EIF3G Polyclonal Antibody Catalog Number:11165-1-AP 1 Publications
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genes with 5' Terminal Oligopyrimidine Tracts Preferentially Escape Global Suppression of Translation by the SARS-Cov-2 NSP1 Protein
Downloaded from rnajournal.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Genes with 5′ terminal oligopyrimidine tracts preferentially escape global suppression of translation by the SARS-CoV-2 Nsp1 protein Shilpa Raoa, Ian Hoskinsa, Tori Tonna, P. Daniela Garciaa, Hakan Ozadama, Elif Sarinay Cenika, Can Cenika,1 a Department of Molecular Biosciences, University of Texas at Austin, Austin, TX 78712, USA 1Corresponding author: [email protected] Key words: SARS-CoV-2, Nsp1, MeTAFlow, translation, ribosome profiling, RNA-Seq, 5′ TOP, Ribo-Seq, gene expression 1 Downloaded from rnajournal.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Abstract Viruses rely on the host translation machinery to synthesize their own proteins. Consequently, they have evolved varied mechanisms to co-opt host translation for their survival. SARS-CoV-2 relies on a non-structural protein, Nsp1, for shutting down host translation. However, it is currently unknown how viral proteins and host factors critical for viral replication can escape a global shutdown of host translation. Here, using a novel FACS-based assay called MeTAFlow, we report a dose-dependent reduction in both nascent protein synthesis and mRNA abundance in cells expressing Nsp1. We perform RNA-Seq and matched ribosome profiling experiments to identify gene-specific changes both at the mRNA expression and translation level. We discover that a functionally-coherent subset of human genes are preferentially translated in the context of Nsp1 expression. These genes include the translation machinery components, RNA binding proteins, and others important for viral pathogenicity. Importantly, we uncovered a remarkable enrichment of 5′ terminal oligo-pyrimidine (TOP) tracts among preferentially translated genes. -

GPRC5A Suppresses Protein Synthesis at the Endoplasmic Reticulum to Prevent Radiation-Induced Lung Tumorigenesis
ARTICLE Received 8 Mar 2016 | Accepted 28 Apr 2016 | Published 8 Jun 2016 DOI: 10.1038/ncomms11795 OPEN GPRC5A suppresses protein synthesis at the endoplasmic reticulum to prevent radiation-induced lung tumorigenesis Jian Wang1, Alton B. Farris2, Kaiming Xu1, Ping Wang1, Xiangming Zhang1, Duc M. Duong3, Hong Yi4, Hui-Kuo Shu1, Shi-Yong Sun5 & Ya Wang1 GPRC5A functions as a lung tumour suppressor to prevent spontaneous and environmentally induced lung carcinogenesis; however, the underlying mechanism remains unclear. Here we reveal that GPRC5A at the endoplasmic reticulum (ER) membrane suppresses synthesis of the secreted or membrane-bound proteins including a number of oncogenes, the most important one being Egfr. The ER-located GPRC5A disturbs the assembly of the eIF4F-mediated translation initiation complex on the mRNA cap through directly binding to the eIF4F complex with its two middle extracellular loops. Particularly, suppression of EGFR by GPRC5A contributes significantly to preventing ionizing radiation (IR)-induced lung tumorigenesis. Thus, GPRC5A deletion enhances IR-promoted EGFR expression through an increased translation rate, thereby significantly increasing lung tumour incidence in Gprc5a À / À mice. Our findings indicate that under-expressed GPRC5A during lung tumor- igenesis enhances any transcriptional stimulation through an active translational status, which can be used to control oncogene expression and potentially the resulting related disease. 1 Department of Radiation Oncology, Emory University School of Medicine and the Winship Cancer Institute, Emory University, Atlanta, Georgia 30322, USA. 2 Department of Pathology, Emory University School of Medicine and the Winship Cancer Institute, Emory University, Atlanta, Georgia 30322, USA. 3 Emory Integrated Proteomics Core and Biochemistry Department, Atlanta, Georgia 30322, USA. -

Membrane Proteomics of Cervical Cancer Cell Lines Reveal Insights on the Process of Cervical Carcinogenesis
INTERNATIONAL JOURNAL OF ONCOLOGY 53: 2111-2122, 2018 Membrane proteomics of cervical cancer cell lines reveal insights on the process of cervical carcinogenesis KALLIOPI I. PAPPA1,2, POLYXENI CHRISTOU3,4, AMARILDO XHOLI3, GEORGE MERMELEKAS3, GEORGIA KONTOSTATHI3,4, VASILIKI LYGIROU3,4, MANOUSOS MAKRIDAKIS3, JEROME ZOIDAKIS3 and NICHOLAS P. ANAGNOU1,4 1Cell and Gene Therapy Laboratory, Centre of Basic Research II, Biomedical Research Foundation of the Academy of Athens, 11527 Athens; 2First Department of Obstetrics and Gynecology, University of Athens School of Medicine, Alexandra Hospital, 11528 Athens; 3Biotechnology Division, Centre of Basic Research, Biomedical Research Foundation of the Academy of Athens; 4Laboratory of Biology, University of Athens School of Medicine, 11527 Athens, Greece Received March 22, 2018; Accepted May 4, 2018 DOI: 10.3892/ijo.2018.4518 Abstract. The available therapeutic approaches for cervical biological pathways relevant to malignancy, including ‘HIPPO cancer can seriously affect the fertility potential of patient; signaling’, ‘PI3K/Akt signaling’, ‘cell cycle: G2/M DNA thus, there is a pressing requirement for less toxic and damage checkpoint regulation’ and ‘EIF2 signaling’. These targeted therapies. The membrane proteome is a potential unique membrane protein identifications offer insights on a source of therapeutic targets; however, despite the signifi- previously inaccessible region of the cervical cancer proteome, cance of membrane proteins in cancer, proteomic analysis and may represent putative -
Type of the Paper (Article
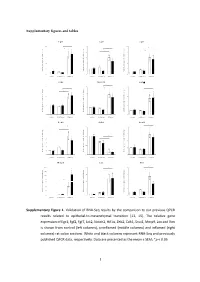
Supplementary figures and tables E g r 1 F g f2 F g f7 1 0 * 5 1 0 * * e e e * g g g * n n n * a a a 8 4 * 8 h h h * c c c d d d * l l l o o o * f f f * n n n o o o 6 3 6 i i i s s s s s s e e e r r r p p p x x x e e e 4 2 4 e e e n n n e e e g g g e e e v v v i i i t t t 2 1 2 a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d J a k 2 N o tc h 2 H if1 * 3 4 6 * * * e e e g g g n n n a a * * a * h h * h c c c 3 * d d * d l l l * o o o f f 2 f 4 n n n o o o i i i s s s s s s e e e r r 2 r p p p x x x e e e e e e n n n e e 1 e 2 g g g e e 1 e v v v i i i t t t a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Z e b 2 C d h 1 S n a i1 * * 7 1 .5 4 * * e e e g g g 6 n n n * a a a * h h h c c c 3 * d d d l l l 5 o o o f f f 1 .0 * n n n * o o o i i i 4 * s s s s s s e e e r r r 2 p p p x x x 3 e e e e e e n n n e e e 0 .5 g g g 2 e e e 1 v v v i i i t t t a a a * l l l e e e 1 * R R R 0 0 .0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d M m p 9 L o x V im 2 0 0 2 0 8 * * * e e e * g g g 1 5 0 * n n n * a a a * h h h * c c c 1 5 * 6 d d d l l l 1 0 0 o o o f f f n n n o o o i i i 5 0 s s s s s s * e e e r r r 1 0 4 3 0 p p p * x x x e e e * e e e n n n e e e 2 0 g g g e e e 5 2 v v v i i i t t t a a a l l l 1 0 e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Supplementary Figure 1. -

Systematically Profiling the Expression of Eif3 Subunits in Glioma Reveals
Chai et al. Cancer Cell Int (2019) 19:155 https://doi.org/10.1186/s12935-019-0867-1 Cancer Cell International PRIMARY RESEARCH Open Access Systematically profling the expression of eIF3 subunits in glioma reveals the expression of eIF3i has prognostic value in IDH-mutant lower grade glioma Rui‑Chao Chai1,4,6†, Ning Wang2†, Yu‑Zhou Chang3, Ke‑Nan Zhang1,6, Jing‑Jun Li1,6, Jun‑Jie Niu5, Fan Wu1,6*, Yu‑Qing Liu1,6* and Yong‑Zhi Wang1,3,4,6* Abstract Background: Abnormal expression of the eukaryotic initiation factor 3 (eIF3) subunits plays critical roles in tumo‑ rigenesis and progression, and also has potential prognostic value in cancers. However, the expression and clinical implications of eIF3 subunits in glioma remain unknown. Methods: Expression data of eIF3 for patients with gliomas were obtained from the Chinese Glioma Genome Atlas (CGGA) (n 272) and The Cancer Genome Atlas (TCGA) (n 595). Cox regression, the receiver operating characteristic (ROC) curves= and Kaplan–Meier analysis were used to study= the prognostic value. Gene oncology (GO) and gene set enrichment analysis (GSEA) were utilized for functional prediction. Results: In both the CGGA and TCGA datasets, the expression levels of eIF3d, eIF3e, eIF3f, eIF3h and eIF3l highly were associated with the IDH mutant status of gliomas. The expression of eIF3b, eIF3i, eIF3k and eIF3m was increased with the tumor grade, and was associated with poorer overall survival [All Hazard ratio (HR) > 1 and P < 0.05]. By contrast, the expression of eIF3a and eIF3l was decreased in higher grade gliomas and was associated with better overall sur‑ vival (Both HR < 1 and P < 0.05). -

Translation Initiation Factor Eif3h Targets Specific Transcripts To
Translation initiation factor eIF3h targets specific transcripts to polysomes during embryogenesis Avik Choudhuria,b, Umadas Maitraa,1, and Todd Evansb,1 aDepartment of Developmental and Molecular Biology, Albert Einstein College of Medicine of Yeshiva University, Bronx, NY 10461; and bDepartment of Surgery, Weill Cornell Medical College, New York, NY 10065 Edited by Igor B. Dawid, The Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, MD, and approved April 30, 2013 (received for review February 14, 2013) Eukaryotic translation initiation factor 3 (eIF3) plays a central role eukaryotes. These—eIF3d, eIF3e, eIF3f, eIF3h, eIF3j, eIF3k, in translation initiation and consists of five core (conserved) sub- eIF3l, and eIF3m—were designated “non-core” subunits (4). units present in both budding yeast and higher eukaryotes. Higher In contrast to the budding yeast, the genome of the fission eukaryotic eIF3 contains additional (noncore or nonconserved) yeast Schizosaccharomyces pombe contains structural homologs subunits of poorly defined function, including sub-unit h (eIF3h), of at least five noncore (nonconserved) eIF3 subunits—eIF3d, which in zebrafish is encoded by two distinct genes (eif3ha and eIF3e, eIF3f, eIF3h, and eIF3m. The gene encoding eIF3f is eif3hb). Previously we showed that eif3ha encodes the predominant essential for growth, whereas eIF3d, eIF3e, and eIF3h are dis- isoform during zebrafish embryogenesis and that depletion of this pensable for growth and viability (5–11). However, deleted factor causes defects in the development of the brain and eyes. To strains show specific phenotypes including defects in meiosis/ investigate the molecular mechanism governing this regulation, we sporulation (6, 9, 11). -

Relevance of Translation Initiation in Diffuse Glioma Biology and Its
cells Review Relevance of Translation Initiation in Diffuse Glioma Biology and its Therapeutic Potential Digregorio Marina 1, Lombard Arnaud 1,2, Lumapat Paul Noel 1, Scholtes Felix 1,2, Rogister Bernard 1,3 and Coppieters Natacha 1,* 1 Laboratory of Nervous System Disorders and Therapy, GIGA-Neurosciences Research Centre, University of Liège, 4000 Liège, Belgium; [email protected] (D.M.); [email protected] (L.A.); [email protected] (L.P.N.); [email protected] (S.F.); [email protected] (R.B.) 2 Department of Neurosurgery, CHU of Liège, 4000 Liège, Belgium 3 Department of Neurology, CHU of Liège, 4000 Liège, Belgium * Correspondence: [email protected] Received: 18 October 2019; Accepted: 26 November 2019; Published: 29 November 2019 Abstract: Cancer cells are continually exposed to environmental stressors forcing them to adapt their protein production to survive. The translational machinery can be recruited by malignant cells to synthesize proteins required to promote their survival, even in times of high physiological and pathological stress. This phenomenon has been described in several cancers including in gliomas. Abnormal regulation of translation has encouraged the development of new therapeutics targeting the protein synthesis pathway. This approach could be meaningful for glioma given the fact that the median survival following diagnosis of the highest grade of glioma remains short despite current therapy. The identification of new targets for the development of novel therapeutics is therefore needed in order to improve this devastating overall survival rate. This review discusses current literature on translation in gliomas with a focus on the initiation step covering both the cap-dependent and cap-independent modes of initiation. -

Inhibition of the MID1 Protein Complex
Matthes et al. Cell Death Discovery (2018) 4:4 DOI 10.1038/s41420-017-0003-8 Cell Death Discovery ARTICLE Open Access Inhibition of the MID1 protein complex: a novel approach targeting APP protein synthesis Frank Matthes1,MoritzM.Hettich1, Judith Schilling1, Diana Flores-Dominguez1, Nelli Blank1, Thomas Wiglenda2, Alexander Buntru2,HannaWolf1, Stephanie Weber1,InaVorberg 1, Alina Dagane2, Gunnar Dittmar2,3,ErichWanker2, Dan Ehninger1 and Sybille Krauss1 Abstract Alzheimer’s disease (AD) is characterized by two neuropathological hallmarks: senile plaques, which are composed of amyloid-β (Aβ) peptides, and neurofibrillary tangles, which are composed of hyperphosphorylated tau protein. Aβ peptides are derived from sequential proteolytic cleavage of the amyloid precursor protein (APP). In this study, we identified a so far unknown mode of regulation of APP protein synthesis involving the MID1 protein complex: MID1 binds to and regulates the translation of APP mRNA. The underlying mode of action of MID1 involves the mTOR pathway. Thus, inhibition of the MID1 complex reduces the APP protein level in cultures of primary neurons. Based on this, we used one compound that we discovered previously to interfere with the MID1 complex, metformin, for in vivo experiments. Indeed, long-term treatment with metformin decreased APP protein expression levels and consequently Aβ in an AD mouse model. Importantly, we have initiated the metformin treatment late in life, at a time-point where mice were in an already progressed state of the disease, and could observe an improved behavioral phenotype. These 1234567890 1234567890 findings together with our previous observation, showing that inhibition of the MID1 complex by metformin also decreases tau phosphorylation, make the MID1 complex a particularly interesting drug target for treating AD. -

Lncrna SNHG8 Is Identified As a Key Regulator of Acute Myocardial
Zhuo et al. Lipids in Health and Disease (2019) 18:201 https://doi.org/10.1186/s12944-019-1142-0 RESEARCH Open Access LncRNA SNHG8 is identified as a key regulator of acute myocardial infarction by RNA-seq analysis Liu-An Zhuo, Yi-Tao Wen, Yong Wang, Zhi-Fang Liang, Gang Wu, Mei-Dan Nong and Liu Miao* Abstract Background: Long noncoding RNAs (lncRNAs) are involved in numerous physiological functions. However, their mechanisms in acute myocardial infarction (AMI) are not well understood. Methods: We performed an RNA-seq analysis to explore the molecular mechanism of AMI by constructing a lncRNA-miRNA-mRNA axis based on the ceRNA hypothesis. The target microRNA data were used to design a global AMI triple network. Thereafter, a functional enrichment analysis and clustering topological analyses were conducted by using the triple network. The expression of lncRNA SNHG8, SOCS3 and ICAM1 was measured by qRT-PCR. The prognostic values of lncRNA SNHG8, SOCS3 and ICAM1 were evaluated using a receiver operating characteristic (ROC) curve. Results: An AMI lncRNA-miRNA-mRNA network was constructed that included two mRNAs, one miRNA and one lncRNA. After RT-PCR validation of lncRNA SNHG8, SOCS3 and ICAM1 between the AMI and normal samples, only lncRNA SNHG8 had significant diagnostic value for further analysis. The ROC curve showed that SNHG8 presented an AUC of 0.850, while the AUC of SOCS3 was 0.633 and that of ICAM1 was 0.594. After a pairwise comparison, we found that SNHG8 was statistically significant (P SNHG8-ICAM1 = 0.002; P SNHG8-SOCS3 = 0.031). -

Nat1 Promotes Translation of Specific Proteins That Induce Differentiation of Mouse Embryonic Stem Cells
Nat1 promotes translation of specific proteins that induce differentiation of mouse embryonic stem cells Hayami Sugiyamaa, Kazutoshi Takahashia,b, Takuya Yamamotoa,c,d, Mio Iwasakia, Megumi Naritaa, Masahiro Nakamuraa, Tim A. Randb, Masato Nakagawaa, Akira Watanabea,c,e, and Shinya Yamanakaa,b,1 aDepartment of Life Science Frontiers, Center for iPS Cell Research and Application, Kyoto University, Kyoto 606-8507, Japan; bGladstone Institute of Cardiovascular Disease, San Francisco, CA 94158; cInstitute for Integrated Cell-Material Sciences, Kyoto University, Kyoto 606-8501, Japan; dJapan Agency for Medical Research and Development-Core Research for Evolutional Science and Technology (AMED-CREST), Tokyo 100-0004, Japan; and eJapan Science and Technology Agency (JST)-CREST, Saitama 332-0012, Japan Contributed by Shinya Yamanaka, November 22, 2016 (sent for review October 18, 2016; reviewed by Katsura Asano and Keisuke Kaji) Novel APOBEC1 target 1 (Nat1) (also known as “p97,”“Dap5,” and To study the role of Nat1 in cell differentiation further, we “Eif4g2”) is a ubiquitously expressed cytoplasmic protein that is homol- generated mouse embryonic stem cells (mES cells) lacking both ogous to the C-terminal two thirds of eukaryotic translation initiation alleles of the Nat1 gene. mES cells were derived from blastocysts factor 4G (Eif4g1). We previously showed that Nat1-null mouse embry- in 1981 (11, 12) and possess two unique properties. First, ES cells onic stem cells (mES cells) are resistant to differentiation. In the current have the potential to self-renew indefinitely (maintenance). Sec- study, we found that NAT1 and eIF4G1 share many binding proteins, ond, ES cells have the potential to differentiate into all somatic such as the eukaryotic translation initiation factors eIF3 and eIF4A and and germ cell types (pluripotency) that make up the body. -

EIF3S4 (EIF3G) (NM 003755) Human Untagged Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for SC319338 EIF3S4 (EIF3G) (NM_003755) Human Untagged Clone Product data: Product Type: Expression Plasmids Product Name: EIF3S4 (EIF3G) (NM_003755) Human Untagged Clone Tag: Tag Free Symbol: EIF3G Synonyms: eIF3-delta; EIF3-P42; eIF3-p44; EIF3S4 Vector: pCMV6-AC (PS100020) E. coli Selection: Ampicillin (100 ug/mL) Cell Selection: Neomycin Fully Sequenced ORF: >OriGene sequence for NM_003755.3 GCTCTCTGGCCGGGCTTGGGCTGCGTGGAGAATACTTTTTGCGATGCCTACTGGAGACTT TGATTCGAAGCCCAGTTGGGCCGACCAGGTGGAGGAGGAGGGGGAGGACGACAAATGTGT CACCAGCGAGCTCCTCAAGGGGATCCCTCTGGCCACAGGTGACACCAGCCCAGAGCCAGA GCTACTGCCGGGAGCTCCACTGCCGCCTCCCAAGGAGGTCATCAACGGAAACATAAAGAC AGTGACAGAGTACAAGATAGATGAGGATGGCAAGAAGTTCAAGATTGTCCGCACCTTCAG GATTGAGACCCGGAAGGCTTCAAAGGCTGTCGCAAGGAGGAAGAACTGGAAGAAGTTCGG GAACTCAGAGTTTGACCCCCCCGGACCCAATGTGGCCACCACCACTGTCAGTGACGATGT CTCTATGACGTTCATCACCAGCAAAGAGGACCTGAACTGCCAGGAGGAGGAGGACCCTAT GAACAAACTCAAGGGCCAGAAGATCGTGTCCTGCCGCATCTGCAAGGGCGACCACTGGAC CACCCGCTGCCCCTACAAGGATACGCTGGGGCCCATGCAGAAGGAGCTGGCCGAGCAGCT GGGCCTGTCTACTGGCGAGAAGGAGAAGCTGCCGGGAGAGCTAGAGCCGGTGCAGGCCAC GCAGAACAAGACAGGGAAGTATGTGCCGCCGAGCCTGCGCGACGGGGCCAGCCGCCGCGG GGAGTCCATGCAGCCCAACCGCAGAGCCGACGACAACGCCACCATCCGTGTCACCAACTT GTCAGAGGACACGCGTGAGACCGACCTGCAGGAGCTCTTCCGGCCTTTCGGCTCTATCTC CCGCATCTACCTGGCTAAGGACAAGACCACTGGCCAATCCAAGGGCTTTGCCTTCATCAG CTTCCACCGCCGCGAGGATGCTGCGCGTGCCATTGCCGGGGTGTCCGGCTTTGGCTACGA -

Comparative Analysis of the Ubiquitin-Proteasome System in Homo Sapiens and Saccharomyces Cerevisiae
Comparative Analysis of the Ubiquitin-proteasome system in Homo sapiens and Saccharomyces cerevisiae Inaugural-Dissertation zur Erlangung des Doktorgrades der Mathematisch-Naturwissenschaftlichen Fakultät der Universität zu Köln vorgelegt von Hartmut Scheel aus Rheinbach Köln, 2005 Berichterstatter: Prof. Dr. R. Jürgen Dohmen Prof. Dr. Thomas Langer Dr. Kay Hofmann Tag der mündlichen Prüfung: 18.07.2005 Zusammenfassung I Zusammenfassung Das Ubiquitin-Proteasom System (UPS) stellt den wichtigsten Abbauweg für intrazelluläre Proteine in eukaryotischen Zellen dar. Das abzubauende Protein wird zunächst über eine Enzym-Kaskade mit einer kovalent gebundenen Ubiquitinkette markiert. Anschließend wird das konjugierte Substrat vom Proteasom erkannt und proteolytisch gespalten. Ubiquitin besitzt eine Reihe von Homologen, die ebenfalls posttranslational an Proteine gekoppelt werden können, wie z.B. SUMO und NEDD8. Die hierbei verwendeten Aktivierungs- und Konjugations-Kaskaden sind vollständig analog zu der des Ubiquitin- Systems. Es ist charakteristisch für das UPS, daß sich die Vielzahl der daran beteiligten Proteine aus nur wenigen Proteinfamilien rekrutiert, die durch gemeinsame, funktionale Homologiedomänen gekennzeichnet sind. Einige dieser funktionalen Domänen sind auch in den Modifikations-Systemen der Ubiquitin-Homologen zu finden, jedoch verfügen diese Systeme zusätzlich über spezifische Domänentypen. Homologiedomänen lassen sich als mathematische Modelle in Form von Domänen- deskriptoren (Profile) beschreiben. Diese Deskriptoren können wiederum dazu verwendet werden, mit Hilfe geeigneter Verfahren eine gegebene Proteinsequenz auf das Vorliegen von entsprechenden Homologiedomänen zu untersuchen. Da die im UPS involvierten Homologie- domänen fast ausschließlich auf dieses System und seine Analoga beschränkt sind, können domänen-spezifische Profile zur Katalogisierung der UPS-relevanten Proteine einer Spezies verwendet werden. Auf dieser Basis können dann die entsprechenden UPS-Repertoires verschiedener Spezies miteinander verglichen werden.