CEACAM1 As a Multi-Purpose Target for Cancer Immunotherapy
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Screening and Identification of Key Biomarkers in Clear Cell Renal Cell Carcinoma Based on Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Screening and identification of key biomarkers in clear cell renal cell carcinoma based on bioinformatics analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Clear cell renal cell carcinoma (ccRCC) is one of the most common types of malignancy of the urinary system. The pathogenesis and effective diagnosis of ccRCC have become popular topics for research in the previous decade. In the current study, an integrated bioinformatics analysis was performed to identify core genes associated in ccRCC. An expression dataset (GSE105261) was downloaded from the Gene Expression Omnibus database, and included 26 ccRCC and 9 normal kideny samples. Assessment of the microarray dataset led to the recognition of differentially expressed genes (DEGs), which was subsequently used for pathway and gene ontology (GO) enrichment analysis. -

Propranolol-Mediated Attenuation of MMP-9 Excretion in Infants with Hemangiomas
Supplementary Online Content Thaivalappil S, Bauman N, Saieg A, Movius E, Brown KJ, Preciado D. Propranolol-mediated attenuation of MMP-9 excretion in infants with hemangiomas. JAMA Otolaryngol Head Neck Surg. doi:10.1001/jamaoto.2013.4773 eTable. List of All of the Proteins Identified by Proteomics This supplementary material has been provided by the authors to give readers additional information about their work. © 2013 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 10/01/2021 eTable. List of All of the Proteins Identified by Proteomics Protein Name Prop 12 mo/4 Pred 12 mo/4 Δ Prop to Pred mo mo Myeloperoxidase OS=Homo sapiens GN=MPO 26.00 143.00 ‐117.00 Lactotransferrin OS=Homo sapiens GN=LTF 114.00 205.50 ‐91.50 Matrix metalloproteinase‐9 OS=Homo sapiens GN=MMP9 5.00 36.00 ‐31.00 Neutrophil elastase OS=Homo sapiens GN=ELANE 24.00 48.00 ‐24.00 Bleomycin hydrolase OS=Homo sapiens GN=BLMH 3.00 25.00 ‐22.00 CAP7_HUMAN Azurocidin OS=Homo sapiens GN=AZU1 PE=1 SV=3 4.00 26.00 ‐22.00 S10A8_HUMAN Protein S100‐A8 OS=Homo sapiens GN=S100A8 PE=1 14.67 30.50 ‐15.83 SV=1 IL1F9_HUMAN Interleukin‐1 family member 9 OS=Homo sapiens 1.00 15.00 ‐14.00 GN=IL1F9 PE=1 SV=1 MUC5B_HUMAN Mucin‐5B OS=Homo sapiens GN=MUC5B PE=1 SV=3 2.00 14.00 ‐12.00 MUC4_HUMAN Mucin‐4 OS=Homo sapiens GN=MUC4 PE=1 SV=3 1.00 12.00 ‐11.00 HRG_HUMAN Histidine‐rich glycoprotein OS=Homo sapiens GN=HRG 1.00 12.00 ‐11.00 PE=1 SV=1 TKT_HUMAN Transketolase OS=Homo sapiens GN=TKT PE=1 SV=3 17.00 28.00 ‐11.00 CATG_HUMAN Cathepsin G OS=Homo -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

The Regulation of Interleukin 7 Receptor Alpha Internalization, Recycling and Degradation by IL-7
Universidade de Lisboa Faculdade de Medicina Unidade de Biologia do Cancro, Instituto de Medicina Molecular The regulation of Interleukin 7 receptor alpha internalization, recycling and degradation by IL-7 - Possible implications in T-cell homeostasis, migration and leukaemogenesis - Catarina Martins de Oliveira Henriques Doutoramento em Ciências Biomédicas For the degree of Doctor of Philosophy 2009 Universidade de Lisboa Faculdade de Medicina Unidade de Biologia do Cancro, Instituto de Medicina Molecular The regulation of Interleukin 7 receptor alpha internalization, recycling and degradation by IL-7 - Possible implications in T-cell homeostasis, migration and leukaemogenesis - Catarina Martins de Oliveira Henriques (Recipient of a scholarship- SFRH7BD/21940/2005 from Fundação para a Ciência e Tecnologia) Tese orientada pelo Doutor João T. Barata, Prof Doutor.Gerard Graham e Prof. Doutora Leonor Parreira Doutoramento em Ciências Biomédicas, especialidade em Ciências Biopatológicas For the degree of Doctor of Philosophy 2009 As opiniões expressas são da exclusiva responsabilidade do seu autor A impressão desta dissertação foi aprovada pela Comissão Coordenadora do Conselho Científico da Faculdade de Medicina de Lisboa em reunião de 13 de Outubro de 2009. Para a Prof. Filomena Mota. Sem o seu apoio e inspiração há muitos anos atrás, eu não seria hoje uma bióloga nem esta tese teria alguma vez existido… Table of Contents Table of contents……………………………………………………….………………….................. i Aknowledgements……………………………………………………………………………………………. -

Combination Immune Therapy
Combination Immune Therapy Translational Medicine Plenary SWOG April 17, 2017 San Francisco T‐cell Activation, Proliferation, and Function is Controlled by Multiple Agonist and Antagonist Signals 1. Co‐stimulation via CD28 ligation 2. CTLA‐4 ligation on activated T 3. T cell function in tissue is subject transduces T cell activating signals cells down‐regulates T cell to feedback inhibition responses T cell Functional Block T cell Activation T cell Activation T cell Proliferative Block T cell T cell APC T cell CTLA‐4 T cell CTLA‐4 TCR TCR PD‐1 PD‐1 PD‐1 IFN‐gamma TCR CD28 Cytokines TCR CD4 MHC MHC PD‐‐L1 MHC B7 MHC CD28 B7 PD‐‐L1 Tumor or Tumor or APC APC Immune cell B7 immune cell APC NK Presence of PD-L1 or TILs1 PPD-L1 /TIL+ PD-L1/TIL + + PD-L1 /TIL+ + PD-L1 /TIL PD-L1 /TIL D-L1 /TIL+ Schalper and Rimm, Yale University NSCLC PD-L1/TIL 45% 17% 26% 12% Type 1 Type 2 Type 3 Type 4 45% 41% 13% 1% Melanoma45% 17% 26% 12% Taube et al Type 1 Type 2 Type 3 Type 4 Tumor‐specific T cells are contained in the PD‐1+ TIL population and are functional after in vitro culture Spectrum of PD‐1/PD‐L1 Antagonist Activity Active • Melanoma Minimal to no activity • Renal cancer (clear cell and non‐clear cell) • Prostate cancer • NSCLC – adenocarcinoma and squamous cell • MMR+ (MSS) colon cancer • Small cell lung cancer • Myeloma • Head and neck cancer • Pancreatic cancer • Gastric and gastroesophageal junction • MMR‐repair deficient tumors (colon, cholangiocarcinoma) • Bladder • Triple negative breast cancer • Ovarian Major PD‐1/PD‐L1 antagonists • Hepatocellular -
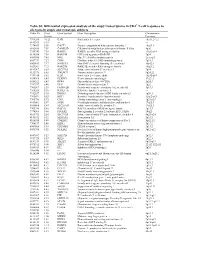
Table S1| Differential Expression Analysis of the Atopy Transcriptome
Table S1| Differential expression analysis of the atopy transcriptome in CD4+ T-cell responses to allergens in atopic and nonatopic subjects Probe ID S.test Gene Symbol Gene Description Chromosome Statistic Location 7994280 10.32 IL4R Interleukin 4 receptor 16p11.2-12.1 8143383 8.95 --- --- --- 7974689 8.50 DACT1 Dapper, antagonist of beta-catenin, homolog 1 14q23.1 8102415 7.59 CAMK2D Calcium/calmodulin-dependent protein kinase II delta 4q26 7950743 7.58 RAB30 RAB30, member RAS oncogene family 11q12-q14 8136580 7.54 RAB19B GTP-binding protein RAB19B 7q34 8043504 7.45 MAL Mal, T-cell differentiation protein 2cen-q13 8087739 7.27 CISH Cytokine inducible SH2-containing protein 3p21.3 8000413 7.17 NSMCE1 Non-SMC element 1 homolog (S. cerevisiae) 16p12.1 8021301 7.15 RAB27B RAB27B, member RAS oncogene family 18q21.2 8143367 6.83 SLC37A3 Solute carrier family 37 member 3 7q34 8152976 6.65 TMEM71 Transmembrane protein 71 8q24.22 7931914 6.56 IL2R Interleukin 2 receptor, alpha 10p15-p14 8014768 6.43 PLXDC1 Plexin domain containing 1 17q21.1 8056222 6.43 DPP4 Dipeptidyl-peptidase 4 (CD26) 2q24.3 7917697 6.40 GFI1 Growth factor independent 1 1p22 7903507 6.39 FAM102B Family with sequence similarity 102, member B 1p13.3 7968236 5.96 RASL11A RAS-like, family 11, member A --- 7912537 5.95 DHRS3 Dehydrogenase/reductase (SDR family) member 3 1p36.1 7963491 5.83 KRT1 Keratin 1 (epidermolytic hyperkeratosis) 12q12-q13 7903786 5.72 CSF1 Colony stimulating factor 1 (macrophage) 1p21-p13 8019061 5.67 SGSH N-sulfoglucosamine sulfohydrolase (sulfamidase) 17q25.3 -

Bioinformatics Method Identifies Potential Biomarkers of Dilated Cardiomyopathy in a Human Induced Pluripotent Stem Cell‑Derived Cardiomyocyte Model
EXPERIMENTAL AND THERAPEUTIC MEDICINE 14: 2771-2778, 2017 Bioinformatics method identifies potential biomarkers of dilated cardiomyopathy in a human induced pluripotent stem cell‑derived cardiomyocyte model YU ZHUANG1, YU-JIA GONG2, BEI-FEN ZHONG3, YI ZHOU1 and LI GONG4 1Department of Cardiovascular Surgery, Shanghai First People's Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai 200080; 2Stomatology Faculty, School of Medicine, Nantong University, Nantong, Jiangsu 226000; 3Department of Urology, Shanghai First People's Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai 200080; 4Department of Cardiothoracic Surgery, The Affiliated Huai'an Hospital of Xuzhou Medical University, Huai'an, Jiangsu 223002, P.R. China Received February 19, 2016; Accepted February 10, 2017 DOI: 10.3892/etm.2017.4850 Abstract. Dilated cardiomyopathy (DCM) is the most common crucial nodes in module 2, which were linked to each other. type of cardiomyopathy that account for the majority of heart In conclusion, several potential biomarkers for DCM were failure cases. The present study aimed to reveal the under- identified, such as MMP2, FLT1, CDH1, ITGB6, COL6A3, lying molecular mechanisms of DCM and provide potential COL6A1, LAMC2, PENK and APLNR. These genes may serve biomarkers for detection of this condition. The public dataset significant roles in DCM via involvement of various BPs, such of GSE35108 was downloaded, and 4 normal induced pluripo- as blood vessel and vasculature development and cell adhe- tent stem cell (iPSC)-derived cardiomyocytes (N samples) and sion, and the ECM-receptor interaction pathway. 4 DCM iPSC-derived cardiomyocytes (DCM samples) were utilized. Raw data were preprocessed, followed by identifica- Introduction tion of differentially expressed genes (DEGs) between N and DCM samples. -

An Attempt to Polarize Human Neutrophils Toward N1 and N2 Phenotypes in Vitro
fimmu-11-00532 April 24, 2020 Time: 17:59 # 1 ORIGINAL RESEARCH published: 28 April 2020 doi: 10.3389/fimmu.2020.00532 An Attempt to Polarize Human Neutrophils Toward N1 and N2 Phenotypes in vitro Mareike Ohms, Sonja Möller and Tamás Laskay* Department of Infectious Diseases and Microbiology, University of Lübeck, Lübeck, Germany Neutrophils act as the first line of defense against invading pathogens. Although traditionally considered in context of their antimicrobial effector functions, the importance of tumor-associated neutrophils (TANs) in the development of cancer has become increasingly clear during the last decade. With regard to their high plasticity, neutrophils were shown to acquire an anti-tumorigenic N1 or a pro-tumorigenic N2 phenotype. Despite the urgent need to get a comprehensive understanding of the interaction of TANs with their tumor microenvironment, most studies still rely on murine tumor models. Here we present for the first time a polarization attempt to generate N1 and N2 neutrophils from primary human neutrophils in vitro. Our results underscore Edited by: that N1-polarized neutrophils have a pro-inflammatory phenotype characterized among Martin Herrmann, University Hospital Erlangen, Germany others by a higher level of intercellular adhesion molecule (ICAM)-1 and high secretion Reviewed by: of interferon (IFN)g-induced protein 10 (IP-10)/C-X-C motif chemokine 10 (CXCL10) Payel Sil, and tumor necrosis factor (TNF). Further, we demonstrate that neutrophils incubated National Institute of Environmental under a tumor-mimicking in vitro environment show a high cell surface expression of Health Sciences (NIEHS), United States C-X-C motif chemokine receptor 2 (CXCR2) and secrete high levels of interleukin (IL)- Mihaela Gadjeva, 8. -

SUPPLEMENTARY METHODS Cell Culture.-Human Peripheral Blood
SUPPLEMENTARY METHODS Cell culture.-Human peripheral blood mononuclear cells (PBMC) were isolated from buffy coats from normal donors over a Lymphoprep (Nycomed Pharma) gradient. Monocytes were purified from PBMC by magnetic cell sorting using CD14 microbeads (Miltenyi Biotech). Monocytes were cultured at 0.5 x 106 cells/ml for 7 days in RPMI 1640 (standard RPMI, which contains 1 mg/L folic acid) supplemented with 10% fetal calf serum, at 37ºC in a humidified atmosphere with 5% CO2, and containing GM-CSF (1000U/ml) or M-CSF (10 ng/ml, ImmunoTools) to generate GM-CSF- polarized macrophages (GM-MØ) or M-CSF-polarized macrophages (M-MØ). MTX pharmacokinetic studies in RA patients administered 25 mg MTX showed peak plasma levels of 1-2 µM MTX two hours after drug administration, but plasma levels decline to 10-50 nM MTX within 24-48 hours [1, 2]. MTX (50 nM), pemetrexed (PMX, 50 nM), folic acid (FA, 50 nM) [3], thymidine (dT, 10 µM), pifithrin-α (PFT, 25-50 µM), nutlin-3 (10 µM, Sigma-Aldrich) was added once on monocytes together with the indicated cytokine, or on monocytes and 7-day differentiated macrophages for 48h. Gene expression profiling.-For long-term MTX treatment, RNA was isolated from three independent preparations of monocytes either unexposed or exposed to MTX (50 nM) and differentiated to GM-MØ or M-MØ for 7-days. For short-term schedule, RNA was isolated from three independent samples of fully differentiated GM-MØ either unexposed or exposed to MTX (50 nM) for 48h, by using RNeasy Mini kit (QIAGEN). -

MALE Protein Name Accession Number Molecular Weight CP1 CP2 H1 H2 PDAC1 PDAC2 CP Mean H Mean PDAC Mean T-Test PDAC Vs. H T-Test
MALE t-test t-test Accession Molecular H PDAC PDAC vs. PDAC vs. Protein Name Number Weight CP1 CP2 H1 H2 PDAC1 PDAC2 CP Mean Mean Mean H CP PDAC/H PDAC/CP - 22 kDa protein IPI00219910 22 kDa 7 5 4 8 1 0 6 6 1 0.1126 0.0456 0.1 0.1 - Cold agglutinin FS-1 L-chain (Fragment) IPI00827773 12 kDa 32 39 34 26 53 57 36 30 55 0.0309 0.0388 1.8 1.5 - HRV Fab 027-VL (Fragment) IPI00827643 12 kDa 4 6 0 0 0 0 5 0 0 - 0.0574 - 0.0 - REV25-2 (Fragment) IPI00816794 15 kDa 8 12 5 7 8 9 10 6 8 0.2225 0.3844 1.3 0.8 A1BG Alpha-1B-glycoprotein precursor IPI00022895 54 kDa 115 109 106 112 111 100 112 109 105 0.6497 0.4138 1.0 0.9 A2M Alpha-2-macroglobulin precursor IPI00478003 163 kDa 62 63 86 72 14 18 63 79 16 0.0120 0.0019 0.2 0.3 ABCB1 Multidrug resistance protein 1 IPI00027481 141 kDa 41 46 23 26 52 64 43 25 58 0.0355 0.1660 2.4 1.3 ABHD14B Isoform 1 of Abhydrolase domain-containing proteinIPI00063827 14B 22 kDa 19 15 19 17 15 9 17 18 12 0.2502 0.3306 0.7 0.7 ABP1 Isoform 1 of Amiloride-sensitive amine oxidase [copper-containing]IPI00020982 precursor85 kDa 1 5 8 8 0 0 3 8 0 0.0001 0.2445 0.0 0.0 ACAN aggrecan isoform 2 precursor IPI00027377 250 kDa 38 30 17 28 34 24 34 22 29 0.4877 0.5109 1.3 0.8 ACE Isoform Somatic-1 of Angiotensin-converting enzyme, somaticIPI00437751 isoform precursor150 kDa 48 34 67 56 28 38 41 61 33 0.0600 0.4301 0.5 0.8 ACE2 Isoform 1 of Angiotensin-converting enzyme 2 precursorIPI00465187 92 kDa 11 16 20 30 4 5 13 25 5 0.0557 0.0847 0.2 0.4 ACO1 Cytoplasmic aconitate hydratase IPI00008485 98 kDa 2 2 0 0 0 0 2 0 0 - 0.0081 - 0.0 -

(Lilrs) on Human Neutrophils: Modulators of Infection and Immunity
MINI REVIEW published: 13 May 2020 doi: 10.3389/fimmu.2020.00857 Leukocyte Immunoglobulin-Like Receptors (LILRs) on Human Neutrophils: Modulators of Infection and Immunity Alexander L. Lewis Marffy and Alex J. McCarthy* MRC Centre for Molecular Bacteriology and Infection, Imperial College London, London, United Kingdom Neutrophils have a crucial role in defense against microbes. Immune receptors allow neutrophils to sense their environment, with many receptors functioning to recognize signs of infection and to promote antimicrobial effector functions. However, the neutrophil Edited by: Nicole Thielens, response must be tightly regulated to prevent excessive inflammation and tissue damage, UMR5075 Institut de Biologie and regulation is achieved by expression of inhibitory receptors that can raise activation Structurale (IBS), France thresholds. The leukocyte immunoglobulin-like receptor (LILR) family contain activating Reviewed by: and inhibitory members that can up- or down-regulate immune cell activity. New ligands Debby Burshtyn, University of Alberta, Canada and functions for LILR continue to emerge. Understanding the role of LILR in neutrophil Tamás Laskay, biology is of general interest as they can activate and suppress antimicrobial responses University of Lübeck, Germany of neutrophils and because several human pathogens exploit these receptors for immune *Correspondence: Alex J. McCarthy evasion. This review focuses on the role of LILR in neutrophil biology. We focus on the [email protected] current knowledge of LILR expression -

Neural Immunoglobulin Superfamily Interaction Networks
Available online at www.sciencedirect.com ScienceDirect Neural immunoglobulin superfamily interaction networks 1 ¨ 2 Kai Zinn and Engin Ozkan The immunoglobulin superfamily (IgSF) encompasses mediates protein–protein interactions, and many proteins hundreds of cell surface proteins containing multiple have both Ig domains and FnIII repeats. immunoglobulin-like (Ig) domains. Among these are neural IgCAMs, which are cell adhesion molecules that mediate Many cell surface IgSF proteins are homophilic or het- interactions between cells in the nervous system. IgCAMs in erophilic adhesion molecules (IgCAMs) containing mul- some vertebrate IgSF subfamilies bind to each other tiple Ig domains. IgCAMs mediate interactions among homophilically and heterophilically, forming small interaction neurons and between neurons and glia. Homophilic networks. In Drosophila, a global ‘interactome’ screen IgCAMs include NCAM and L1 in mammals and Fasci- identified two larger networks in which proteins in one IgSF clin II and Dscam in Drosophila. IgCAM-like proteins are subfamily selectively interact with proteins in a different receptors for axon guidance cues such as Netrins and Slits. subfamily. One of these networks, the ‘Dpr-ome’, includes Finally, neuronal transmembrane signal transduction 30 IgSF proteins, each of which is expressed in a unique subset molecules, such as receptor tyrosine kinases and phos- of neurons. Recent evidence shows that one interacting protein phatases, can have IgCAM-like extracellular (XC) pair within the Dpr-ome network is required for development of domains. the brain and neuromuscular system. This mini review does not cover homophilic IgCAMs, Addresses axon guidance receptors, or signaling receptors, but 1 Division of Biology and Biological Engineering, California Institute of focuses on subfamilies of the IgSF that participate in Technology, Pasadena, CA 91125, United States 2 complex networks of interactions.