Induction of P57 Expression by P73ß
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The P53/P73 - P21cip1 Tumor Suppressor Axis Guards Against Chromosomal Instability by Restraining CDK1 in Human Cancer Cells
Oncogene (2021) 40:436–451 https://doi.org/10.1038/s41388-020-01524-4 ARTICLE The p53/p73 - p21CIP1 tumor suppressor axis guards against chromosomal instability by restraining CDK1 in human cancer cells 1 1 2 1 2 Ann-Kathrin Schmidt ● Karoline Pudelko ● Jan-Eric Boekenkamp ● Katharina Berger ● Maik Kschischo ● Holger Bastians 1 Received: 2 July 2020 / Revised: 2 October 2020 / Accepted: 13 October 2020 / Published online: 9 November 2020 © The Author(s) 2020. This article is published with open access Abstract Whole chromosome instability (W-CIN) is a hallmark of human cancer and contributes to the evolvement of aneuploidy. W-CIN can be induced by abnormally increased microtubule plus end assembly rates during mitosis leading to the generation of lagging chromosomes during anaphase as a major form of mitotic errors in human cancer cells. Here, we show that loss of the tumor suppressor genes TP53 and TP73 can trigger increased mitotic microtubule assembly rates, lagging chromosomes, and W-CIN. CDKN1A, encoding for the CDK inhibitor p21CIP1, represents a critical target gene of p53/p73. Loss of p21CIP1 unleashes CDK1 activity which causes W-CIN in otherwise chromosomally stable cancer cells. fi Vice versa 1234567890();,: 1234567890();,: Consequently, induction of CDK1 is suf cient to induce abnormal microtubule assembly rates and W-CIN. , partial inhibition of CDK1 activity in chromosomally unstable cancer cells corrects abnormal microtubule behavior and suppresses W-CIN. Thus, our study shows that the p53/p73 - p21CIP1 tumor suppressor axis, whose loss is associated with W-CIN in human cancer, safeguards against chromosome missegregation and aneuploidy by preventing abnormally increased CDK1 activity. -

P14ARF Inhibits Human Glioblastoma–Induced Angiogenesis by Upregulating the Expression of TIMP3
P14ARF inhibits human glioblastoma–induced angiogenesis by upregulating the expression of TIMP3 Abdessamad Zerrouqi, … , Daniel J. Brat, Erwin G. Van Meir J Clin Invest. 2012;122(4):1283-1295. https://doi.org/10.1172/JCI38596. Research Article Oncology Malignant gliomas are the most common and the most lethal primary brain tumors in adults. Among malignant gliomas, 60%–80% show loss of P14ARF tumor suppressor activity due to somatic alterations of the INK4A/ARF genetic locus. The tumor suppressor activity of P14ARF is in part a result of its ability to prevent the degradation of P53 by binding to and sequestering HDM2. However, the subsequent finding of P14ARF loss in conjunction with TP53 gene loss in some tumors suggests the protein may have other P53-independent tumor suppressor functions. Here, we report what we believe to be a novel tumor suppressor function for P14ARF as an inhibitor of tumor-induced angiogenesis. We found that P14ARF mediates antiangiogenic effects by upregulating expression of tissue inhibitor of metalloproteinase–3 (TIMP3) in a P53-independent fashion. Mechanistically, this regulation occurred at the gene transcription level and was controlled by HDM2-SP1 interplay, where P14ARF relieved a dominant negative interaction of HDM2 with SP1. P14ARF-induced expression of TIMP3 inhibited endothelial cell migration and vessel formation in response to angiogenic stimuli produced by cancer cells. The discovery of this angiogenesis regulatory pathway may provide new insights into P53-independent P14ARF tumor-suppressive mechanisms that have implications for the development of novel therapies directed at tumors and other diseases characterized by vascular pathology. Find the latest version: https://jci.me/38596/pdf Research article P14ARF inhibits human glioblastoma–induced angiogenesis by upregulating the expression of TIMP3 Abdessamad Zerrouqi,1 Beata Pyrzynska,1,2 Maria Febbraio,3 Daniel J. -

Deciphering the Nature of Trp73 Isoforms in Mouse
cancers Article Deciphering the Nature of Trp73 Isoforms in Mouse Embryonic Stem Cell Models: Generation of Isoform-Specific Deficient Cell Lines Using the CRISPR/Cas9 Gene Editing System Lorena López-Ferreras 1,2,†, Nicole Martínez-García 1,3,†, Laura Maeso-Alonso 1,2,‡, Marta Martín-López 1,4,‡, Ángela Díez-Matilla 1,‡ , Javier Villoch-Fernandez 1,2, Hugo Alonso-Olivares 1,2, Margarita M. Marques 3,5,* and Maria C. Marin 1,2,* 1 Instituto de Biomedicina (IBIOMED), Universidad de León, 24071 León, Spain; [email protected] (L.L.-F.); [email protected] (N.M.-G.); [email protected] (L.M.-A.); [email protected] (M.M.-L.); [email protected] (Á.D.-M.); [email protected] (J.V.-F.); [email protected] (H.A.-O.) 2 Departamento de Biología Molecular, Universidad de León, 24071 León, Spain 3 Departamento de Producción Animal, Universidad de León, 24071 León, Spain 4 Biomar Microbial Technologies, Parque Tecnológico de León, Armunia, 24009 León, Spain 5 Instituto de Desarrollo Ganadero y Sanidad Animal (INDEGSAL), Universidad de León, 24071 León, Spain * Correspondence: [email protected] (M.M.M.); [email protected] (M.C.M.); Tel.: +34-987-291757 Citation: López-Ferreras, L.; (M.M.M.); +34-987-291490 (M.C.M.) Martínez-García, N.; Maeso-Alonso, † Equal contribution. ‡ Equal contribution. L.; Martín-López, M.; Díez-Matilla, Á.; Villoch-Fernandez, J.; Simple Summary: The Trp73 gene is involved in the regulation of multiple biological processes Alonso-Olivares, H.; Marques, M.M.; Marin, M.C. Deciphering the Nature such as response to stress, differentiation and tissue architecture. -

TP63 Is Significantly Upregulated in Diabetic Kidney
International Journal of Molecular Sciences Article TP63 Is Significantly Upregulated in Diabetic Kidney Sitai Liang 1, Bijaya K. Nayak 1, Kristine S. Vogel 1 and Samy L. Habib 1,2,* 1 Department of Cell Systems and Anatomy, The University of Texas Health Science Center, San Antonio, TX 78229, USA; [email protected] (S.L.); [email protected] (B.K.N.); [email protected] (K.S.V.) 2 South Texas, Veterans Healthcare System, San Antonio, TX 78229, USA * Correspondence: [email protected]; Tel.: +1-21-0567-3816; Fax: +1-21-0567-3802 Abstract: The role of tumor protein 63 (TP63) in regulating insulin receptor substrate 1 (IRS-1) and other downstream signal proteins in diabetes has not been characterized. RNAs extracted from kidneys of diabetic mice (db/db) were sequenced to identify genes that are involved in kidney complications. RNA sequence analysis showed more than 4- to 6-fold increases in TP63 expression in the diabetic mice’s kidneys, compared to wild-type mice at age 10 and 12 months old. In addition, the kidneys from diabetic mice showed significant increases in TP63 mRNA and protein expression compared to WT mice. Mouse proximal tubular cells exposed to high glucose (HG) for 48 h showed significant decreases in IRS-1 expression and increases in TP63, compared to cells grown in normal glucose (NG). When TP63 was downregulated by siRNA, significant increases in IRS-1 and activation of AMP-activated protein kinase (AMPK (p-AMPK-Th172)) occurred under NG and HG conditions. Moreover, activation of AMPK by pretreating the cells with AICAR resulted in significant down- regulation of TP63 and increased IRS-1 expression. -

Are Interactions with P63 and P73 Involved in Mutant P53 Gain of Oncogenic Function?
Oncogene (2007) 26, 2220–2225 & 2007 Nature Publishing Group All rights reserved 0950-9232/07 $30.00 www.nature.com/onc REVIEW Are interactions with p63 and p73 involved in mutant p53 gain of oncogenic function? Y Li and C Prives Department of Biological Sciences, Columbia University, New York, NY, USA Although still controversial, the presence of mutant p53 in deletion or frameshift mutations resulting in their lack cancer cells mayresult in more aggressive tumors and of expression, however, many tumor-derived p53 muta- correspondinglyworse outcomes. The means bywhich tions are of the missense variety. These mutations are mutant p53 exerts such pro-oncogenic activityare usually located within the central conserved region of currentlyunder extensive investigation and different p53 that binds to DNA in a sequence-specific manner models have been proposed. We focus here on a proposed and result in either direct loss of DNA contact or gross mechanism bywhich a subset of tumor-derived p53 mut- change of the conformation that no longer permits p53 ants physically interact with p53 family members, p63 and to recognize its wild-type DNA targets (reviewed by p73, and negativelyregulate their proapoptotic function. Vousden and Lu, 2002). Approximately 30% of these Both cell-based assays and knock-in mice expressing tumor-derived p53 mutations affect only six ‘hot-spot’ mutant forms of p53 support this model. As more than half residues (175, 245, 248, 249, 273 and 282). In many cases of human tumors harbor mutant forms of p53 protein, tumor-derived p53 mutants are expressed at far higher approaches aimed at disrupting the pathological interac- levels than wild-type p53 protein due in part to their tions among p53 familymembers might be of clinical failure to be degraded by Mdm2. -

Alterations of P14arf, P53, and P73 Genes Involved in the E2F-1-Mediated Apoptotic Pathways in Non-Small Cell Lung Carcinoma
[CANCER RESEARCH 61, 5636–5643, July 15, 2001] Alterations of p14ARF, p53, and p73 Genes Involved in the E2F-1-mediated Apoptotic Pathways in Non-Small Cell Lung Carcinoma Siobhan A. Nicholson,1 Nader T. Okby,1 Mohammed A. Khan, Judith A. Welsh, Mary G. McMenamin, William D. Travis, James R. Jett, Henry D. Tazelaar, Victor Trastek, Peter C. Pairolero, Paul G. Corn, James G. Herman, Lance A. Liotta, Neil E. Caporaso, and Curtis C. Harris2 Laboratory of Human Carcinogenesis, National Cancer Institute, Bethesda, Maryland 20892 [S. A. N., M. A. K., J. A. W., M. G. M., L. A. L., N. E. C., C. C. H.]; Orange Pathology Associates, Middleton, New York 10940 [N. T. O.]; Armed Forces Institute of Pathology, Washington, DC 20306 [S. A. N., W. D. T.]; Mayo Clinic, Rochester, Minnesota 55905 [J. R. J., H. D. T., V. T., P. C. P.]; and The Johns Hopkins Oncology Center, Baltimore, Maryland 21231 [P. G. C., J. G. H.] ABSTRACT encoded by a separate exon 1 that lies ϳ20 kb upstream of exon 1␣ and shares exons 2 and 3 as read in an ARF, giving rise to a protein ARF Overexpression of E2F-1 induces apoptosis by both a p14 -p53- and INK4a ARF completely unrelated to p16 (14). Despite its unrelated structure, a p73-mediated pathway. p14 is the alternate tumor suppressor prod- ARF p14 also is capable of causing cell cycle arrest in G1 and G2. uct of the INK4a/ARF locus that is inactivated frequently in lung carci- ARF nogenesis. Because p14ARF stabilizes p53, it has been proposed that the loss p14 binds to and antagonizes the actions of MDM2, a negative of p14ARF is functionally equivalent to a p53 mutation. -

Regulation of P27kip1 and P57kip2 Functions by Natural Polyphenols
biomolecules Review Regulation of p27Kip1 and p57Kip2 Functions by Natural Polyphenols Gian Luigi Russo 1,* , Emanuela Stampone 2 , Carmen Cervellera 1 and Adriana Borriello 2,* 1 National Research Council, Institute of Food Sciences, 83100 Avellino, Italy; [email protected] 2 Department of Precision Medicine, University of Campania “Luigi Vanvitelli”, 81031 Napoli, Italy; [email protected] * Correspondence: [email protected] (G.L.R.); [email protected] (A.B.); Tel.: +39-0825-299-331 (G.L.R.) Received: 31 July 2020; Accepted: 9 September 2020; Published: 13 September 2020 Abstract: In numerous instances, the fate of a single cell not only represents its peculiar outcome but also contributes to the overall status of an organism. In turn, the cell division cycle and its control strongly influence cell destiny, playing a critical role in targeting it towards a specific phenotype. Several factors participate in the control of growth, and among them, p27Kip1 and p57Kip2, two proteins modulating various transitions of the cell cycle, appear to play key functions. In this review, the major features of p27 and p57 will be described, focusing, in particular, on their recently identified roles not directly correlated with cell cycle modulation. Then, their possible roles as molecular effectors of polyphenols’ activities will be discussed. Polyphenols represent a large family of natural bioactive molecules that have been demonstrated to exhibit promising protective activities against several human diseases. Their use has also been proposed in association with classical therapies for improving their clinical effects and for diminishing their negative side activities. The importance of p27Kip1 and p57Kip2 in polyphenols’ cellular effects will be discussed with the aim of identifying novel therapeutic strategies for the treatment of important human diseases, such as cancers, characterized by an altered control of growth. -
Beyond Traditional Morphological Characterization of Lung
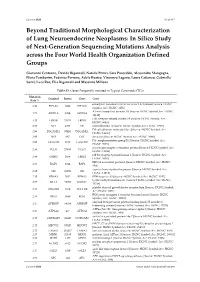
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -

P63 and P73 Isoform Expression in Non-Small Cell Lung Cancer and Corresponding Morphological Normal Lung Tissue
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector ORIGINAL ARTICLE p63 and p73 Isoform Expression in Non-small Cell Lung Cancer and Corresponding Morphological Normal Lung Tissue Marco Lo Iacono, PhD, Valentina Monica, MS, Silvia Saviozzi, PhD, Paolo Ceppi, MS, Enrico Bracco, PhD, Mauro Papotti, MD, and Giorgio V. Scagliotti, MD he tumor suppressor p53 is a key transcription factor Background: The TP73 and TP63 genes are members of the p53 regulating the expression of genes influencing cell senes- tumor suppressor family and are expressed in different N-terminal T cence, proliferation, apoptosis, and differentiation. The TP73 isoforms either with proapoptotic (transactivation domain, TA) and and TP63 genes are members of the p53 tumor suppressor antiapoptotic (N-terminally truncated, ⌬N) function. Unlike p53, the family, based on substantial structural and functional homol- role of p73 and p63 in tumor is controversial. It has been recently ogies. Unlike p53, that is a tumor suppressor, the role of p73 hypothesized that altered ⌬N:TA expression ratio, rather than single and p63 in tumor is controversial because of their genomic isoform overexpression, plays a role in the pathogenesis of many locus complexity. Through either alternative exon splicing, or diseases, including lung cancer. a second promoter, TP73 and TP63 genes generate several N- Methods: Isoform-specific, real-time polymerase chain reaction and and C-terminal isoforms. The N-terminal isoforms can be immunohistochemistry analysis on matched cancer and correspond- clustered into two groups: the transactivation competent TA ing normal tissues from surgically resected non-small cell lung proteins (TAp63 and TAp73) and the transactivation-defec- cancers (NSCLCs) have been performed aiming to explore the tive, N-terminally truncated ⌬N proteins (⌬Np63, ⌬NЈp73, expression levels of each p63 and p73 N-terminal isoforms and their ⌬2p73, ⌬2/3p73, and ⌬Np73).1 TAp63 and TAp73, and p53, ⌬N:TA expression ratio. -

The Potential Tumor Suppressor P73 Differentially Regulates Cellular P53 Target Genes1
[CANCER RESEARCH 58, 5061-5065. November 15. 1998] Advances in Brief The Potential Tumor Suppressor p73 Differentially Regulates Cellular p53 Target Genes1 Jianhui Zhu,2 Jieyuan Jiang,2 Wenjing Zhou, and Xinbin Chen3 Institute of Molecular Medicine and Genetics, Medical College of Georgia. Augusta, Georgia 30912 Abstract activation and DNA binding domains between p53 and p73, the p53 and p73 signaling pathways leading to tumor suppression are both p73, a potential tumor suppressor, is a p53 homologue. Transient over similar and different. expression of p73 in cells can induce apoptosis and p21, a cellular p53 target gene primarily responsible for p53-dependent cell cycle arrest. To further characterize the role of p73 in tumor suppression, we established Materials and Methods several groups of cell lines that inducibly express p73 under a tetracycline- regulated promoter. By using these cell lines, we found that p73 can Plasmids. cDNAs for p73a. p73ß.and p73«292 (Ref. 3; kindly provided by W. Kaelin) were cloned separately into a tetraeycline-regulated expression induce both cell cycle arrest and apoptosis. We also found that p73 can vector. 10-3, at its Eco RI and Xba I siles, and the resulting plasmids were used activate some but not all of the previously identified p53 cellular target genes. Furthermore, we found that the transcriptional activities of p53, to generate cell lines that inducibly express p73. p73 proteins were tagged at p73a, and p73/3 to induce their common cellular target genes differ among their N-termini with influenza hemagglutinin peptide. one another. These results suggest that p73 is both similar to and different Generation of H1299 Cell Lines that Inducibly Express p73. -

P73 Overexpression Is Associated with Resistance to Treatment With
[CANCER RESEARCH 61, 935–938, February 1, 2001] Advances in Brief p73␣ Overexpression Is Associated with Resistance to Treatment with DNA-damaging Agents in a Human Ovarian Cancer Cell Line1 Faina Vikhanskaya,2 Sergio Marchini,2 Mirko Marabese, Emanuela Galliera, and Massimo Broggini3 Laboratory of Molecular Pharmacology, Department of Oncology, Istituto di Ricerche Farmacologiche “Mario Negri,” 20157 Milan, Italy Abstract changes in gene expression and the response of these clones to different chemical and physical DNA-damaging agents. We examined the consequences of p73␣ overexpression on gene expres- sion and cellular response to anticancer agents in clones from the human Materials and Methods ovarian cancer cell line A2780. Using microarray filters, the expression of ␣ 588 genes in two clones overexpressing p73 (A2780/p73.4 and A2780/ Cell Culture and Treatment. Clones derived from the human ovarian p73.5) in comparison with empty vector-transfected cells was evaluated. carcinoma cell lines A2780, A2780/pCDNA3, A2780/p73.4, and A2780/p73.5 There were clear differences in gene expression profiles. Both of the clones were obtained and grown as reported previously (12). DDP4 was obtained from showed a marked increase in the expression of genes involved in DNA Bristol Myers Squibb (Syracuse, NY). Topotecan was obtained from Smith, repair, including genes participating in nucleotide excision repair and Kline & Beecham (Brentford, United Kingdom) and MNNG was purchased mismatch repair. This was confirmed by reverse transcription-PCR and from Sigma (Milan, Italy). For clonogenic assay, cells were plated at 250 Northern blot analysis and was associated with an increase in the ability cells/well. -

Structure and Apoptotic Function of P73
BMB Rep. 2015; 48(2): 81-90 BMB www.bmbreports.org Reports Invited Mini Review Structure and apoptotic function of p73 Mi-Kyung Yoon1, Ji-Hyang Ha1, Min-Sung Lee1,2 & Seung-Wook Chi1,2,* 1Structural Biology & Nanopore Research Laboratory, Functional Genomics Research Center, KRIBB, Daejeon 305-806, 2Department of Bio-Analytical Science, University of Science and Technology, Daejeon 305-350, Korea p73 is a structural and functional homologue of the p53 tumor cers in which p53 is mutated or inactive. suppressor protein. Like p53, p73 induces apoptosis and cell More than 10 p73 variants have been identified, generated cycle arrest and transactivates p53-responsive genes, confer- using two promoters at the N-terminus and alternative splicing ring its tumor suppressive activity. In addition, p73 has unique at the C-terminus (4). Among these, however, p73 exists as roles in neuronal development and differentiation. The im- two major isoforms with opposing activities; the pro-apoptotic portance of p73-induced apoptosis lies in its capability to sub- transcriptionally active (TA) isoform and the anti-apoptotic stitute the pro-apoptotic activity of p53 in various human can- N-terminal transactivation domain (TAD)-truncated delta-N cer cells in which p53 is mutated or inactive. Despite the great (DN) isoform (5). The DNp73 isoform is dominant negative to- importance of p73-induced apoptosis in cancer therapy, little ward TAp73 and wild-type p53, thus inhibiting pro-apoptotic is known about the molecular basis of p73-induced apoptosis. activities. In this review, we discuss the p73 structures reported to date, The stability and activity of p73 are primarily regulated by detailed structural comparisons between p73 and p53, and cur- post-translational modifications (PTMs).