Coding Strategy of the Genome of Chilo Iridescent Virus
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Artemia Spp., a Susceptible Host and Vector for Lymphocystis Disease Virus
viruses Article Artemia spp., a Susceptible Host and Vector for Lymphocystis Disease Virus Estefania J. Valverde, Alejandro M. Labella, Juan J. Borrego and Dolores Castro * Departamento de Microbiología, Facultad de Ciencias, Universidad de Málaga, 29017 Málaga, Spain; [email protected] (E.J.V.); [email protected] (A.M.L.); [email protected] (J.J.B.) * Correspondence: [email protected]; Tel.: +34-952134214 Received: 8 May 2019; Accepted: 30 May 2019; Published: 1 June 2019 Abstract: Different developmental stages of Artemia spp. (metanauplii, juveniles and adults) were bath-challenged with two isolates of the Lymphocystis disease virus (LCDV), namely, LCDV SA25 (belonging to the species Lymphocystis disease virus 3) and ATCC VR-342 (an unclassified member of the genus Lymphocystivirus). Viral quantification and gene expression were analyzed by qPCR at different times post-inoculation (pi). In addition, infectious titres were determined at 8 dpi by integrated cell culture (ICC)-RT-PCR, an assay that detects viral mRNA in inoculated cell cultures. In LCDV-challenged Artemia, the viral load increased by 2–3 orders of magnitude (depending on developmental stage and viral isolate) during the first 8–12 dpi, with viral titres up to 2.3 102 × Most Probable Number of Infectious Units (MPNIU)/mg. Viral transcripts were detected in the infected Artemia, relative expression values showed a similar temporal evolution in the different experimental groups. Moreover, gilthead seabream (Sparus aurata) fingerlings were challenged by feeding on LCDV-infected metanauplii. Although no Lymphocystis symptoms were observed in the fish, the number of viral DNA copies was significantly higher at the end of the experimental trial and major capsid protein (mcp) gene expression was consistently detected. -

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

Disease of Aquatic Organisms 105:1
Vol. 105: 1–8, 2013 DISEASES OF AQUATIC ORGANISMS Published July 9 doi: 10.3354/dao02594 Dis Aquat Org Megalocytivirus infection in orbiculate batfish Platax orbicularis Preeyanan Sriwanayos1, Ruth Francis-Floyd1,2, Mark F. Stidworthy3, Barbara D. Petty1,2, Karen Kelley4, Thomas B. Waltzek5,* 1Program in Fisheries and Aquatic Sciences, School of Forestry Resources and Conservation, University of Florida, Gainesville, Florida 32653, USA 2Department of Large Animal Clinical Sciences, College of Veterinary Medicine, University of Florida, Gainesville, Florida 32610, USA 3International Zoo Veterinary Group, Station House, Parkwood Street, Keighley, West Yorkshire BD21 4NQ, UK 4Interdisciplinary Center for Biotechnology Research (ICBR), Cellomics Division, Electron Microscopy and Bio-imaging Core Laboratory, University of Florida, Gainesville, Florida 32611, USA 5Department of Infectious Diseases and Pathology, College of Veterinary Medicine, University of Florida, Gainesville, Florida 32608, USA ABSTRACT: Megalocytiviruses cause systemic disease in both marine and freshwater fishes, neg- atively impacting ornamental and food fish aquaculture. In this report, we characterize a megalo- cytivirus infection in a captive marine ornamental fish, the orbiculate batfish Platax orbicularis. Histologic examination revealed cytomegalic cells characterized by strongly basophilic granular intracytoplasmic inclusions within various organs. Transmission electron microscopy revealed icosahedral virus particles within the cytoplasm of cytomegalic cells consistent -

Enzymatic Manufacture of Deoxythymidine-5'-Triphosphate with Permeable Intact Cells of E. Coli Coexpressing Thymidylate Kinase
J. Microbiol. Biotechnol. (2015), 25(12), 2034–2042 http://dx.doi.org/10.4014/jmb.1506.06020 Research Article Review jmb Enzymatic Manufacture of Deoxythymidine-5’-Triphosphate with Permeable Intact Cells of E. coli Coexpressing Thymidylate Kinase and Acetate Kinase Jiao Zhang, Yahui Qian, Qingbao Ding*, and Ling Ou* State Key Laboratory of Bioreactor Engineering, East China University of Science and Technology, Shanghai 200237, P.R. China Received: June 8, 2015 Revised: July 26, 2015 A one-pot process of enzymatic synthesis of deoxythymidine-5’-triphosphate (5’-dTTP) Accepted: September 11, 2015 employing whole cells of recombinant Escherichia coli coexpressing thymidylate kinase (TMKase) and acetate kinase (ACKase) was developed. Genes tmk and ack from E. coli were First published online cloned and inserted into pET28a(+), and then transduced into E. coli BL21 (DE3) to form September 15, 2015 recombinant strain pTA in which TMKase and ACKase were simultaneously overexpressed. It *Corresponding authors was found that the relative residual specific activities of TMKase and ACKase, in pTA Q.D. pretreated with 20 mM ethylene diamine tetraacetic acid (EDTA) at 25oC for 30 min, were 94% Phone: +86-21-64250676; Fax: +86-21-64250676; and 96%, respectively. The yield of 5’-dTTP reached above 94% from 5 mM deoxythymidine E-mail: [email protected] 5’-monophosphate (5’-dTMP) and 15 mM acetyl phosphate catalyzed with intact cells of pTA L.O. Phone: +86-21-64253257; pretreated with EDTA. The process was so effective that only 0.125 mM adenosine-5’- Fax: +86-21-64253257; triphosphate was sufficient to deliver the phosphate group from acetyl phosphate to dTMP E-mail: [email protected] and dTDP. -

The DNA Virus Invertebrate Iridescent Virus 6 Is a Target of the Drosophila Rnai Machinery
The DNA virus Invertebrate iridescent virus 6 is a target of the Drosophila RNAi machinery Alfred W. Bronkhorsta,1, Koen W. R. van Cleefa,1, Nicolas Vodovarb,2, Ikbal_ Agah Ince_ c,d,e, Hervé Blancb, Just M. Vlakc, Maria-Carla Salehb,3, and Ronald P. van Rija,3 aDepartment of Medical Microbiology, Nijmegen Centre for Molecular Life Sciences, Nijmegen Institute for Infection, Inflammation, and Immunity, Radboud University Nijmegen Medical Centre, 6500 HB Nijmegen, The Netherlands; bViruses and RNA Interference Group, Institut Pasteur, Centre National de la Recherche Scientifique, Unité de Recherche Associée 3015, 75015 Paris, France; cLaboratory of Virology, Wageningen University, 6708 PB Wageningen, The Netherlands; dDepartment of Genetics and Bioengineering, Yeditepe University, Istanbul 34755, Turkey; and eDepartment of Biosystems Engineering, Faculty of Engineering, Giresun University, Giresun 28100, Turkey Edited by Peter Palese, Mount Sinai School of Medicine, New York, NY, and approved October 19, 2012 (received for review April 28, 2012) RNA viruses in insects are targets of an RNA interference (RNAi)- sequently, are hypersensitive to virus infection and succumb more based antiviral immune response, in which viral replication inter- rapidly than their wild-type (WT) controls (11–14). mediates or viral dsRNA genomes are processed by Dicer-2 (Dcr-2) Small RNA cloning and next-generation sequencing provide into viral small interfering RNAs (vsiRNAs). Whether dsDNA virus detailed insights into vsiRNA biogenesis. In several studies in infections are controlled by the RNAi pathway remains to be insects, the polarity of the vsiRNA population deviates strongly determined. Here, we analyzed the role of RNAi in DNA virus from the highly skewed distribution of positive strand (+) over infection using Drosophila melanogaster infected with Invertebrate negative (−) viral RNAs that is generally observed in (+) RNA iridescent virus 6 (IIV-6) as a model. -

Table S1. List of Oligonucleotide Primers Used
Table S1. List of oligonucleotide primers used. Cla4 LF-5' GTAGGATCCGCTCTGTCAAGCCTCCGACC M629Arev CCTCCCTCCATGTACTCcgcGATGACCCAgAGCTCGTTG M629Afwd CAACGAGCTcTGGGTCATCgcgGAGTACATGGAGGGAGG LF-3' GTAGGCCATCTAGGCCGCAATCTCGTCAAGTAAAGTCG RF-5' GTAGGCCTGAGTGGCCCGAGATTGCAACGTGTAACC RF-3' GTAGGATCCCGTACGCTGCGATCGCTTGC Ukc1 LF-5' GCAATATTATGTCTACTTTGAGCG M398Arev CCGCCGGGCAAgAAtTCcgcGAGAAGGTACAGATACGc M398Afwd gCGTATCTGTACCTTCTCgcgGAaTTcTTGCCCGGCGG LF-3' GAGGCCATCTAGGCCATTTACGATGGCAGACAAAGG RF-5' GTGGCCTGAGTGGCCATTGGTTTGGGCGAATGGC RF-3' GCAATATTCGTACGTCAACAGCGCG Nrc2 LF-5' GCAATATTTCGAAAAGGGTCGTTCC M454Grev GCCACCCATGCAGTAcTCgccGCAGAGGTAGAGGTAATC M454Gfwd GATTACCTCTACCTCTGCggcGAgTACTGCATGGGTGGC LF-3' GAGGCCATCTAGGCCGACGAGTGAAGCTTTCGAGCG RF-5' GAGGCCTGAGTGGCCTAAGCATCTTGGCTTCTGC RF-3' GCAATATTCGGTCAACGCTTTTCAGATACC Ipl1 LF-5' GTCAATATTCTACTTTGTGAAGACGCTGC M629Arev GCTCCCCACGACCAGCgAATTCGATagcGAGGAAGACTCGGCCCTCATC M629Afwd GATGAGGGCCGAGTCTTCCTCgctATCGAATTcGCTGGTCGTGGGGAGC LF-3' TGAGGCCATCTAGGCCGGTGCCTTAGATTCCGTATAGC RF-5' CATGGCCTGAGTGGCCGATTCTTCTTCTGTCATCGAC RF-3' GACAATATTGCTGACCTTGTCTACTTGG Ire1 LF-5' GCAATATTAAAGCACAACTCAACGC D1014Arev CCGTAGCCAAGCACCTCGgCCGAtATcGTGAGCGAAG D1014Afwd CTTCGCTCACgATaTCGGcCGAGGTGCTTGGCTACGG LF-3' GAGGCCATCTAGGCCAACTGGGCAAAGGAGATGGA RF-5' GAGGCCTGAGTGGCCGTGCGCCTGTGTATCTCTTTG RF-3' GCAATATTGGCCATCTGAGGGCTGAC Kin28 LF-5' GACAATATTCATCTTTCACCCTTCCAAAG L94Arev TGATGAGTGCTTCTAGATTGGTGTCggcGAAcTCgAGCACCAGGTTG L94Afwd CAACCTGGTGCTcGAgTTCgccGACACCAATCTAGAAGCACTCATCA LF-3' TGAGGCCATCTAGGCCCACAGAGATCCGCTTTAATGC RF-5' CATGGCCTGAGTGGCCAGGGCTAGTACGACCTCG -

Genome Analysis of Ranavirus Frog Virus 3Isolated from American Bullfrog
www.nature.com/scientificreports OPEN Genome analysis of Ranavirus frog virus 3 isolated from American Bullfrog (Lithobates catesbeianus) in South America Marcelo Candido 1*, Loiane Sampaio Tavares1, Anna Luiza Farias Alencar2, Cláudia Maris Ferreira3, Sabrina Ribeiro de Almeida Queiroz1, Andrezza Maria Fernandes1 & Ricardo Luiz Moro de Sousa1 Ranaviruses (family Iridoviridae) cause important diseases in cold-blooded vertebrates. In addition, some occurrences indicate that, in this genus, the same virus can infect animals from diferent taxonomic groups. A strain isolated from a Ranavirus outbreak (2012) in the state of Sao Paulo, Brazil, had its genome sequenced and presented 99.26% and 36.85% identity with samples of Frog virus 3 (FV3) and Singapore grouper iridovirus (SGIV) ranaviruses, respectively. Eight potential recombination events among the analyzed sample and reference FV3 samples were identifed, including a recombination with Bohle iridovirus (BIV) sample from Oceania. The analyzed sample presented several rearrangements compared to FV3 reference samples from North America and European continent. We report for the frst time the complete genome of Ranavirus FV3 isolated from South America, these results contribute to a greater knowledge related to evolutionary events of potentially lethal infectious agent for cold-blooded animals. Among the major viral pathogens, worldwide distributed and recent history, Ranavirus (Rv) is highlighted, on which, studies in South America remain limited. Rv are part of the family Iridoviridae that is divided into fve genera, of which three are considered more relevant by infectious severity in aquatic and semi-aquatic animals: Lymphocystivirus, Megalocytivirus and Rv. Tey are enveloped and unenveloped viruses, showing double-stranded DNA whose genome ranges from 103 to 220 kbp. -

Overexpression of DNA Polymerase @Jsensitizes Mammalian Cells to 2',3'
ICANCERRESEARCH57.I10-I 16,JanuaryI. I997J Overexpression of DNA Polymerase @JSensitizesMammalian Cells to 2',3'-Deoxycytidine and 3'-Azido-3' .deoxythymidine' Khalil Bouayadi, Jean-Sébastien Hoffmann, Pascal Fons, Michele Tiraby, Jean-Paul Reynes, and Christophe Cazaux2 Laboratoire de Microbiologic et de Génétique,UniversitéPaul Sabatier, 118 route de Narbonne, 31062 Toulouse cédex(K. B., P. F., C. C.]; Institut de Pharmacologic et Biologic Structurale. Centre National de Ia Recherche ScienujIque, UPR 9062, 205 route de Narbonne, 31077 Toulouse cédexfl. S. H.!; and Laboratoire CAYLA, ZI. Montaudran, 5 rue J. Rodier, 3/400 Toulouse (M. T., J. P. R.J ABSTRACT nator AZT-MP was also observed (10, 11). In vivo, AZT-MP is incorporated into cellular DNA (12), and it has been suggested that Mammalian DNA polymerase @3is a DNA repair enzyme expressed polymerase 13may play a role in this process (1 1). @ constitutively at a low level. In vitro, purified DNA polymerase (Pol) incorporates the nucleotide analogues 2'-3' deoxycytidine (ddC)-triphos AZT and ddC are antiviral agents currently used in the treatment phate and 3'-azido-3'-deoxythymidine (AZT)-triphosphate Into DNA, of AIDS. These chemotherapeutic drugs target HIV reverse tran causing chain termination. We have tested the possibility ofenhancing the scriptase which incorporates them into HIV DNA, terminating cytotoxicity of these chain terminators against mammalian cells by in DNA polymerization (13). In this study, we hypothesized that creasing the level of Pol (I. Chinese hamster ovary AA8 and murine overexpression of Pol f3, a cellular target of ddC and AZT, might melanoma B16 cell lines were stably transfected with rat pol fi cDNA render these analogues effective also against tumor cells. -

The Lymphocystis Diseases in the Olive Flounder, Paralichthys Olivaceus
Univ. j. zool. Rajshahi Univ. Vol. 26, 2007. pp. 59-62 ISSN 1023-6104 http://journals.sfu.ca/bd/index.php/UJZRU © Rajshahi University Zoological Society The lymphocystis diseases in the Olive flounder, Paralichthys olivaceus Mosharrof Hossain, Seok Ryel Kim and Myung Joo Oh* Division of Food Science and Aqualife Medicine, Chonnam National University Yeosu-550-749, Korea. Abstract: Lymphocystis disease virus (LCDV) is the causative agent of lymphocystis disease, affecting more than 100 teleost species worldwide. Characteristically, LCD is chronic, self limiting and species specific. The greatly hypertrophied cells, called lymphocystis tumor cells, typically occur on the skin, fins and oral region. Lymphocystis cells were ovoid to circular and varied in sizes ranging from 200-250 mm. The lymphocystis disease infected flounder have unsightly appearances that discourage the commercial values. A PCR detection technique was developed to amplify a fragment of LCDV major capsid protein gene (1347bp) which is shortcoming and useful. The PCR result proved that the LCD-virus replicated in the epidermis (fins and skin) not in the spleen, kidney, intestine or brain of Paralichthys olivaceus. Keyword: Lymphocystis disease, LCDV, PCR, Paralichthys olivaceus Introduction diseases have been isolated from more than 100 teleost Lymphocystis disease (LCD) is a chronic, self-limiting, species (Anders, 1989), however the infections and viral disease affecting many species of teleosts virus replication is unknown. LCDV has been studied worldwide. Freshwater, estuarine, and marine fish in for the different isolation and characterization warm-water, and cold-water environments are techniques (Iwamoto et al., 2002; Alonso et al., 2005; susceptible to this disease. In general, lymphocystis is a Cano et al., 2006) that helping shortcoming detection disease of more evolutionarily advanced species of of the disease and to take initiatives to a disease free teleosts, like perches, seabreams and flounders. -
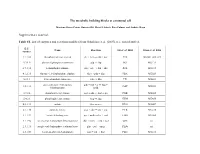
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

Direct Induction of Gj-Specific Transcripts Following Reactivation of the Cdc28 Kinase in the Absence of De Novo Protein Synthesis
Downloaded from genesdev.cshlp.org on October 4, 2021 - Published by Cold Spring Harbor Laboratory Press Direct induction of Gj-specific transcripts following reactivation of the Cdc28 kinase in the absence of de novo protein synthesis Nicholas J. Marini and Steven I. Reed* The Scripps Research Institute, Department of Molecular Biology, La Jolla, California 92037 USA In Saccharomyces cerevisiae, the genes encoding the HO endonuclease, G,-specific cyclins CLNl and CLN2, as well as most proteins involved in DNA synthesis, are periodically transcribed with maximal levels reached in late G,. For HO and the DNA replication genes, cell cycle stage-specific expression has been shown to be dependent on the Cdc28 kinase and passage through START. Here, we show that cells released from cdc28*^ arrest in the presence of cycloheximide show wild-type levels of induction for HO, CLNl, and CDC9 (DNA ligase). Induction is gradual with a significant lag not seen in untreated cells where transcript levels fluctuate coordinately with the cell cycle. This lag may be due, at least in part, to association of the Cdc28 peptide with Gj cyclins to form an active kinase complex because overexpression of CLN2 prior to release in cycloheximide increases the rate of induction for CDC9 and HO. Consistent with this, release from pheromone arrest (where CLNl and CLN2 are not expressed) in cycloheximide shows no induction at all, Transcriptonal activation of CDC9 is likely to be mediated through a conserved promoter element also present in genes for other DNA synthesis enzymes similarly cell cycle regulated. The element contains an intact Mlul restriction enzyme recognition site (consensus ~5'-A/TPuACGCGTNA/T-3'), Insertion of a 20-bp fragment from the CDC9 promoter (containing a Mlul element) upstream of LacZ confers both periodic expression and transcriptional induction in cycloheximide following release from cdc28"' arrest. -

Comparative Analysis of Transcriptional Regulation Patterns: Understanding the Gene Expression Profile in Nucleocytoviricota
pathogens Review Comparative Analysis of Transcriptional Regulation Patterns: Understanding the Gene Expression Profile in Nucleocytoviricota Fernanda Gil de Souza †,Jônatas Santos Abrahão * and Rodrigo Araújo Lima Rodrigues *,† Laboratório de Vírus, Departamento de Microbiologia, Instituto de Ciências Biológicas, Universidade Federal de Minas Gerais, Belo Horizonte, Minas Gerais 31270-901, Brazil; [email protected] * Correspondence: [email protected] (J.S.A.); [email protected] (R.A.L.R.) † These authors contributed equally to this work. Abstract: The nucleocytoplasmic large DNA viruses (NCLDV) possess unique characteristics that have drawn the attention of the scientific community, and they are now classified in the phylum Nucleocytoviricota. They are characterized by sharing many genes and have their own transcriptional apparatus, which provides certain independence from their host’s machinery. Thus, the presence of a robust transcriptional apparatus has raised much discussion about the evolutionary aspects of these viruses and their genomes. Understanding the transcriptional process in NCLDV would provide information regarding their evolutionary history and a better comprehension of the biology of these viruses and their interaction with hosts. In this work, we reviewed NCLDV transcription and performed a comparative functional analysis of the groups of genes expressed at different times of infection of representatives of six different viral families of giant viruses. With this analysis, it was possible to observe