6Genomics and Epigenomics of Cancer Prediction Program, Institute of Predictive and Personalized Medicine of Cancer (IMPPC), 08916 Badalona (Barcelona), Spain. List of Supplementary Materials Supplementary Materials and Methods Fig. S1. Hierarchical clustering of based on CpG sites differentially-methylated in Stage I ADC compared to non-tumor adjacent tissues. Fig. S2. Confirmatory pyrosequencing analysis of DNA methylation at the HOXA9 locus in Stage I ADC from a subset of the NCI microarray cohort. 1 Fig. S3. Methylation Beta-values for HOXA9 probe cg26521404 in Stage I ADC samples from Japan. Fig. S4. Kaplan-Meier analysis of HOXA9 promoter methylation in a published cohort of Stage I lung ADC (J Clin Oncol 2013;31(32):4140-7). Fig. S5. Kaplan-Meier analysis of a combined prognostic biomarker in Stage I lung ADC. -

Microarray Characterization of Human Embryonic Stem Cell–Derived Retinal Cultures
Retinal Cell Biology Microarray Characterization of Human Embryonic Stem Cell–Derived Retinal Cultures Deepak A. Lamba1,2 and Thomas A. Reh2,3 PURPOSE. A number of protocols have been published to induce photoreceptor cells to cause significant visual impairment retinal determination from human embryonic stem cells (hESC) and blindness. The mammalian retina does not have an and induced pluripotent stem cells (iPSC). Although all these innate capacity to regenerate the dying cells (see Ref. 1 for studies have shown some degree of expression of markers of review); visual prosthetics or cell replacement strategies are retinal cells, fewer than 30 markers are typically used to char- therefore being pursued in an effort to restore some vision acterize the ESC-derived retinal cells. Hence, it is not known to the most severely affected patients. Human embryonic whether they express all the genes present in normal develop- stem cells (hESCs), because of their property of pluripo- ing retinal cells. To assess the efficiency of their retinal deter- tency, may have a significant role in cell replacement ther- mination protocol at the transcriptome level and to understand apies in the eye. However, this same property requires that the changes in human retinal gene expression patterns during the cells be directed in their differentiation to retinal cell development, the authors conducted a microarray-based anal- fates before use in therapies to minimize the risks of devel- ysis comparing human retina to hESC-derived retinal cells. oping teratomas. Several groups, including our own, have METHODS. The authors extracted total RNA from 60-day, 80-day, previously shown that a combination of signaling molecules and 96-day human fetal retina and hESC-derived retinal cells at can direct ESCs to retinal cell fates (see Discussion). -
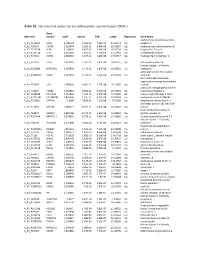
Table S1. the Statistical Metrics for Key Differentially Expressed Genes (Degs)
Table S1. The statistical metrics for key differentially expressed genes (DEGs) Gene Agilent Id Symbol logFC pValue FDR tvalue Regulation Gene Name oxidized low density lipoprotein A_24_P124624 OLR1 2.458429 1.19E-13 7.25E-10 24.04241 Up receptor 1 A_23_P90273 CHST8 2.622464 3.85E-12 6.96E-09 19.05867 Up carbohydrate sulfotransferase 8 A_23_P217528 KLF8 2.109007 4.85E-12 7.64E-09 18.76234 Up Kruppel like factor 8 A_23_P114740 CFH 2.651636 1.85E-11 1.79E-08 17.13652 Up complement factor H A_23_P34031 XAGE2 2.000935 2.04E-11 1.81E-08 17.02457 Up X antigen family member 2 A_23_P27332 TCF4 1.613097 2.32E-11 1.87E-08 16.87275 Up transcription factor 4 histone cluster 1 H1 family A_23_P250385 HIST1H1B 2.298658 2.47E-11 1.87E-08 16.80362 Up member b abnormal spindle microtubule A_33_P3288159 ASPM 2.162032 2.79E-11 2.01E-08 16.66292 Up assembly H19, imprinted maternally expressed transcript (non-protein A_24_P52697 H19 1.499364 4.09E-11 2.76E-08 16.23387 Up coding) potassium voltage-gated channel A_24_P31627 KCNB1 2.289689 6.65E-11 3.97E-08 15.70253 Up subfamily B member 1 A_23_P214168 COL12A1 2.155835 7.59E-11 4.15E-08 15.56005 Up collagen type XII alpha 1 chain A_33_P3271341 LOC388282 2.859496 7.61E-11 4.15E-08 15.55704 Up uncharacterized LOC388282 A_32_P150891 DIAPH3 2.2068 7.83E-11 4.22E-08 15.5268 Up diaphanous related formin 3 zinc finger protein 185 with LIM A_23_P11025 ZNF185 1.385721 8.74E-11 4.59E-08 15.41041 Up domain heat shock protein family B A_23_P96872 HSPB11 1.887166 8.94E-11 4.64E-08 15.38599 Up (small) member 11 A_23_P107454