Age-Dependent Decline in Fate Switch from NG2 Cells to Astrocytes After Olig2 Deletion
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The GATA2 Transcription Factor Negatively Regulates the Proliferation of Neuronal Progenitors
RESEARCH ARTICLE 2155 Development 133, 2155-2165 (2006) doi:10.1242/dev.02377 The GATA2 transcription factor negatively regulates the proliferation of neuronal progenitors Abeer El Wakil*, Cédric Francius*,†, Annie Wolff, Jocelyne Pleau-Varet† and Jeannette Nardelli†,§ Postmitotic neurons are produced from a pool of cycling progenitors in an orderly fashion that requires proper spatial and temporal coordination of proliferation, fate determination, differentiation and morphogenesis. This probably relies on complex interplay between mechanisms that control cell cycle, specification and differentiation. In this respect, we have studied the possible implication of GATA2, a transcription factor that is involved in several neuronal specification pathways, in the control of the proliferation of neural progenitors in the embryonic spinal cord. Using gain- and loss-of-function manipulations, we have shown that Gata2 can drive neural progenitors out of the cycle and, to some extent, into differentiation. This correlates with the control of cyclin D1 transcription and of the expression of the p27/Kip1 protein. Interestingly, this functional aspect is not only associated with silencing of the Notch pathway but also appears to be independent of proneural function. Consistently, GATA2 also controls the proliferation capacity of mouse embryonic neuroepithelial cells in culture. Indeed, Gata2 inactivation enhances the proliferation rate in these cells. By contrast, GATA2 overexpression is sufficient to force such cells and neuroblastoma cells to stop dividing but not to drive either type of cell into differentiation. Furthermore, a non-cell autonomous effect of Gata2 expression was observed in vivo as well as in vitro. Hence, our data have provided evidence for the ability of Gata2 to inhibit the proliferation of neural progenitors, and they further suggest that, in this regard, Gata2 can operate independently of neuronal differentiation. -

Olig1 and Sox10 Interact Synergistically to Drivemyelin Basic
The Journal of Neuroscience, December 26, 2007 • 27(52):14375–14382 • 14375 Cellular/Molecular Olig1 and Sox10 Interact Synergistically to Drive Myelin Basic Protein Transcription in Oligodendrocytes Huiliang Li,1 Yan Lu,2 Hazel K. Smith,1 and William D. Richardson1 1Wolfson Institute for Biomedical Research and Department of Biology, University College London, London WC1E 6BT, United Kingdom, and 2Medical Research Council, Clinical Sciences Centre, Imperial College London, London W12 0NN, United Kingdom The oligodendrocyte lineage genes (Olig1/2), encoding basic helix-loop-helix transcription factors, were first identified in screens for master regulators of oligodendrocyte development. OLIG1 is important for differentiation of oligodendrocyte precursors into myelin- forming oligodendrocytes during development and is thought to play a crucial role in remyelination during multiple sclerosis. However, itisstillunclearhowOLIG1interactswithitstranscriptionalcofactorsandDNAtargets.OLIG1wasreportedlyrestrictedtomammals,but we demonstrate here that zebrafish and other teleosts also possess an OLIG1 homolog. In zebrafish, as in mammals, Olig1 is expressed in the oligodendrocyte lineage. Olig1 associates physically with another myelin-associated transcription factor, Sox10, and the Olig1/Sox10 complex activates mbp (myelin basic protein) transcription via conserved DNA sequence motifs in the mbp promoter region. In contrast, Olig2 does not bind to Sox10 in zebrafish, although both OLIG1 and OLIG2 bind SOX10 in mouse. Key words: Olig1; Olig2; Sox10; Mbp; oligodendrocyte; myelin; zebrafish; mouse; evolution; development Introduction directly regulates Mbp transcription (Stolt et al., 2002), and over- Myelin, the multilayered glial sheath around axons, is one of the expression of SOX10 alone is sufficient to induce myelin gene defining features of jawed vertebrates (gnathostomes). It is expression in embryonic chick spinal cord (Liu et al., 2007). -

SUPPLEMENTARY MATERIAL Bone Morphogenetic Protein 4 Promotes
www.intjdevbiol.com doi: 10.1387/ijdb.160040mk SUPPLEMENTARY MATERIAL corresponding to: Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells SUMIYO MIMURA, MIKA SUGA, KAORI OKADA, MASAKI KINEHARA, HIROKI NIKAWA and MIHO K. FURUE* *Address correspondence to: Miho Kusuda Furue. Laboratory of Stem Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, 7-6-8, Saito-Asagi, Ibaraki, Osaka 567-0085, Japan. Tel: 81-72-641-9819. Fax: 81-72-641-9812. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.160040mk TABLE S1 PRIMER LIST FOR QRT-PCR Gene forward reverse AP2α AATTTCTCAACCGACAACATT ATCTGTTTTGTAGCCAGGAGC CDX2 CTGGAGCTGGAGAAGGAGTTTC ATTTTAACCTGCCTCTCAGAGAGC DLX1 AGTTTGCAGTTGCAGGCTTT CCCTGCTTCATCAGCTTCTT FOXD3 CAGCGGTTCGGCGGGAGG TGAGTGAGAGGTTGTGGCGGATG GAPDH CAAAGTTGTCATGGATGACC CCATGGAGAAGGCTGGGG MSX1 GGATCAGACTTCGGAGAGTGAACT GCCTTCCCTTTAACCCTCACA NANOG TGAACCTCAGCTACAAACAG TGGTGGTAGGAAGAGTAAAG OCT4 GACAGGGGGAGGGGAGGAGCTAGG CTTCCCTCCAACCAGTTGCCCCAAA PAX3 TTGCAATGGCCTCTCAC AGGGGAGAGCGCGTAATC PAX6 GTCCATCTTTGCTTGGGAAA TAGCCAGGTTGCGAAGAACT p75 TCATCCCTGTCTATTGCTCCA TGTTCTGCTTGCAGCTGTTC SOX9 AATGGAGCAGCGAAATCAAC CAGAGAGATTTAGCACACTGATC SOX10 GACCAGTACCCGCACCTG CGCTTGTCACTTTCGTTCAG Suppl. Fig. S1. Comparison of the gene expression profiles of the ES cells and the cells induced by NC and NC-B condition. Scatter plots compares the normalized expression of every gene on the array (refer to Table S3). The central line -

Hormonal Regulation of Oligodendrogenesis I: Effects Across the Lifespan
biomolecules Review Hormonal Regulation of Oligodendrogenesis I: Effects across the Lifespan Kimberly L. P. Long 1,*,†,‡ , Jocelyn M. Breton 1,‡,§ , Matthew K. Barraza 2 , Olga S. Perloff 3 and Daniela Kaufer 1,4,5 1 Helen Wills Neuroscience Institute, University of California, Berkeley, CA 94720, USA; [email protected] (J.M.B.); [email protected] (D.K.) 2 Department of Molecular and Cellular Biology, University of California, Berkeley, CA 94720, USA; [email protected] 3 Memory and Aging Center, Department of Neurology, University of California, San Francisco, CA 94143, USA; [email protected] 4 Department of Integrative Biology, University of California, Berkeley, CA 94720, USA 5 Canadian Institute for Advanced Research, Toronto, ON M5G 1M1, Canada * Correspondence: [email protected] † Current address: Department of Psychiatry and Behavioral Sciences, University of California, San Francisco, CA 94143, USA. ‡ These authors contributed equally to this work. § Current address: Department of Psychiatry, Columbia University, New York, NY 10027, USA. Abstract: The brain’s capacity to respond to changing environments via hormonal signaling is critical to fine-tuned function. An emerging body of literature highlights a role for myelin plasticity as a prominent type of experience-dependent plasticity in the adult brain. Myelin plasticity is driven by oligodendrocytes (OLs) and their precursor cells (OPCs). OPC differentiation regulates the trajectory of myelin production throughout development, and importantly, OPCs maintain the ability to proliferate and generate new OLs throughout adulthood. The process of oligodendrogenesis, Citation: Long, K.L.P.; Breton, J.M.; the‘creation of new OLs, can be dramatically influenced during early development and in adulthood Barraza, M.K.; Perloff, O.S.; Kaufer, D. -
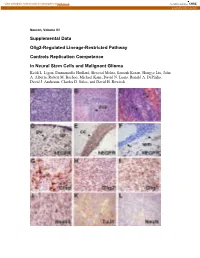
Supplemental Data Olig2-Regulated Lineage-Restricted Pathway Controls Replication Competence in Neural Stem Cells and Malignant
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Caltech Authors Neuron, Volume 53 Supplemental Data Olig2-Regulated Lineage-Restricted Pathway Controls Replication Competence in Neural Stem Cells and Malignant Glioma Keith L. Ligon, Emmanuelle Huillard, Shwetal Mehta, Santosh Kesari, Hongye Liu, John A. Alberta, Robert M. Bachoo, Michael Kane, David N. Louis, Ronald A. DePinho, David J. Anderson, Charles D. Stiles, and David H. Rowitch Figure S1. Olig1/2+/- Ink4a/Arf-/-EGFRvIII Gliomas Express Characteristic Morphologic and Immunophenotypic Features of Human Malignant Gliomas (A) Tumors exhibit dense cellularity and atypia. (B) Characteristic infiltration of host SVZ stem cell niche. (C) Although occasional tumors exhibited pseudopalisading necrosis (n, arrowhead) and hemorrhage (h, arrowhead) similar to human GBM (Astrocytoma WHO Grade IV), most tumors lacked these features. (D) IHC for hEGFR highlights hallmark feature of human gliomas, including perivascular (pv) and subpial (sp) accumulation of tumor cells, (E) striking white matter tropism of tumor cells in corpus callosum (cc), and (F) distant single cell infiltration of cerebellar white matter (wm, arrows). (G-J) Immunohistochemical markers characteristic of human tumors (Gfap, Olig2, Olig1, Nestin) are present. (K and L) Weak staining for the early neuronal marker TuJ1 and absence of the differentiated neuronal marker, NeuN similar to human glioma and consistent with heterogeneous lines of partial differentiation. Figure S2. Olig1/2+/- Ink4a/Arf-/-EGFRvIII Neurospheres Are Multipotent and Olig1/2-/- Ink4a/Arf-/-EGFRvIII Neurospheres Are Bipotent Olig1/2+/- or Olig1/2-/- Ink4a/Arf-/-EGFRvIII neurospheres were allowed to differentiate for 6 days in medium without EGF. -

Characterization of Transcription Factor Networks Involved in Umbilical Cord Blood CD34+ Stem Cells-Derived Erythropoiesis
Characterization of Transcription Factor Networks Involved in Umbilical Cord Blood CD34+ Stem Cells-Derived Erythropoiesis Biaoru Li1, Lianghao Ding2, Chinrang Yang2, Baolin Kang1, Li Liu3, Michael D. Story2, Betty S. Pace1* 1 Department of Pediatrics, Hematology/Oncology Division, Georgia Regents University, Augusta, Georgia, United States of America, 2 Department of Radiation Oncology and Simmons Comprehensive Cancer Center, University of Texas Southwestern Medical Center, Dallas, Texas, United States of America, 3 Department of Molecular and Cell Biology, University of Texas at Dallas, Richardson, Texas, United States of America Abstract Fetal stem cells isolated from umbilical cord blood (UCB) possess a great capacity for proliferation and differentiation and serve as a valuable model system to study gene regulation. Expanded knowledge of the molecular control of hemoglobin synthesis will provide a basis for rational design of therapies for b-hemoglobinopathies. Transcriptome data are available for erythroid progenitors derived from adult stem cells, however studies to define molecular mechanisms controlling globin gene regulation during fetal erythropoiesis are limited. Here, we utilize UCB-CD34+ stem cells induced to undergo erythroid differentiation to characterize the transcriptome and transcription factor networks (TFNs) associated with the c/b-globin switch during fetal erythropoiesis. UCB-CD34+ stem cells grown in the one-phase liquid culture system displayed a higher proliferative capacity than adult CD34+ stem cells. The c/b-globin switch was observed after day 42 during fetal erythropoiesis in contrast to adult progenitors where the switch occurred around day 21. To gain insights into transcription factors involved in globin gene regulation, microarray analysis was performed on RNA isolated from UCB-CD34+ cell-derived erythroid progenitors harvested on day 21, 42, 49 and 56 using the HumanHT-12 Expression BeadChip. -

Transcriptional and Epigenetic Characterization of Early Striosomes
bioRxiv preprint doi: https://doi.org/10.1101/2020.05.19.105171; this version posted May 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Transcriptional and epigenetic characterization of early striosomes identifies Foxf2 and Olig2 as factors required for development of striatal compartmentation and neuronal phenotypic differentiation Maria-Daniela Cirnaru1, Sicheng Song2, Kizito-Tshitoko Tshilenge3, Chuhyon Corwin1, Justyna Mleczko1, Carlos Galicia Aguirre3, Houda Benlhabib2, Jaroslav Bendl4,5,6, Jordi Creus Muncunill1, Pasha Apontes4,5,6, John F. Fullard4,5,6, Peter Carlsson7, Panos Roussos4,5,6,8, Sean D. Mooney2, Lisa M. Ellerby3, Michelle E. Ehrlich1* 1. Department of Neurology, Icahn School of Medicine at Mount Sinai, New York, New York 10029, USA. 2. Department of Epidemiology, University of Washington, Seattle, WA, USA. 3. Buck Institute for Research on Aging, Novato, CA, 94945, USA. 4. Pamela Sklar Division of Psychiatric Genomics, Icahn School of Medicine at Mount Sinai, One Gustave L. Levy Place, New York, NY, 10029, USA. 5. Institute for Genomics and Multiscale Biology, Department of Genetics and Genomic Sciences, Icahn School of Medicine at Mount Sinai, One Gustave L. Levy Place, New York, NY, 10029, USA. 6. Department of Psychiatry, Icahn School of Medicine at Mount Sinai, One Gustave L. Levy Place, New York, NY, 10029, USA. 7. Department of Chemistry and Molecular Biology, University of Gothenburg, Gothenburg, Sweden. 8. Mental Illness Research, Education, and Clinical Center (VISN 2 South), James J. Peters VA Medical Center, Bronx, NY 10468, USA. -

Combinatorial Profiles of Oligodendrocyte-Selective Classes of Transcriptional Regulators Differentially Modulate Myelin Basic Protein Gene Expression
The Journal of Neuroscience, September 7, 2005 • 25(36):8311–8321 • 8311 Development/Plasticity/Repair Combinatorial Profiles of Oligodendrocyte-Selective Classes of Transcriptional Regulators Differentially Modulate Myelin Basic Protein Gene Expression Solen Gokhan,1,2,3,7* Mireya Marin-Husstege,8* Shau Yu Yung,4 Darah Fontanez,8,9 Patrizia Casaccia-Bonnefil,8,9 and Mark F. Mehler1,2,3,4,5,6,7 1Institute for Brain Disorders and Neural Regeneration, 2F. M. Kirby Program in Neural Protection and Repair, Departments of 3Neurology, 4Neuroscience, and 5Psychiatry and Behavioral Sciences, 6Einstein Cancer Center, and 7Rose F. Kennedy Center for Research in Mental Retardation and Developmental Disabilities, Albert Einstein College of Medicine, Bronx, New York 10461, and 8Department of Neuroscience and Cell Biology, University of Medicine and Dentistry of New Jersey, Robert Wood Johnson Medical School, and 9Undergraduate Research Program RISE at Rutgers, The State University of New Jersey, Piscataway, New Jersey 08854 Recent studies suggest that specific neural basic helix-loop-helix (HLH; i.e., Olig1 and Olig2, Mash1), associated inhibitory HLH (i.e., Id2 andId4),high-mobilitygroupdomain(i.e.,Sox10),andhomeodomain(i.e.,Nkx2.2)transcriptionfactorsareinvolvedinoligodendrocyte (OL) lineage specification and progressive stages of maturation including myelination. However, the developmental interplay among theselineage-selectivedeterminants,inacell-andmaturationalstage-specificcontext,hasnotyetbeendefined.Weshowhereinvivoand in vitro developmental -

Investigating the Gene Regulatory Network Underlying Caudal Hindbrain Specification in Embryonic Zebrafish
University of Massachusetts Medical School eScholarship@UMMS GSBS Dissertations and Theses Graduate School of Biomedical Sciences 2018-06-13 Investigating the Gene Regulatory Network Underlying Caudal Hindbrain Specification in Embryonic Zebrafish Priyanjali Ghosh University of Massachusetts Medical School Let us know how access to this document benefits ou.y Follow this and additional works at: https://escholarship.umassmed.edu/gsbs_diss Part of the Developmental Biology Commons Repository Citation Ghosh P. (2018). Investigating the Gene Regulatory Network Underlying Caudal Hindbrain Specification in Embryonic Zebrafish. GSBS Dissertations and Theses. https://doi.org/10.13028/M2S971. Retrieved from https://escholarship.umassmed.edu/gsbs_diss/979 This material is brought to you by eScholarship@UMMS. It has been accepted for inclusion in GSBS Dissertations and Theses by an authorized administrator of eScholarship@UMMS. For more information, please contact [email protected]. INVESTIGATING THE GENE REGULATORY NETWORK UNDERLYING CAUDAL HINDBRAIN SPECIFICATION IN EMBRYONIC ZEBRAFISH A Dissertation Presented By Priyanjali Ghosh Submitted to the Faculty of the University of Massachusetts Graduate School of Biomedical Sciences, Worcester in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY June 13, 2018 Department of Biochemistry and Molecular Pharmacology INVESTIGATING THE GENE REGULATORY NETWORK UNDERLYING CAUDAL HINDBRAIN SPECIFICATION IN EMBRYONIC ZEBRAFISH A Dissertation Presented By Priyanjali Ghosh This work was undertaken in the Graduate School of Biomedical Sciences Department of Biochemistry and Molecular Pharmacology Under the mentorship of Charles G. Sagerström, PhD, Thesis Advisor Michael M. Francis, PhD Member of Committee Lawrence J. Hayward, MD, PhD, Member of Committee Scot A. Wolfe, PhD, Member of Committee Sarah McMenamin, PhD, External Member of Committee Alonzo H. -

Frequency and Clinical Characteristics of Intrachromosomal Amplification of Chromosome 21 in Korean Childhood B-Lineage Acute Lymphoblastic Leukemia
Original Article Diagnostic Genetics Ann Lab Med 2016;36:475-480 http://dx.doi.org/10.3343/alm.2016.36.5.475 ISSN 2234-3806 • eISSN 2234-3814 Frequency and Clinical Characteristics of Intrachromosomal Amplification of Chromosome 21 in Korean Childhood B-lineage Acute Lymphoblastic Leukemia Jieun Kim, M.D.1, Chuhl Joo Lyu, M.D.2, Saeam Shin, M.D.3, Seung-Tae Lee, M.D.1, and Jong Rak Choi, M.D.1 Department of Laboratory Medicine1, Yonsei University College of Medicine, Seoul; Department of Pediatrics2, Yonsei Cancer Research Center, Yonsei University College of Medicine, Seoul; Department of Laboratory Medicine3, Hallym University College of Medicine, Kangnam Sacred Heart Hospital, Seoul, Korea Background: Intrachromosomal amplification of chromosome 21 (iAMP21) is known to Received: December 1, 2015 be associated with poor prognosis in B-cell ALL (B-ALL). To determine the frequency and Revision received: March 30, 2016 Accepted: May 11, 2016 clinical characteristics of iAMP21 in Korean B-ALL patients, we performed FISH and mul- tiplex ligation-dependent probe amplification (MLPA) analyses. Corresponding author: Seung-Tae Lee Department of Laboratory Medicine, Yonsei Methods: A total of 102 childhood B-ALL patients were screened with ETV6-RUNX1 FISH University College of Medicine, 50-1 probes (Abbott Molecular, USA). The presence of an iAMP21 was confirmed by using Yonsei-ro, Seodaemun-gu, Seoul 03722, Korea MLPA P327 iAMP21-ERG probemix (MRC Holland, The Netherlands). Tel: +82-2-2228-2450 Results: iAMP21 was detected in one of the screened B-ALL patients (1/102 patients, Fax: +82-2-364-1583 E-mail: [email protected] 1.0%) who presented the ALL immunophenotype and complex karyotype at initial diagno- sis. -

Genome-Wide Screen Reveals WNT11, a Non-Canonical WNT Gene, As a Direct Target of ETS Transcription Factor ERG
Oncogene (2011) 30, 2044–2056 & 2011 Macmillan Publishers Limited All rights reserved 0950-9232/11 www.nature.com/onc ORIGINAL ARTICLE Genome-wide screen reveals WNT11, a non-canonical WNT gene, as a direct target of ETS transcription factor ERG LH Mochmann1, J Bock1, J Ortiz-Ta´nchez1, C Schlee1, A Bohne1, K Neumann2, WK Hofmann3, E Thiel1 and CD Baldus1 1Department of Hematology and Oncology, Charite´, Campus Benjamin Franklin, Berlin, Germany; 2Institute for Biometrics and Clinical Epidemiology, Charite´, Campus Mitte, Berlin, Germany and 3Department of Hematology and Oncology, University Hospital Mannheim, Mannheim, Germany E26 transforming sequence-related gene (ERG) is a current therapies in acute leukemia patients with poor transcription factor involved in normal hematopoiesis prognosis characterized by high ERG mRNA expression. and is dysregulated in leukemia. ERG mRNA over- Oncogene (2011) 30, 2044–2056; doi:10.1038/onc.2010.582; expression was associated with poor prognosis in a subset published online 17 January 2011 of patients with T-cell acute lymphoblastic leukemia (T-ALL) and acute myeloid leukemia (AML). Herein, a Keywords: ETS-related gene (ERG); WNT11; acute genome-wide screen of ERG target genes was conducted leukemia; ERG target genes; 6-bromoindirubin-3-oxime by chromatin immunoprecipitation-on-chip (ChIP-chip) in (BIO); morphological transformation Jurkat cells. In this screen, 342 significant annotated genes were derived from this global approach. Notably, ERG-enriched targets included WNT signaling genes: WNT11, WNT2, WNT9A, CCND1 and FZD7. Further- more, chromatin immunoprecipitation (ChIP) of normal Introduction and primary leukemia bone marrow material also confirmed WNT11 as a target of ERG in six of seven The erythroblastosis virus E26 transforming sequence patient samples. -

Oxygen Impairs Oligodendroglial Development Via Oxidative Stress
www.nature.com/scientificreports OPEN Oxygen impairs oligodendroglial development via oxidative stress and reduced expression of HIF-1α Received: 20 June 2016 Christina Brill1, Till Scheuer1,2, Christoph Bührer1, Stefanie Endesfelder1 & Thomas Schmitz1 Accepted: 18 January 2017 The premature increase of oxygen tension may contribute to oligodendroglial precursor cell (OPC) Published: 23 February 2017 damage in preterm infants. Fetal OPCs are exposed to low oxygen tissue tensions not matched when cells are cultured in room air. Maturation (A2B5, O4, O1, MBP, CNP, arborization), oxidative stress (nitrotyrosine Western blot, NRF2 and SOD2 expression), apoptosis (TUNEL), proliferation (Ki67), and expression of transcription factors regulated by Hypoxia-Inducible-Factor-1-alpha (Hif-1α) expressed in OPCs (Olig1, Olig2, Sox9, Sox10) were assessed in rat OPCs and OLN93 cells cultured at 5% O2 and 21% O2. Influences of Hif-1α were investigated by Hif-1α luciferase reporter assays and Hif-1α- knockdown experiments. At 21% O2, cell proliferation was decreased and process arborization of OPCs was reduced. Expression of MBP, CNP, Olig1, Sox9 and Sox10 was lower at 21% O2, while Nrf2, SOD2, nitrotyrosine were increased. Apoptosis was unchanged. Luciferease reporter assay in OLN93 cells indicated increased Hif-1α activity at 5% O2. In OLN93 cells at 5% O2, Hif-1α knockdown decreased the expression of MBP and CNP, similar to that observed at 21% O2. These data indicate that culturing OPCs at 21% O2 negatively affects development and maturation. Both enhanced oxidative stress and reduced expression of Hif-1α-regulated genes contribute to these hyperoxia-induced changes. In addition to its fundamental role in energy metabolism, oxygen serves as a regulator of cellular develop- ment.