Fe(II) Fe(III)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supporting Information
Supporting Information Lozupone et al. 10.1073/pnas.0807339105 SI Methods nococcus, and Eubacterium grouped with members of other Determining the Environmental Distribution of Sequenced Genomes. named genera with high bootstrap support (Fig. 1A). One To obtain information on the lifestyle of the isolate and its reported member of the Bacteroidetes (Bacteroides capillosus) source, we looked at descriptive information from NCBI grouped firmly within the Firmicutes. This taxonomic error was (www.ncbi.nlm.nih.gov/genomes/lproks.cgi) and other related not surprising because gut isolates have often been classified as publications. We also determined which 16S rRNA-based envi- Bacteroides based on an obligate anaerobe, Gram-negative, ronmental surveys of microbial assemblages deposited near- nonsporulating phenotype alone (6, 7). A more recent 16S identical sequences in GenBank. We first downloaded the gbenv rRNA-based analysis of the genus Clostridium defined phylo- files from the NCBI ftp site on December 31, 2007, and used genetically related clusters (4, 5), and these designations were them to create a BLAST database. These files contain GenBank supported in our phylogenetic analysis of the Clostridium species in the HGMI pipeline. We thus designated these Clostridium records for the ENV database, a component of the nonredun- species, along with the species from other named genera that dant nucleotide database (nt) where 16S rRNA environmental cluster with them in bootstrap supported nodes, as being within survey data are deposited. GenBank records for hits with Ͼ98% these clusters. sequence identity over 400 bp to the 16S rRNA sequence of each of the 67 genomes were parsed to get a list of study titles Annotation of GTs and GHs. -

26Sor Em2rray /Mmucd
D4006- Gut Microbiota Analysis UC Davis MMPC - Microbiome & Host Response Core Contents 1 Methods: 1 1.1 Sequencing . .1 1.2 Data processing . .1 2 Summary of Findings: 2 2.1 Sequencing analysis . .2 2.2 Microbial diversity . .2 2.2.1 Alpha Diversity . .2 2.2.2 Beta Diversity . 10 2.3 Data analysis using taxa abundance data . 13 2.3.1 Stacked bar graphs of Taxa abundances at each level . 14 A Appendix 1 (Taxa Abundance Tables) 30 B Appendix 2 (Taxa removed from filtered datasets at each level) 37 References: 43 Core Contacts: Helen E. Raybould, Ph.D., Core Leader ([email protected]) Trina A. Knotts, Ph.D., Core Co-Leader ([email protected]) Michael L. Goodson, Ph.D., Core Scientist ([email protected]) Client(s): Kent Lloyd, DVM PhD ;MMRRC; UC Davis Project #: MBP-2079 MMRRC strain ID: MMRRC_043514 Animal Information: The strain was donated to the MMRRC by Russell Ray at Baylor College of Medicine. Fecal samples were obtained from animals housed under the care of Russell Ray at Baylor College of Medicine. 1 Methods: Brief Project Description: MMRRC strains are often contributed to the MMRRC to fulfill the resource sharing aspects of NIH grants. Since transporting mice to another facilty often causes a microbiota shift, having a record of the original fecal microbiota from the donor institution where the original phenotyping or testing was performed may prove helpful if a phenotype is lost after transfer. Several MMRRC mouse lines were selected for fecal microbiota profiling of the microbiota. Table 1: Animal-Strain Information X.SampleID TreatmentGroup Animal_ID Genotype Line Sex MMRRC.043514.Mx054 MMRRC.043514_Het_M Mx054 Het MMRRC.043514 M MMRRC.043514.Mx415 MMRRC.043514_Het_M Mx415 Het MMRRC.043514 M 1 1.1 Sequencing Frozen fecal or regional gut samples were shipped on dry ice to UC Davis MMPC and Host Microbe Systems Biology Core. -

Supplementary Material Impacts of Radiation on the Bacterial And
Supplementary Material Impacts of radiation on the bacterial and fungal microbiome of small mammals in the Chernobyl Exclusion Zone Rachael E. Antwis, Nicholas A. Beresford, Joseph A. Jackson, Ross Fawkes, Catherine L. Barnett, Elaine Potter, Lee Walker, Sergey Gaschak, Michael D. Wood Table S1 Sample sizes of host species used in the study, and associated sex, total absorbed dose rate, burn and site categories for these samples. Animals estimated to receive total absorbed dose rates of <4 µGy h-1 were assigned ‘low’, those with estimated dose rates of 4-42 µGy h-1 assigned ‘medium’, and those >42 µGy h-1 assigned to the ‘high’ category. All samples from 2017 were collected within the Red Forest but from areas that had experienced different degrees of damage from forest fires. Samples from 2018 were either collected within or outside the Red Forest. Sampling information Sex Total absorbed dose category Burn category Site category Year Sample Host species Total n Female Male Unidentified Low Medium High Burnt Burnt Unburnt Inside Red Outside type (regrowth) (minimal Forest Red regrowth) Forest 2017 Faeces Bank vole 22 5 16 1 0 0 22 3 1 18 All - 2017 Faeces Striped field mouse 29 10 18 1 0 7 22 15 0 14 All - 2017 Faeces Wood mouse 27 6 20 1 0 0 27 2 25 0 All - 2017 Faeces Yellow-necked mouse 58 28 29 1 0 7 51 19 17 22 All - 2018 Gut Bank vole 142 59 83 0 42 64 36 - - - 34 108 (caecum) RESULTS Differences in community composition between host species and sample types Wood mice and yellow-necked mice had a similar bacterial community composition to one another (Figure 1a), with the exception of the family Helicobacteraceae (Z = 2.982, p = 0.023; Figure 1c). -
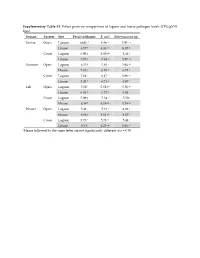
Supplementary Table S1. Fisher Pairwise Comparisons of Lagoon and House Pathogen Levels (CFU/Gvss Log10) Season System Site Fecal Coliforms E
Supplementary Table S1. Fisher pairwise comparisons of lagoon and house pathogen levels (CFU/gVSS log10) Season System Site Fecal coliforms E. coli Enterococcus sp. Spring Open Lagoon 6.00 g1 5.46 hi 5.91 cd House 6.57 e 6.33 cd 6.35 b Cover Lagoon 6.00 g 5.60 gh 5.44 f House 7.73 a 7.63 a 5.87 cd Summer Open Lagoon 6.35 f 5.83 f 5.94 cd House 7.83 a 6.10 e 6.73 a Cover Lagoon 7.04 c 6.47 c 5.86 cd House 7.42 b 6.71 b 6.07 c Fall Open Lagoon 5.58 i 5.58 gh 5.56 ef House 6.81 d 6.72 b 6.01 c Cover Lagoon 5.99 g 5.34 i 5.50 f House 6.38 f 6.19 de 5.74 de Winter Open Lagoon 5.41 j 5.13 j 4.88 g House 6.06 g 5.61 gh 6.67 a Cover Lagoon 5.73 h 5.73 fg 5.44 f House 6.34 f 6.25 de 5.86 cd 1Means followed by the same letter are not significantly different at p = 0.05 Supplementary Table S2. Relative abundances of OTUs identified using universal bacterial primer set, presented as relative abundances (%) Only bacterial families are counted in Figure 5 and discussion involving bacterial family identification. # Identified in # Identified from all 8 All 16 All #OTU Table SpOLRA SuOLRA FOLRA WOLRA SpCLRA SuCLRA FCLRA WCLRA SpOHRA SuOHRA FOHRA WOHRA SpCHRA SuCHRA FCHRA WCHRA all samples lagoon samples Samples Lagoons Notes Key Patulibacteraceae 0 9.93739E‐05* 0.000134 0.000116 0 0 0 0 0 0 0.000452 0 0 0 0 0 4 3 Sp = Spring O = Open Ruaniaceae 0 0 0 0 0 0 0 0 0 0 0 0 0 0.000221577 0 7.73E‐05 2 0 Su = Summer C = Covered Thermoactinomycetaceae 0 0 0.000267 0.000116 0 0.00037092 0.000529 0.000526 0 0.000342922 0.000271 0.000197 0 0 0 0 8 5 F = Fall L = Lagoon Beutenbergiaceae 0.000226334 0.000331246 0.000579 0.001663 0.000620176 0 0.000106 0.000807 0.002320743 0.000571537 0.000723 0.000591 0.000169731 0 0.000606 0 13 7 W = Winter H = House Thermoanaerobacterales Family III. -

Molecular Analyses of Changes Induced in the Microbial Populations of Murine Colon After As(III) Exposure Rishu Dheer
Duquesne University Duquesne Scholarship Collection Electronic Theses and Dissertations Summer 2011 Molecular Analyses of Changes Induced in the Microbial Populations of Murine Colon After As(III) Exposure Rishu Dheer Follow this and additional works at: https://dsc.duq.edu/etd Recommended Citation Dheer, R. (2011). Molecular Analyses of Changes Induced in the Microbial Populations of Murine Colon After As(III) Exposure (Doctoral dissertation, Duquesne University). Retrieved from https://dsc.duq.edu/etd/484 This Immediate Access is brought to you for free and open access by Duquesne Scholarship Collection. It has been accepted for inclusion in Electronic Theses and Dissertations by an authorized administrator of Duquesne Scholarship Collection. For more information, please contact [email protected]. MOLECULAR ANALYSES OF CHANGES INDUCED IN THE MICROBIAL POPULATIONS OF MURINE COLON AFTER As (III) EXPOSURE A Dissertation Submitted to Bayer School of Natural and Environmental Sciences Duquesne University In partial fulfillment of the requirements for the degree of Doctor of Philosophy By Rishu Dheer August, 2011 Copyright by Rishu Dheer 2011 MOLECULAR ANALYSES OF CHANGES INDUCED IN THE MICROBIAL POPULATIONS OF MURINE COLON AFTER As (III) EXPOSURE By Rishu Dheer Approved June 14, 2011 ________________________________ ________________________________ Dr. John F. Stolz Dr. Aaron Barchowsky Director, Center of Environmental Associate Professor of Biology Research and Education (Committee Member) Professor of Biology (Committee Chair) ________________________________ -

Oral Sub-Chronic Ochratoxin a Exposure Induces Gut Microbiota Alterations in Mice
toxins Article Oral Sub-Chronic Ochratoxin a Exposure Induces Gut Microbiota Alterations in Mice María Izco 1 , Ariane Vettorazzi 2,3 , Maria de Toro 4, Yolanda Sáenz 5 and Lydia Alvarez-Erviti 1,* 1 Laboratory of Molecular Neurobiology, Center for Biomedical Research of La Rioja (CIBIR), 26006 Logroño, Spain; [email protected] 2 MITOX Group, Department of Pharmacology and Toxicology, Faculty of Pharmacy, Universidad de Navarra, 31008 Pamplona, Spain; [email protected] 3 IdiSNA, Navarra Institute for Health Research, 31008 Pamplona, Spain 4 Center for Biomedical Research of La Rioja (CIBIR), Genomics and Bioinformatics Core Facility, 26006 Logroño, Spain; [email protected] 5 Molecular Microbiology Area, Center for Biomedical Research of La Rioja (CIBIR), 26006 Logroño, Spain; [email protected] * Correspondence: [email protected]; Tel.: +34-941-278-875 Abstract: Gut microbiota plays crucial roles in maintaining host health. External factors, such as diet, medicines, and environmental toxins, influence the composition of gut microbiota. Ochratoxin A (OTA) is one of the most prevalent and relevant mycotoxins and is a highly abundant food and animal feed contaminant. In the present study, we aimed to investigate OTA gut microbiome toxicity in mice sub-chronically exposed to low doses of OTA (0.21, 0.5, and 1.5 mg/kg body weight) by daily oral gavage for 28 days. Fecal microbiota from control and OTA-treated mice was analyzed using 16S ribosomal RNA (rRNA) gene sequencing followed by metagenomics. OTA exposure caused marked changes in gut microbial community structure, including the decrease in the diversity of fecal microbiota and the relative abundance of Firmicutes, as well as the increase in the relative abundance of Bacteroidetes at the phylum level. -

Pan-Genome Analyses of 24 Shewanella Strains Re-Emphasize
Zhong et al. Biotechnol Biofuels (2018) 11:193 https://doi.org/10.1186/s13068-018-1201-1 Biotechnology for Biofuels RESEARCH Open Access Pan‑genome analyses of 24 Shewanella strains re‑emphasize the diversifcation of their functions yet evolutionary dynamics of metal‑reducing pathway Chaofang Zhong, Maozhen Han, Shaojun Yu, Pengshuo Yang, Hongjun Li and Kang Ning* Abstract Background: Shewanella strains are important dissimilatory metal-reducing bacteria which are widely distributed in diverse habitats. Despite eforts to genomically characterize Shewanella, knowledge of the molecular components, functional information and evolutionary patterns remain lacking, especially for their compatibility in the metal- reducing pathway. The increasing number of genome sequences of Shewanella strains ofers a basis for pan-genome studies. Results: A comparative pan-genome analysis was conducted to study genomic diversity and evolutionary relation- ships among 24 Shewanella strains. Results revealed an open pan-genome of 13,406 non-redundant genes and a core-genome of 1878 non-redundant genes. Selective pressure acted on the invariant members of core genome, in which purifying selection drove evolution in the housekeeping mechanisms. Shewanella strains exhibited extensive genome variability, with high levels of gene gain and loss during the evolution, which afected variable gene sets and facilitated the rapid evolution. Additionally, genes related to metal reduction were diversely distributed in Shewanella strains and evolved under purifying selection, which highlighted the basic conserved functionality and specifcity of respiratory systems. Conclusions: The diversity of genes present in the accessory and specifc genomes of Shewanella strains indicates that each strain uses diferent strategies to adapt to diverse environments. -

Gut Microbiota Analysis UC Davis MMPC - Microbiome & Host Response Core
D4006- Gut Microbiota Analysis UC Davis MMPC - Microbiome & Host Response Core Contents 1 Methods: 1 1.1 Sequencing . .1 1.2 Data processing . .1 2 Summary of Findings: 2 2.1 Sequencing analysis . .2 2.2 Microbial diversity . .2 2.2.1 Alpha Diversity . .2 2.2.2 Beta Diversity . 10 2.3 Data analysis using taxa abundance data . 13 2.3.1 Stacked bar graphs of Taxa abundances at each level . 14 A Appendix 1 (Taxa Abundance Tables) 30 B Appendix 2 (Taxa removed from filtered datasets at each level) 37 References: 43 Core Contacts: Helen E. Raybould, Ph.D., Core Leader ([email protected]) Trina A. Knotts, Ph.D., Core Co-Leader ([email protected]) Michael L. Goodson, Ph.D., Core Scientist ([email protected]) Client(s): Kent Lloyd, DVM PhD ;MMRRC; UC Davis Project #: MBP-2079 MMRRC strain ID: MMRRC_043568 Animal Information: The strain was donated to the MMRRC by Michael Mitchell at Inserm. Fecal samples were obtained from animals housed under the care of Arthur Arnold at UCLA. 1 Methods: Brief Project Description: MMRRC strains are often contributed to the MMRRC to fulfill the resource sharing aspects of NIH grants. Since transporting mice to another facilty often causes a microbiota shift, having a record of the original fecal microbiota from the donor institution where the original phenotyping or testing was performed may prove helpful if a phenotype is lost after transfer. Several MMRRC mouse lines were selected for fecal microbiota profiling of the microbiota. Table 1: Animal-Strain Information X.SampleID TreatmentGroup Animal_ID Genotype Line Sex MMRRC.043568.M93054 MMRRC.043568_Tg.plus_M M93054 Tg.plus MMRRC.043568 M MMRRC.043568.M06285 MMRRC.043568_Tg.plus_M M06285 Tg.plus MMRRC.043568 M 1 1.1 Sequencing Frozen fecal or regional gut samples were shipped on dry ice to UC Davis MMPC and Host Microbe Systems Biology Core. -

Comprehensive Comparative Genomics Reveals Over 50 Phyla of Free‑Living and Pathogenic Bacteria Are Associated with Diverse Members of the Amoebozoa Yonas I
www.nature.com/scientificreports OPEN Comprehensive comparative genomics reveals over 50 phyla of free‑living and pathogenic bacteria are associated with diverse members of the amoebozoa Yonas I. Tekle*, Janae M. Lyttle, Maya G. Blasingame & Fang Wang The Amoebozoa, a group containing predominantly amoeboid unicellular protists has been shown to play an important ecological role in controlling environmental bacteria. Amoebozoans not only graze bacteria but also serve as a safe niche for bacterial replication and harbor endosymbiotic bacteria including dangerous human pathogens. Despite their importance, only a few lineages of Amoebozoa have been studied in this regard. In this research, we conducted a comprehensive genomic and transcriptomic study with expansive taxon sampling by including representatives from the three known clades of the Amoebozoa. We used culture independent whole culture and single cell genomics/transcriptomics to investigate the association of bacteria with diverse amoebozoans. Relative to current published evidence, we recovered the largest number of bacterial phyla (64) and human pathogen genera (51) associated with the Amoebozoa. Using single cell genomics/ transcriptomics we were able to determine up to 24 potential endosymbiotic bacterial phyla, some potentially endosymbionts. This includes the majority of multi‑drug resistant pathogens designated as major public health threats. Our study demonstrates amoebozoans are associated with many more phylogenetically diverse bacterial phyla than previously recognized. It also shows that all amoebozoans are capable of harboring far more dangerous human pathogens than presently documented, making them of primal public health concern. Te study of microbial interactions is a complex and fascinating feld of research 1–3. Microorganisms occupy diverse ecological niches and are usually found in large communities that result in inherent interactions. -

Phylogeny Reveals Novel Hipa-Homologous Kinase Families
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.14.439929; this version posted April 16, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Phylogeny Reveals Novel HipA-Homologous Kinase Families and Toxin – Antitoxin Gene Organizations Kenn Gerdes1*, Rene Bærentsen2 & Ditlev E. Brodersen2* 1) Centre of Excellence for Bacterial Stress Response and Persistence, Section for Functional Genomics, Department of Biology, University of Copenhagen, DK-2200, Copenhagen N, Denmark. 2) Department of Molecular Biology and Genetics, Aarhus University, DK-8000 Aarhus C, Denmark. * To whom correspondence should be addressed: [email protected] or [email protected] Running Head: Phylogeny of High Persister A-Homologous Kinases Keywords: High Persister A, HipB, HipS, HipT, HIRAN, Stl, GltX, TrpS bioRxiv preprint doi: https://doi.org/10.1101/2021.04.14.439929; this version posted April 16, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract Toxin – Antitoxin modules function in the genetic stability of mobile genetic elements, bacteriophage defense, and antibiotic tolerance. A gain-of-function mutation of the Escherichia coli K-12 hipBA module can induce antibiotic tolerance in a subpopulation of bacterial cells, a phenomenon known as persistence. HipA is a Ser/Thr kinase that phosphorylates and inactivates glutamyl tRNA synthetase, inhibiting cellular translation and inducing the stringent response. -

Comparative Genomics and Experimental Characterization of N
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 281, NO. 40, pp. 29872–29885, October 6, 2006 Printed in the U.S.A. Comparative Genomics and Experimental Characterization of N-Acetylglucosamine Utilization Pathway of Shewanella oneidensis*□S Received for publication, May 26, 2006, and in revised form, July 20, 2006 Published, JBC Papers in Press, July 20, 2006, DOI 10.1074/jbc.M605052200 Chen Yang‡1, Dmitry A. Rodionov‡§1,2, Xiaoqing Li‡, Olga N. Laikova¶, Mikhail S. Gelfand§ʈ, Olga P. Zagnitko**, Margaret F. Romine‡‡3, Anna Y. Obraztsova§§, Kenneth H. Nealson§§3, and Andrei L. Osterman‡** From the ‡Burnham Institute for Medical Research, La Jolla, California 92037, §Institute for Information Transmission Problems, Russian Academy of Sciences, Moscow 127994, Russia, ¶State Scientific Center GosNIIGenetika, Moscow 117545, Russia, ʈDepartment of Bioengineering and Bioinformatics, Moscow State University, Moscow 119992, Russia, **Fellowship for ‡‡ Interpretation of Genomes, Burr Ridge, Illinois 60527, Pacific Northwest National Laboratory, Richland, Washington 99352, Downloaded from and §§Department of Earth Sciences, University of Southern California, Los Angeles, California 90089 We used a comparative genomics approach implemented in the aquatic environments and abundant in environments where SEED annotation environment to reconstruct the chitin and GlcNAc nutrient levels are high and redox interfaces are found. utilization subsystem and regulatory network in most proteobacteria, Shewanella oneidensis MR-1 was originally isolated from fresh- www.jbc.org including 11 species of Shewanella with completely sequenced water sediments of Oneida Lake, NY (1), whereas other species genomes. Comparative analysis of candidate regulatory sites allowed have been isolated from marine sediments, marine waters, and us to characterize three different GlcNAc-specific regulons, NagC, a variety of other environments (2). -

The Shewanella Genus: Ubiquitous Organisms Sustaining and Preserving Aquatic Ecosystems Olivier Lemaire, Vincent Méjean, Chantal Iobbi-Nivol
The Shewanella genus: ubiquitous organisms sustaining and preserving aquatic ecosystems Olivier Lemaire, Vincent Méjean, Chantal Iobbi-Nivol To cite this version: Olivier Lemaire, Vincent Méjean, Chantal Iobbi-Nivol. The Shewanella genus: ubiquitous organisms sustaining and preserving aquatic ecosystems. FEMS Microbiology Reviews, Wiley-Blackwell, 2020, 44 (2), pp.155-170. 10.1093/femsre/fuz031. hal-02936277 HAL Id: hal-02936277 https://hal.archives-ouvertes.fr/hal-02936277 Submitted on 11 Mar 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. The Shewanella genus: ubiquitous organisms sustaining and preserving aquatic ecosystems. Olivier N. Lemaire*, Vincent Méjean and Chantal Iobbi-Nivol Aix-Marseille Université, Laboratoire de Bioénergétique et Ingénierie des Protéines, UMR 7281, Institut de Microbiologie de la Méditerranée, Centre National de la Recherche Scientifique, 13402 Marseille, France. *Corresponding author. Email: [email protected] Keywords Bacteria, Microbial Physiology, Ecological Network, Microflora, Symbiosis, Biotechnology