Polygenic Scores for Major Depressive Disorder and Depressive Symptoms Predict Response to Lithium in Patients with Bipolar Disorder
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

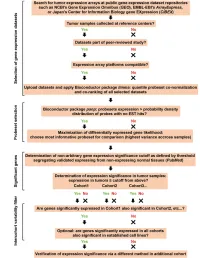
A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Bioinformatic Analysis of Autism Positional Candidate Genes Using Biological Databases and Computational Gene Network Prediction
Genes, Brain and Behavior (2003) 2: 303–320 Copyright # Blackwell Munksgaard 2003 Bioinformatic analysis of autism positional candidate genes using biological databases and computational gene network prediction A. L. Yonan†,‡, A. A. Palmer†, K. C. Smith†, inform studies of autism, and to illustrate and explore I. Feldman†,††, H. K. Lee†, J. M. Yonan§, the increasing potential of bioinformatic approaches as † †,†† *,†,‡, S. G. Fischer , P. Pavlidis and T. C. Gilliam { a compliment to linkage analysis. Keywords: 17q, AGRE sample, autism, association studies, †Columbia Genome Center, Columbia University, New York, bioinformatics, candidate genes ‡ Department of Genetics and Development, Columbia University, Received 30 June 2003, revised 20 August 2003, accepted New York, for publication 21 August 2003 ¶Department of Psychiatry, Columbia University and New York State Psychiatric Institute, New York, §Division of Molecular Genetics, Departments of Pediatrics and Medicine, Columbia University, New York, Autism is a pervasive neurodevelopmental disorder that ††Department of Biomedical Informatics, Columbia University, severely impairs development of normal social and emotional New York, USA interactions and related forms of communication. Disease *Corresponding author: T. C. Gilliam, Columbia Genome Center, symptoms characteristically include unusually restricted and 1150 St. Nicholas Avenue, Room 508, New York, NY 10032, USA. E-mail: [email protected] stereotyped patterns of behaviors and interests. Autism describes the most severe manifestation -

Characterizing Genomic Duplication in Autism Spectrum Disorder by Edward James Higginbotham a Thesis Submitted in Conformity
Characterizing Genomic Duplication in Autism Spectrum Disorder by Edward James Higginbotham A thesis submitted in conformity with the requirements for the degree of Master of Science Graduate Department of Molecular Genetics University of Toronto © Copyright by Edward James Higginbotham 2020 i Abstract Characterizing Genomic Duplication in Autism Spectrum Disorder Edward James Higginbotham Master of Science Graduate Department of Molecular Genetics University of Toronto 2020 Duplication, the gain of additional copies of genomic material relative to its ancestral diploid state is yet to achieve full appreciation for its role in human traits and disease. Challenges include accurately genotyping, annotating, and characterizing the properties of duplications, and resolving duplication mechanisms. Whole genome sequencing, in principle, should enable accurate detection of duplications in a single experiment. This thesis makes use of the technology to catalogue disease relevant duplications in the genomes of 2,739 individuals with Autism Spectrum Disorder (ASD) who enrolled in the Autism Speaks MSSNG Project. Fine-mapping the breakpoint junctions of 259 ASD-relevant duplications identified 34 (13.1%) variants with complex genomic structures as well as tandem (193/259, 74.5%) and NAHR- mediated (6/259, 2.3%) duplications. As whole genome sequencing-based studies expand in scale and reach, a continued focus on generating high-quality, standardized duplication data will be prerequisite to addressing their associated biological mechanisms. ii Acknowledgements I thank Dr. Stephen Scherer for his leadership par excellence, his generosity, and for giving me a chance. I am grateful for his investment and the opportunities afforded me, from which I have learned and benefited. I would next thank Drs. -

A Chromosome-Centric Human Proteome Project (C-HPP) To
computational proteomics Laboratory for Computational Proteomics www.FenyoLab.org E-mail: [email protected] Facebook: NYUMC Computational Proteomics Laboratory Twitter: @CompProteomics Perspective pubs.acs.org/jpr A Chromosome-centric Human Proteome Project (C-HPP) to Characterize the Sets of Proteins Encoded in Chromosome 17 † ‡ § ∥ ‡ ⊥ Suli Liu, Hogune Im, Amos Bairoch, Massimo Cristofanilli, Rui Chen, Eric W. Deutsch, # ¶ △ ● § † Stephen Dalton, David Fenyo, Susan Fanayan,$ Chris Gates, , Pascale Gaudet, Marina Hincapie, ○ ■ △ ⬡ ‡ ⊥ ⬢ Samir Hanash, Hoguen Kim, Seul-Ki Jeong, Emma Lundberg, George Mias, Rajasree Menon, , ∥ □ △ # ⬡ ▲ † Zhaomei Mu, Edouard Nice, Young-Ki Paik, , Mathias Uhlen, Lance Wells, Shiaw-Lin Wu, † † † ‡ ⊥ ⬢ ⬡ Fangfei Yan, Fan Zhang, Yue Zhang, Michael Snyder, Gilbert S. Omenn, , Ronald C. Beavis, † # and William S. Hancock*, ,$, † Barnett Institute and Department of Chemistry and Chemical Biology, Northeastern University, Boston, Massachusetts 02115, United States ‡ Stanford University, Palo Alto, California, United States § Swiss Institute of Bioinformatics (SIB) and University of Geneva, Geneva, Switzerland ∥ Fox Chase Cancer Center, Philadelphia, Pennsylvania, United States ⊥ Institute for System Biology, Seattle, Washington, United States ¶ School of Medicine, New York University, New York, United States $Department of Chemistry and Biomolecular Sciences, Macquarie University, Sydney, NSW, Australia ○ MD Anderson Cancer Center, Houston, Texas, United States ■ Yonsei University College of Medicine, Yonsei University, -

(12) United States Patent (10) Patent No.: US 8,148,129 B2 Frankel Et Al
US008148129B2 (12) United States Patent (10) Patent No.: US 8,148,129 B2 Frankel et al. (45) Date of Patent: Apr. 3, 2012 (54) GENERATION OF POTENT DOMINANT 6,824,978 B1 1 1/2004 Cox, III et al. NEGATIVE TRANSCRIPTIONAL 6,933,113 B2 8, 2005 Case et al. 6,979,539 B2 12/2005 Cox, III et al. INHIBITORS 7,013,219 B2 3/2006 Case et al. 7,070,934 B2 7/2006 Cox, III et al. (75) Inventors: Alan Frankel, Mill Valley, CA (US); 7,163,824 B2 1/2007 Cox, III et al. Robert Nakamura, San Francisco, CA 7,220,719 B2 5/2007 Case et al. (US); Chandreyee Das, Brookline, MA 7,235,354 B2 6/2007 Case et al. 7,262,054 B2 8/2007 Jamieson et al. (US); Ivan D’Orso, San Francisco, CA 7,273,923 B2 9/2007 Jamieson et al. (US); Jocelyn Grunwell, San Mateo, 2003, OO82552 A1* 5, 2003 Wolffe et al. ..................... 435/6 CA (US) (73) Assignee: The Regents of the University of OTHER PUBLICATIONS California, Oakland, CA (US) Cramer et al., Coupling of Transcription with Alternative Splicing: RNA Pol II Promoters Modulate SF2. ASF and 9G8 Effects on an (*) Notice: Subject to any disclaimer, the term of this Exonic Splicing Enhancer, Molecular Cell, 1999, 4:251-258.* patent is extended or adjusted under 35 Cama-Carvalho et al., Nucleocytoplasmic shuttling of heterodimeric U.S.C. 154(b) by 806 days. splicing factor U2AF, JBC. Published on Dec. 15, 2000 as Manu script M008759200.* (21) Appl. No.: 11/765,592 Rosonina et al., Gene Expression: The Close Coupling of Transcrip tion and Splicing, Current Biology, vol. -

Autocrine IFN Signaling Inducing Profibrotic Fibroblast Responses By
Downloaded from http://www.jimmunol.org/ by guest on September 23, 2021 Inducing is online at: average * The Journal of Immunology , 11 of which you can access for free at: 2013; 191:2956-2966; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication August 2013; doi: 10.4049/jimmunol.1300376 http://www.jimmunol.org/content/191/6/2956 A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Autocrine IFN Signaling Feng Fang, Kohtaro Ooka, Xiaoyong Sun, Ruchi Shah, Swati Bhattacharyya, Jun Wei and John Varga J Immunol cites 49 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130037 6.DC1 This article http://www.jimmunol.org/content/191/6/2956.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 23, 2021. The Journal of Immunology A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Inducing Autocrine IFN Signaling Feng Fang,* Kohtaro Ooka,* Xiaoyong Sun,† Ruchi Shah,* Swati Bhattacharyya,* Jun Wei,* and John Varga* Activation of TLR3 by exogenous microbial ligands or endogenous injury-associated ligands leads to production of type I IFN. -
Supplementary Data
SUPPLEMENTAL INFORMATION A study restricted to chemokine receptors as well as a genome-wide transcript analysis uncovered CXCR4 as preferentially expressed in Ewing's sarcoma (Ewing's sarcoma) cells of metastatic origin (Figure 4). Transcriptome analyses showed that in addition to CXCR4, genes known to support cell motility and invasion topped the list of genes preferentially expressed in metastasis-derived cells (Figure 4D). These included kynurenine 3-monooxygenase (KMO), galectin-1 (LGALS1), gastrin-releasing peptide (GRP), procollagen C-endopeptidase enhancer (PCOLCE), and ephrin receptor B (EPHB3). KMO, a key enzyme of tryptophan catabolism, has not been linked to metastasis. Tryptophan and its catabolites, however, are involved in immune evasion by tumors, a process that can assist in tumor progression and metastasis (1). LGALS1, GRP, PCOLCE and EPHB3 have been linked to tumor progression and metastasis of several cancers (2-4). Top genes preferentially expressed in L-EDCL included genes that suppress cell motility and/or potentiate cell adhesion such as plakophilin 1 (PKP1), neuropeptide Y (NPY), or the metastasis suppressor TXNIP (5-7) (Figure 4D). Overall, L-EDCL were enriched in gene sets geared at optimizing nutrient transport and usage (Figure 4D; Supplementary Table 3), a state that may support the early stages of tumor growth. Once tumor growth outpaces nutrient and oxygen supplies, gene expression programs are usually switched to hypoxic response and neoangiogenesis, which ultimately lead to tumor egress and metastasis. Accordingly, gene sets involved in extracellular matrix remodeling, MAPK signaling, and response to hypoxia were up-regulated in M-EDCL (Figure 4D; Supplementary Table 4), consistent with their association to metastasis in other cancers (8, 9). -

WO 2015/149034 A2 1 October 2015 (01.10.2015) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2015/149034 A2 1 October 2015 (01.10.2015) P O P C T (51) International Patent Classification: (71) Applicant: LIFE TECHNOLOGIES CORPORATION C12Q 1/68 (2006.01) [US/US]; c/o IP Legal Department Docketing, 5791 Van Allen Way, Carlsbad, California 92008 (US). (21) International Application Number: PCT/US2015/023 197 (72) Inventors: RHODES, Daniel; Life Technologies Corpora tion, c/o IP Legal Department Docketing, 579 1 Van Allen (22) International Filing Date: Way, Carlsbad, California 92008 (US). SADIS, Seth; Life 27 March 2015 (27.03.2015) Technologies Corporation, c/o IP Legal Department Dock (25) Filing Language: English eting, 5791 Van Allen Way, Carlsbad, California 92008 (US). WYNGAARD, Peter; Life Technologies Corpora (26) Publication Language: English tion, c/o IP Legal Department Docketing, 579 1 Van Allen (30) Priority Data: Way, Carlsbad, California 92008 (US). KHAZANOV, 61/971,455 27 March 2014 (27.03.2014) US Nikolay; Life Technologies Corporation, c/o IP Legal De 61/993,732 15 May 2014 (15.05.2014) US partment Docketing, 5791 Van Allen Way, Carlsbad, Cali 62/004,727 29 May 2014 (29.05.2014) us fornia 92008 (US). BANDLA, Santhoshi; Life Technolo 62/092,898 17 December 2014 (17. 12.2014) us gies Corporation, c/o IP Legal Department Docketing, 5791 Van Allen Way, Carlsbad, California 92008 (US). [Continued on nextpage] (54) Title: GENE FUSIONS AND GENE VARIANTS ASSOCIATED WITH CANCER (57) Abstract: The disclosure provides gene fusions, gene variants, and novel as RNASeq sociations with disease states, as well as kits, probes, and methods of using the (.bam) same. -

Complex Pathogenesis of Hirschsprung's Disease in a Patient
European Journal of Human Genetics (2009) 17, 483 – 490 & 2009 Macmillan Publishers Limited All rights reserved 1018-4813/09 $32.00 www.nature.com/ejhg ARTICLE Complex pathogenesis of Hirschsprung’s disease in a patient with hydrocephalus, vesico-ureteral reflux and a balanced translocation t(3;17)(p12;q11) Paola Griseri1, Yvonne Vos2, Roberto Giorda3, Stefania Gimelli4, Silvana Beri3, Giuseppe Santamaria1, Guendalina Mognato5, Robert MW Hofstra2, Giorgio Gimelli1 and Isabella Ceccherini*1 1Laboratory Molecular Genetics and Cytogenetics, Ist. G. Gaslini, Genova, Italy; 2Department of Genetics, University Medical Center Groningen, University of Groningen, Groningen, The Netherlands; 3‘E. Medea’ Scientific Institute, Bosisio Parini, Lecco, Italy; 4Dipartimento di Patologia Umana ed Ereditaria, Biologia Generale e Genetica Medica, Universita` di Pavia, Pavia, Italy; 5Clinica Chirurgica Pediatrica, Dipartimento di Pediatria, Universita` di Padova, Padova, Italy Hirschsprung’s disease (HSCR), a congenital complex disorder of intestinal innervation, is often associated with other inherited syndromes. Identifying genes involved in syndromic HSCR cases will not only help understanding the specific underlying diseases, but it will also give an insight into the development of the most frequent isolated HSCR. The association between hydrocephalus and HSCR is not surprising as a large number of patients have been reported to show the same clinical association, most of them showing mutations in the L1CAM gene, encoding a neural adhesion molecule often involved in isolated X-linked hydrocephalus. L1 defects are believed to be necessary but not sufficient for the occurrence of the intestinal phenotype in syndromic cases. In this paper, we have carried out the molecular characterization of a patient affected with Hirschsprung’s disease and X-linked hydrocephalus, with a de novo reciprocal balanced translocation t(3;17)(p12;q21). -

Data Set 1. Biological Analysis of the Genes Found to Be Significant in the Endotoxin Study
Data Set 1. Biological analysis of the genes found to be significant in the endotoxin study. Pages 2 – 105: Q values, gene names and annotation on the genes significant at FDR = 0.1%. Pages 106 – 127: Global functional analysis of the down-regulated genes. Pages 128 – 169: Global functional analysis of the up-regulated genes. Probe Set q-value Direction Gene Annotation 117_at 9.60E-05 up HSPA6 heat shock 70kDa protein 6 (HSP70B') 1405_i_at 2.00E-06 down CCL5 chemokine (C-C motif) ligand 5 PRP8 pre-mRNA processing factor 8 200000_s_at 2.50E-05 down PRPF8 homolog (yeast) PRP8 pre-mRNA processing factor 8 200000_s_at 0.000712 down PRPF8 homolog (yeast) 200001_at 0.000658 down CAPNS1 calpain, small subunit 1 200002_at 2.00E-06 down RPL35 ribosomal protein L35 200002_at 2.00E-06 down RPL35 ribosomal protein L35 200003_s_at 1.10E-05 down RPL28 ribosomal protein L28 200003_s_at 1.10E-05 down RPL28 ribosomal protein L28 eukaryotic translation initiation factor 3, 200005_at 2.00E-06 down EIF3S7 subunit 7 zeta, 66/67kDa eukaryotic translation initiation factor 3, 200005_at 2.00E-06 down EIF3S7 subunit 7 zeta, 66/67kDa Parkinson disease (autosomal 200006_at 4.70E-05 down PARK7 recessive, early onset) 7 Parkinson disease (autosomal 200006_at 8.70E-05 down PARK7 recessive, early onset) 7 200008_s_at 5.40E-05 up GDI2 GDP dissociation inhibitor 2 200008_s_at 0.000361 up GDI2 GDP dissociation inhibitor 2 200009_at 0.000171 down GDI2 GDP dissociation inhibitor 2 200010_at 2.00E-06 down RPL11 ribosomal protein L11 200010_at 2.00E-06 down RPL11 ribosomal -

Advances in Analysis of Novel Protein Therapeutics and Clinical Samples with LC-MS/MS Methods
ADVANCES IN ANALYSIS OF NOVEL PROTEIN THERAPEUTICS AND CLINICAL SAMPLES WITH LC-MS/MS METHODS A Dissertation Presented by Fangfei Yan to The Department of Chemistry and Chemical Biology In partial fulfillment of the requirements for the degree of Doctor of Philosophy in the field of Chemistry Northeastern University Boston, Massachusetts April 9, 2013 ADVANCES IN ANALYSIS OF NOVEL PROTEIN THERAPEUTICS AND CLINICAL SAMPLES WITH LC-MS/MS METHODS by Fangfei Yan ABSTRACT OF DISSERTATION Submitted in partial fulfillment of the requirements for the Degree of Doctor of Philosophy in Chemistry in the College of Science of Northeastern University April 9, 2013 2 Abstract Analytical characterization is essential for drug efficacy, stability and safety. Significant concern in protein stability and potential immunogenicity requires detailed characterization of protein therapeutics. Therefore, the ability to detect product variants from non-clonal cell lines is very important. To fulfill this job, LC-MS/MS becomes the perfect choice. This Ph.D. research discusses advances in analysis of novel protein therapeutics and clinical samples with LC- MS/MS methods. Recombinant protein therapeutics is a major focus of my research work. In Chapter 1, an overview of production of recombinant protein therapeutics and the potential heterogeneity, in particular post-translational modifications, is reviewed. Common techniques for protein characterization are discussed, including electrophoresis, chromatography, and mass spectrometry. Because the major focus of my research is about mass spectrometry, further details about ionization method, mass analyzer, hybrid mass spectrometer, fragmentation method and LC-MS/MS analysis are reviewed. In addition, common strategies for protein characterization are discussed. Furthermore, biology part for proteomic analysis is reviewed. -

Regulated T Cell Pre-Mrna Splicing As Genetic Marker of T Cell Suppression
Regulated T cell pre-mRNA splicing as genetic marker of T cell suppression by Boitumelo Mofolo Thesis presented for the degree of Master of Science (Bioinformatics) Institute of Infectious Disease and MolecularTown Medicine University of Cape Town (UCT) Supervisor:Cape Assoc. Prof. ofNicola Mulder August 2012 University The copyright of this thesis vests in the author. No quotation from it or information derived from it is to be published without full acknowledgementTown of the source. The thesis is to be used for private study or non- commercial research purposes only. Cape Published by the University ofof Cape Town (UCT) in terms of the non-exclusive license granted to UCT by the author. University Declaration I, Boitumelo Mofolo, declare that all the work in this thesis, excluding that has been cited and referenced, is my own. Signature Boitumelo Mofolo Town Cape of University Copyright©2012 University of Cape Town All rights reserved 1 ABSTRACT T CELL NORMAL T CELL SUPPRESSION p110 p110 MV p85 PI3K AKT p85 PI3K PHOSPHORYLATION NO PHOSHORYLATION HIV cytoplasm HCMV LCK-011 PRMT5-006 SHIP145 SIP110 Town RV LCK-010 VCL-204 ATM-016 PRMT5-018 ATM-002 CALD1-008 LCK-006 MXI1-001 VCL-202 NRP1-201 MXI1-007 CALD1-004Cape nucleus of Background: Measles is a highly contagious disease that mainly affects children and according to the World Health Organisation (WHO), was responsible for over 164000 deaths in 2008, despite the availability of a safe and cost-effective vaccine [56]. The Measles virus (MV) inactivates T- cells, rendering them dysfunctional, and results in virally induced immunosuppression which shares certain features with thatUniversity induced by HIV.