Associated Fibroblasts in Non-Small Cell Lung Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Análise Integrativa De Perfis Transcricionais De Pacientes Com
UNIVERSIDADE DE SÃO PAULO FACULDADE DE MEDICINA DE RIBEIRÃO PRETO PROGRAMA DE PÓS-GRADUAÇÃO EM GENÉTICA ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Ribeirão Preto – 2012 ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Tese apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo para obtenção do título de Doutor em Ciências. Área de Concentração: Genética Orientador: Prof. Dr. Eduardo Antonio Donadi Co-orientador: Prof. Dr. Geraldo A. S. Passos Ribeirão Preto – 2012 AUTORIZO A REPRODUÇÃO E DIVULGAÇÃO TOTAL OU PARCIAL DESTE TRABALHO, POR QUALQUER MEIO CONVENCIONAL OU ELETRÔNICO, PARA FINS DE ESTUDO E PESQUISA, DESDE QUE CITADA A FONTE. FICHA CATALOGRÁFICA Evangelista, Adriane Feijó Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. Ribeirão Preto, 2012 192p. Tese de Doutorado apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo. Área de Concentração: Genética. Orientador: Donadi, Eduardo Antonio Co-orientador: Passos, Geraldo A. 1. Expressão gênica – microarrays 2. Análise bioinformática por module maps 3. Diabetes mellitus tipo 1 4. Diabetes mellitus tipo 2 5. Diabetes mellitus gestacional FOLHA DE APROVAÇÃO ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. -

Methylglyoxal Down-Regulates the Expression of Cell Cycle Associated
www.nature.com/scientificreports OPEN Methylglyoxal down-regulates the expression of cell cycle associated genes and activates the p53 Received: 8 May 2018 Accepted: 12 December 2018 pathway in human umbilical vein Published: xx xx xxxx endothelial cells Jana D. Braun1, Diego O. Pastene1, Annette Breedijk1, Angelica Rodriguez1, Björn B. Hofmann1, Carsten Sticht2, Elke von Ochsenstein3, Heike Allgayer3, Jacob van den Born4, Stephan Bakker4, Sibylle J. Hauske1, Bernhard K. Krämer1, Benito A. Yard1 & Thomas Albrecht1 Although methylglyoxal (MGO) has emerged as key mediator of diabetic microvascular complications, the infuence of MGO on the vascular transcriptome has not thoroughly been assessed. Since diabetes is associated with low grade infammation causing sustained nuclear factor-kappa B (NF-κB) activation, the current study addressed 1) to what extent MGO changes the transcriptome of human umbilical vein endothelial cells (HUVECs) exposed to an infammatory milieu, 2) what are the dominant pathways by which these changes occur and 3) to what extent is this afected by carnosine, a putative scavenger of MGO. Microarray analysis revealed that exposure of HUVECs to high MGO concentrations signifcantly changes gene expression, characterized by prominent down-regulation of cell cycle associated genes and up-regulation of heme oxygenase-1 (HO-1). KEGG-based pathway analysis identifed six signifcantly enriched pathways of which the p53 pathway was the most afected. No signifcant enrichment of infammatory pathways was found, yet, MGO did inhibit VCAM-1 expression in Western blot analysis. Carnosine signifcantly counteracted MGO-mediated changes in a subset of diferentially expressed genes. Collectively, our results suggest that MGO initiates distinct transcriptional changes in cell cycle/apoptosis genes, which may explain MGO toxicity at high concentrations. -

Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name Protein EVI2B Gene Symbol Evi2b Organism Mouse Gene Summary Description Not Available Gene Aliases Evi-2, Evi2 RefSeq Accession No. NC_000077.6, NT_096135.6 UniGene ID Not Available Ensembl Gene ID ENSMUSG00000093938 Entrez Gene ID 216984 Assay Information Unique Assay ID qMmuCEP0062720 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENSMUST00000179322, ENSMUST00000170422 Amplicon Context Sequence ACAGTATAATCATGAGTATAGCTACCAACATAGAAATTATAATAGTACCGATTAAT ATGGCAGCAATTGCATTGTGGTTGTTTTTATGAGCAGTTCCTTTGCCACTTGGTG GTTCTGAAGTGAGTGGTGGTGGTGGTGGTGGTG Amplicon Length (bp) 114 Chromosome Location 11:79516072-79516215 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 95 R2 0.9997 cDNA Cq 22.69 cDNA Tm (Celsius) 79 gDNA Cq 24.61 Specificity (%) 49.93 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report Evi2b, Mouse Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Mouse Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. Detected Coding Transcript(s) This is a list of the Ensembl transcript ID(s) that this assay will detect. -

CALIFORNIA STATE UNIVERSITY, NORTHRIDGE Bioinformatic
CALIFORNIA STATE UNIVERSITY, NORTHRIDGE Bioinformatic Comparison of the EVI2A Promoter and Coding Regions A thesis submitted in partial fulfillment of the requirements for the degree of Master of Science in Biology By Max Weinstein May 2020 The thesis of Max Weinstein is approved: Professor Rheem D. Medh Date Professor Virginia Oberholzer Vandergon Date Professor Cindy Malone, Chair Date California State University Northridge ii Table of Contents Signature Page ii List of Figures v Abstract vi Introduction 1 Ecotropic Viral Integration Site 2A, a Gene within a Gene 7 Materials and Methods 11 PCR and Cloning of Recombinant Plasmid 11 Transformation and Cell Culture 13 Generation of Deletion Constructs 15 Transfection and Luciferase Assay 16 Identification of Transcription Factor Binding Sites 17 Determination of Region for Analysis 17 Multiple Sequence Alignment (MSA) 17 Model Testing 18 Tree Construction 18 Promoter and CDS Conserved Motif Search 19 Results and Discussion 20 Choice of Species for Analysis 20 Mapping of Potential Transcription Factor Binding Sites 20 Confirmation of Plasmid Generation through Gel Electrophoresis 21 Analysis of Deletion Constructs by Transient Transfection 22 EVI2A Coding DNA Sequence Phylogenetics 23 EVI2A Promoter Phylogenetics 23 EVI2A Conserved Leucine Zipper 25 iii EVI2A Conserved Casein kinase II phosphorylation site 26 EVI2A Conserved Sox-5 Binding Site 26 EVI2A Conserved HLF Binding Site 27 EVI2A Conserved cREL Binding Site 28 EVI2A Conserved CREB Binding Site 28 Summary 29 Appendix: Figures 30 Literature Cited 44 iv List of Figures Figure 1. EVI2A is nested within the gene NF1 30 Figure 2 Putative Transcriptions Start Site 31 Figure 3. Gel Electrophoresis of pGC Blue cloned, Restriction Digested Plasmid 32 Figure 4. -

Identification and Validation of Genes Affecting Aortic Lesions in Mice
Identification and validation of genes affecting aortic lesions in mice Xia Yang, … , Aldons J. Lusis, Pek Yee Lum J Clin Invest. 2010;120(7):2414-2422. https://doi.org/10.1172/JCI42742. Research Article Cardiology Atherosclerosis represents the most significant risk factor for coronary artery disease (CAD), the leading cause of death in developed countries. To better understand the pathogenesis of atherosclerosis, we applied a likelih ood-based model selection method to infer gene-disease causality relationships for the aortic lesion trait in a segregating mouse population demonstrating a spectrum of susceptibility to developing atherosclerotic lesions. We identified 292 genes that tested causal for aortic lesions from liver and adipose tissues of these mice, and we experimentally validated one of these candidate causal genes, complement component 3a receptor 1 (C3ar1), using a knockout mouse model. We also found that genes identified by this method overlapped with genes progressively regulated in the aortic arches of 2 mouse models of atherosclerosis during atherosclerotic lesion development. By comparing our gene set with findings from public human genome-wide association studies (GWAS) of CAD and related traits, we found that 5 genes identified by our study overlapped with published studies in humans in which they were identified as risk factors for multiple atherosclerosis- related pathologies, including myocardial infarction, serum uric acid levels, mean platelet volume, aortic root size, and heart failure. Candidate causal genes were also found to be enriched with CAD risk polymorphisms identified by the Wellcome Trust Case Control Consortium (WTCCC). Our findings therefore validate the ability of causality testing procedures to […] Find the latest version: https://jci.me/42742/pdf Research article Identification and validation of genes affecting aortic lesions in mice Xia Yang,1,2 Larry Peterson,3 Rolf Thieringer,4 Joshua L. -

WNT16 Is a New Marker of Senescence
Table S1. A. Complete list of 177 genes overexpressed in replicative senescence Value Gene Description UniGene RefSeq 2.440 WNT16 wingless-type MMTV integration site family, member 16 (WNT16), transcript variant 2, mRNA. Hs.272375 NM_016087 2.355 MMP10 matrix metallopeptidase 10 (stromelysin 2) (MMP10), mRNA. Hs.2258 NM_002425 2.344 MMP3 matrix metallopeptidase 3 (stromelysin 1, progelatinase) (MMP3), mRNA. Hs.375129 NM_002422 2.300 HIST1H2AC Histone cluster 1, H2ac Hs.484950 2.134 CLDN1 claudin 1 (CLDN1), mRNA. Hs.439060 NM_021101 2.119 TSPAN13 tetraspanin 13 (TSPAN13), mRNA. Hs.364544 NM_014399 2.112 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 2.070 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 2.026 DCBLD2 discoidin, CUB and LCCL domain containing 2 (DCBLD2), mRNA. Hs.203691 NM_080927 2.007 SERPINB2 serpin peptidase inhibitor, clade B (ovalbumin), member 2 (SERPINB2), mRNA. Hs.594481 NM_002575 2.004 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.989 OBFC2A Oligonucleotide/oligosaccharide-binding fold containing 2A Hs.591610 1.962 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.947 PLCB4 phospholipase C, beta 4 (PLCB4), transcript variant 2, mRNA. Hs.472101 NM_182797 1.934 PLCB4 phospholipase C, beta 4 (PLCB4), transcript variant 1, mRNA. Hs.472101 NM_000933 1.933 KRTAP1-5 keratin associated protein 1-5 (KRTAP1-5), mRNA. Hs.534499 NM_031957 1.894 HIST2H2BE histone cluster 2, H2be (HIST2H2BE), mRNA. Hs.2178 NM_003528 1.884 CYTL1 cytokine-like 1 (CYTL1), mRNA. Hs.13872 NM_018659 tumor necrosis factor receptor superfamily, member 10d, decoy with truncated death domain (TNFRSF10D), 1.848 TNFRSF10D Hs.213467 NM_003840 mRNA. -

Review Article J Med Genet: First Published As 10.1136/Jmg.33.1.2 on 1 January 1996
2 J7 Med Genet 1996;33:2-17 Review article J Med Genet: first published as 10.1136/jmg.33.1.2 on 1 January 1996. Downloaded from Molecular genetics of neurofibromatosis type 1 (NFl) Ming Hong Shen, Peter S Harper, Meena Upadhyaya Abstract several discrete entities.'` At least two dis- Neurofibromatosis type 1 (NF1), also tinctive forms, neurofibromatosis type 1 (NF1) called von Recklinghausen disease or peri- and neurofibromatosis type 2 (NF2), were re- pheral neurofibromatosis, is a common cognised and later confirmed by the cloning of autosomal dominant disorder char- two separate genes, the NF1 gene on chro- acterised by multiple neurofibromas, cafe mosome 1746 and the NF2 gene on chro- au lait spots, and Lisch nodules ofthe iris, mosome 22.78 Other NF related syndromes with a variable clinical expression. The were also reported, such as segmental NF,9 gene responsible for this condition, NF1, Watson syndrome,'0 Noonan syndrome," has been isolated by positional cloning. spinal NF,'2 familial cafe au lait spots It spans over 350kb of genomic DNA in (CLS)," and Schwannomatosis.'4 It remains chromosomal region 17qll.2 and encodes to be determined whether such disorders are an mRNA of 11-13 kb containing at least genetically discrete. 59 exons. NFI is widely expressed in a NFl, also called von Recklinghausen disease variety of human and rat tissues. Four or peripheral neurofibromatosis, is a common alternatively spliced NFI transcripts have autosomal dominant disorder affecting about been identified. Three of these transcript 1 in 3000 to 5000 people. It exhibits full pene- isoforms (each with an extra exon: 9br, trance and a high mutation rate with 30 to 23a, and 48a, respectively) show differ- 50% ofNFl patients representing a new muta- ential expression to some extent in various tion. -

Identification of Expression Qtls Targeting Candidate Genes For
ISSN: 2378-3648 Salleh et al. J Genet Genome Res 2018, 5:035 DOI: 10.23937/2378-3648/1410035 Volume 5 | Issue 1 Journal of Open Access Genetics and Genome Research RESEARCH ARTICLE Identification of Expression QTLs Targeting Candidate Genes for Residual Feed Intake in Dairy Cattle Using Systems Genomics Salleh MS1,2, Mazzoni G2, Nielsen MO1, Løvendahl P3 and Kadarmideen HN2,4* 1Department of Veterinary and Animal Sciences, Faculty of Health and Medical Sciences, University of Copenhagen, Denmark Check for 2Department of Bio and Health Informatics, Technical University of Denmark, Lyngby, Denmark updates 3Department of Molecular Biology and Genetics-Center for Quantitative Genetics and Genomics, Aarhus University, AU Foulum, Tjele, Denmark 4Department of Applied Mathematics and Computer Science, Technical University of Denmark, Lyngby, Denmark *Corresponding author: Kadarmideen HN, Department of Applied Mathematics and Computer Science, Technical University of Denmark, DK-2800, Kgs. Lyngby, Denmark, E-mail: [email protected] Abstract body weight gain and net merit). The eQTLs and biological pathways identified in this study improve our understanding Background: Residual feed intake (RFI) is the difference of the complex biological and genetic mechanisms that de- between actual and predicted feed intake and an important termine FE traits in dairy cattle. The identified eQTLs/genet- factor determining feed efficiency (FE). Recently, 170 can- ic variants can potentially be used in new genomic selection didate genes were associated with RFI, but no expression methods that include biological/functional information on quantitative trait loci (eQTL) mapping has hitherto been per- SNPs. formed on FE related genes in dairy cows. In this study, an integrative systems genetics approach was applied to map Keywords eQTLs in Holstein and Jersey cows fed two different diets to eQTL, RNA-seq, Genotype, Data integration, Systems improve identification of candidate genes for FE. -

ID AKI Vs Control Fold Change P Value Symbol Entrez Gene Name *In
ID AKI vs control P value Symbol Entrez Gene Name *In case of multiple probesets per gene, one with the highest fold change was selected. Fold Change 208083_s_at 7.88 0.000932 ITGB6 integrin, beta 6 202376_at 6.12 0.000518 SERPINA3 serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 3 1553575_at 5.62 0.0033 MT-ND6 NADH dehydrogenase, subunit 6 (complex I) 212768_s_at 5.50 0.000896 OLFM4 olfactomedin 4 206157_at 5.26 0.00177 PTX3 pentraxin 3, long 212531_at 4.26 0.00405 LCN2 lipocalin 2 215646_s_at 4.13 0.00408 VCAN versican 202018_s_at 4.12 0.0318 LTF lactotransferrin 203021_at 4.05 0.0129 SLPI secretory leukocyte peptidase inhibitor 222486_s_at 4.03 0.000329 ADAMTS1 ADAM metallopeptidase with thrombospondin type 1 motif, 1 1552439_s_at 3.82 0.000714 MEGF11 multiple EGF-like-domains 11 210602_s_at 3.74 0.000408 CDH6 cadherin 6, type 2, K-cadherin (fetal kidney) 229947_at 3.62 0.00843 PI15 peptidase inhibitor 15 204006_s_at 3.39 0.00241 FCGR3A Fc fragment of IgG, low affinity IIIa, receptor (CD16a) 202238_s_at 3.29 0.00492 NNMT nicotinamide N-methyltransferase 202917_s_at 3.20 0.00369 S100A8 S100 calcium binding protein A8 215223_s_at 3.17 0.000516 SOD2 superoxide dismutase 2, mitochondrial 204627_s_at 3.04 0.00619 ITGB3 integrin, beta 3 (platelet glycoprotein IIIa, antigen CD61) 223217_s_at 2.99 0.00397 NFKBIZ nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta 231067_s_at 2.97 0.00681 AKAP12 A kinase (PRKA) anchor protein 12 224917_at 2.94 0.00256 VMP1/ mir-21likely ortholog -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

Microarray Bioinformatics and Its Applications to Clinical Research
Microarray Bioinformatics and Its Applications to Clinical Research A dissertation presented to the School of Electrical and Information Engineering of the University of Sydney in fulfillment of the requirements for the degree of Doctor of Philosophy i JLI ··_L - -> ...·. ...,. by Ilene Y. Chen Acknowledgment This thesis owes its existence to the mercy, support and inspiration of many people. In the first place, having suffering from adult-onset asthma, interstitial cystitis and cold agglutinin disease, I would like to express my deepest sense of appreciation and gratitude to Professors Hong Yan and David Levy for harbouring me these last three years and providing me a place at the University of Sydney to pursue a very meaningful course of research. I am also indebted to Dr. Craig Jin, who has been a source of enthusiasm and encouragement on my research over many years. In the second place, for contexts concerning biological and medical aspects covered in this thesis, I am very indebted to Dr. Ling-Hong Tseng, Dr. Shian-Sehn Shie, Dr. Wen-Hung Chung and Professor Chyi-Long Lee at Change Gung Memorial Hospital and University of Chang Gung School of Medicine (Taoyuan, Taiwan) as well as Professor Keith Lloyd at University of Alabama School of Medicine (AL, USA). All of them have contributed substantially to this work. In the third place, I would like to thank Mrs. Inge Rogers and Mr. William Ballinger for their helpful comments and suggestions for the writing of my papers and thesis. In the fourth place, I would like to thank my swim coach, Hirota Homma. -
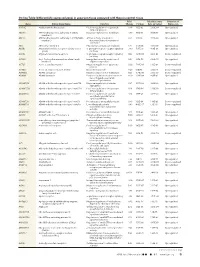
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18