A New AURKC Mutation Causing Macrozoospermia
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Evolution, Expression and Meiotic Behavior of Genes Involved in Chromosome Segregation of Monotremes
G C A T T A C G G C A T genes Article Evolution, Expression and Meiotic Behavior of Genes Involved in Chromosome Segregation of Monotremes Filip Pajpach , Linda Shearwin-Whyatt and Frank Grützner * School of Biological Sciences, The University of Adelaide, Adelaide, SA 5005, Australia; fi[email protected] (F.P.); [email protected] (L.S.-W.) * Correspondence: [email protected] Abstract: Chromosome segregation at mitosis and meiosis is a highly dynamic and tightly regulated process that involves a large number of components. Due to the fundamental nature of chromosome segregation, many genes involved in this process are evolutionarily highly conserved, but duplica- tions and functional diversification has occurred in various lineages. In order to better understand the evolution of genes involved in chromosome segregation in mammals, we analyzed some of the key components in the basal mammalian lineage of egg-laying mammals. The chromosome passenger complex is a multiprotein complex central to chromosome segregation during both mitosis and meio- sis. It consists of survivin, borealin, inner centromere protein, and Aurora kinase B or C. We confirm the absence of Aurora kinase C in marsupials and show its absence in both platypus and echidna, which supports the current model of the evolution of Aurora kinases. High expression of AURKBC, an ancestor of AURKB and AURKC present in monotremes, suggests that this gene is performing all necessary meiotic functions in monotremes. Other genes of the chromosome passenger complex complex are present and conserved in monotremes, suggesting that their function has been preserved Citation: Pajpach, F.; in mammals. -

Program Nr: 1 from the 2004 ASHG Annual Meeting Mutations in A
Program Nr: 1 from the 2004 ASHG Annual Meeting Mutations in a novel member of the chromodomain gene family cause CHARGE syndrome. L.E.L.M. Vissers1, C.M.A. van Ravenswaaij1, R. Admiraal2, J.A. Hurst3, B.B.A. de Vries1, I.M. Janssen1, W.A. van der Vliet1, E.H.L.P.G. Huys1, P.J. de Jong4, B.C.J. Hamel1, E.F.P.M. Schoenmakers1, H.G. Brunner1, A. Geurts van Kessel1, J.A. Veltman1. 1) Dept Human Genetics, UMC Nijmegen, Nijmegen, Netherlands; 2) Dept Otorhinolaryngology, UMC Nijmegen, Nijmegen, Netherlands; 3) Dept Clinical Genetics, The Churchill Hospital, Oxford, United Kingdom; 4) Children's Hospital Oakland Research Institute, BACPAC Resources, Oakland, CA. CHARGE association denotes the non-random occurrence of ocular coloboma, heart defects, choanal atresia, retarded growth and development, genital hypoplasia, ear anomalies and deafness (OMIM #214800). Almost all patients with CHARGE association are sporadic and its cause was unknown. We and others hypothesized that CHARGE association is due to a genomic microdeletion or to a mutation in a gene affecting early embryonic development. In this study array- based comparative genomic hybridization (array CGH) was used to screen patients with CHARGE association for submicroscopic DNA copy number alterations. De novo overlapping microdeletions in 8q12 were identified in two patients on a genome-wide 1 Mb resolution BAC array. A 2.3 Mb region of deletion overlap was defined using a tiling resolution chromosome 8 microarray. Sequence analysis of genes residing within this critical region revealed mutations in the CHD7 gene in 10 of the 17 CHARGE patients without microdeletions, including 7 heterozygous stop-codon mutations. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplementary Information Material and Methods
MCT-11-0474 BKM120: a potent and specific pan-PI3K inhibitor Supplementary Information Material and methods Chemicals The EGFR inhibitor NVP-AEE788 (Novartis), the Jak inhibitor I (Merck Calbiochem, #420099) and anisomycin (Alomone labs, # A-520) were prepared as 50 mM stock solutions in 100% DMSO. Doxorubicin (Adriablastin, Pfizer), EGF (Sigma Ref: E9644), PDGF (Sigma, Ref: P4306) and IL-4 (Sigma, Ref: I-4269) stock solutions were prepared as recommended by the manufacturer. For in vivo administration: Temodal (20 mg Temozolomide capsules, Essex Chemie AG, Luzern) was dissolved in 4 mL KZI/glucose (20/80, vol/vol); Taxotere was bought as 40 mg/mL solution (Sanofi Aventis, France), and prepared in KZI/glucose. Antibodies The primary antibodies used were as follows: anti-S473P-Akt (#9271), anti-T308P-Akt (#9276,), anti-S9P-GSK3β (#9336), anti-T389P-p70S6K (#9205), anti-YP/TP-Erk1/2 (#9101), anti-YP/TP-p38 (#9215), anti-YP/TP-JNK1/2 (#9101), anti-Y751P-PDGFR (#3161), anti- p21Cip1/Waf1 (#2946), anti-p27Kip1 (#2552) and anti-Ser15-p53 (#9284) antibodies were from Cell Signaling Technologies; anti-Akt (#05-591), anti-T32P-FKHRL1 (#06-952) and anti- PDGFR (#06-495) antibodies were from Upstate; anti-IGF-1R (#SC-713) and anti-EGFR (#SC-03) antibodies were from Santa Cruz; anti-GSK3α/β (#44610), anti-Y641P-Stat6 (#611566), anti-S1981P-ATM (#200-301), anti-T2609 DNA-PKcs (#GTX24194) and anti- 1 MCT-11-0474 BKM120: a potent and specific pan-PI3K inhibitor Y1316P-IGF-1R were from Bio-Source International, Becton-Dickinson, Rockland, GenTex and internal production, respectively. The 4G10 antibody was from Millipore (#05-321MG). -

Kinase Profiling Book
Custom and Pre-Selected Kinase Prof iling to f it your Budget and Needs! As of July 1, 2021 19.8653 mm 128 196 12 Tyrosine Serine/Threonine Lipid Kinases Kinases Kinases Carna Biosciences, Inc. 2007 Carna Biosciences, Inc. Profiling Assays available from Carna Biosciences, Inc. As of July 1, 2021 Page Kinase Name Assay Platform Page Kinase Name Assay Platform 4 ABL(ABL1) MSA 21 EGFR[T790M/C797S/L858R] MSA 4 ABL(ABL1)[E255K] MSA 21 EGFR[T790M/L858R] MSA 4 ABL(ABL1)[T315I] MSA 21 EPHA1 MSA 4 ACK(TNK2) MSA 21 EPHA2 MSA 4 AKT1 MSA 21 EPHA3 MSA 5 AKT2 MSA 22 EPHA4 MSA 5 AKT3 MSA 22 EPHA5 MSA 5 ALK MSA 22 EPHA6 MSA 5 ALK[C1156Y] MSA 22 EPHA7 MSA 5 ALK[F1174L] MSA 22 EPHA8 MSA 6 ALK[G1202R] MSA 23 EPHB1 MSA 6 ALK[G1269A] MSA 23 EPHB2 MSA 6 ALK[L1196M] MSA 23 EPHB3 MSA 6 ALK[R1275Q] MSA 23 EPHB4 MSA 6 ALK[T1151_L1152insT] MSA 23 Erk1(MAPK3) MSA 7 EML4-ALK MSA 24 Erk2(MAPK1) MSA 7 NPM1-ALK MSA 24 Erk5(MAPK7) MSA 7 AMPKα1/β1/γ1(PRKAA1/B1/G1) MSA 24 FAK(PTK2) MSA 7 AMPKα2/β1/γ1(PRKAA2/B1/G1) MSA 24 FER MSA 7 ARG(ABL2) MSA 24 FES MSA 8 AurA(AURKA) MSA 25 FGFR1 MSA 8 AurA(AURKA)/TPX2 MSA 25 FGFR1[V561M] MSA 8 AurB(AURKB)/INCENP MSA 25 FGFR2 MSA 8 AurC(AURKC) MSA 25 FGFR2[V564I] MSA 8 AXL MSA 25 FGFR3 MSA 9 BLK MSA 26 FGFR3[K650E] MSA 9 BMX MSA 26 FGFR3[K650M] MSA 9 BRK(PTK6) MSA 26 FGFR3[V555L] MSA 9 BRSK1 MSA 26 FGFR3[V555M] MSA 9 BRSK2 MSA 26 FGFR4 MSA 10 BTK MSA 27 FGFR4[N535K] MSA 10 BTK[C481S] MSA 27 FGFR4[V550E] MSA 10 BUB1/BUB3 MSA 27 FGFR4[V550L] MSA 10 CaMK1α(CAMK1) MSA 27 FGR MSA 10 CaMK1δ(CAMK1D) MSA 27 FLT1 MSA 11 CaMK2α(CAMK2A) MSA 28 -

Identification of a Novel Deletion Mutation in DPY19L2 from An
Li et al. Molecular Cytogenetics (2020) 13:24 https://doi.org/10.1186/s13039-020-00495-1 CASE REPORT Open Access Identification of a novel deletion mutation in DPY19L2 from an infertile patient with globozoospermia: a case report You-zhu Li1†, Rong-feng Wu1†, Xing-shen Zhu2†, Wen-sheng Liu2, Yuan-yuan Ye1, Zhong-xian Lu2* and Na Li3* Abstract Background: Male infertility is an increasing medical concern worldwide. In most cases, genetic factors are considered as the main cause of the disease. Globozoospermia (MIM102530) (also known as round-headed sperm) is a rare and severe malformed spermatospermia caused by acrosome deficiency or severe malformation. A subset of genetic mutations, such as DNAH6, SPATA16, DPY19L2, PICK1, and CCIN related to globozoospermia, have been reported in the past few years. The DPY19L2 mutation is commonly found in patients with globozoospermia. Herein, a 180-kbp homozygote deletion at 12q14.2 (g.63950001–64130000) was identified by copy number variation sequencing (CNVseq) in a patient with a globozoospermia, including the complete deletion of DPY19L2. Case presentation: A 27-year-old patient at the First Affiliated Hospital of Xiamen University was diagnosed with infertility because, despite normal sexual activity for 4 years, his wife did not conceive. The patient was in good health with no obvious discomfort, no history of adverse chemical exposure, and no vices, such as smoking and drinking. The physical examination revealed normal genital development. However, semen tests showed a normal sperm count of 0% and the morphology was the round head. Sperm cytology showed that acrosomal enzyme was lower than normal. -

Phosphorylation of Threonine 3 on Histone H3 by Haspin Kinase Is Required for Meiosis I in Mouse Oocytes
ß 2014. Published by The Company of Biologists Ltd | Journal of Cell Science (2014) 127, 5066–5078 doi:10.1242/jcs.158840 RESEARCH ARTICLE Phosphorylation of threonine 3 on histone H3 by haspin kinase is required for meiosis I in mouse oocytes Alexandra L. Nguyen, Amanda S. Gentilello, Ahmed Z. Balboula*, Vibha Shrivastava, Jacob Ohring and Karen Schindler` ABSTRACT a lesser extent. However, it is not known whether haspin is required for meiosis in oocytes. To date, the only known haspin substrates Meiosis I (MI), the division that generates haploids, is prone to are threonine 3 of histone H3 (H3T3), serine 137 of macroH2A and errors that lead to aneuploidy in females. Haspin is a kinase that threonine 57 of CENP-T (Maiolica et al., 2014). Knockdown or phosphorylates histone H3 on threonine 3, thereby recruiting Aurora inhibition of haspin in mitotically dividing tissue culture cell kinase B (AURKB) and the chromosomal passenger complex lines reveal that phosphorylation of H3T3 is essential for the (CPC) to kinetochores to regulate mitosis. Haspin and AURKC, an alignment of chromosomes at the metaphase plate (Dai and AURKB homolog, are enriched in germ cells, yet their significance Higgins, 2005; Dai et al., 2005), regulation of chromosome in regulating MI is not fully understood. Using inhibitors and cohesion (Dai et al., 2009) and establishing a bipolar spindle (Dai overexpression approaches, we show a role for haspin during MI et al., 2009). In mitotic metaphase, phosphorylation of H3T3 is in mouse oocytes. Haspin-perturbed oocytes display abnormalities restricted to kinetochores, and this mark signals recruitment of the in chromosome morphology and alignment, improper kinetochore– chromosomal passenger complex (CPC) (Dai et al., 2005; Wang microtubule attachments at metaphase I and aneuploidy at et al., 2010; Yamagishi et al., 2010). -

Maternally Recruited Aurora C Kinase Is More Stable Than Aurora B to Support Mouse Oocyte Maturation and Early Development
Maternally recruited Aurora C kinase is more stable PNAS PLUS than Aurora B to support mouse oocyte maturation and early development Karen Schindler1,2, Olga Davydenko2, Brianna Fram, Michael A. Lampson3, and Richard M. Schultz3 Department of Biology, University of Pennsylvania, Philadelphia, PA 19104 Edited* by John J. Eppig, The Jackson Laboratory, Bar Harbor, ME, and approved June 14, 2012 (received for review December 14, 2011) Aurora kinases are highly conserved, essential regulators of cell including abnormally condensed chromatin and abnormally division. Two Aurora kinase isoforms, A and B (AURKA and shaped acrosomes, but females were not examined (11). Muta- AURKB), are expressed ubiquitously in mammals, whereas a third tions in human AURKC cause meiotic arrest and formation of isoform, Aurora C (AURKC), is largely restricted to germ cells. tetraploid sperm (12), suggesting an essential role in cytokinesis Because AURKC is very similar to AURKB, based on sequence and in male meiosis. Experiments in mouse oocytes using a chemical functional analyses, why germ cells express AURKC is unclear. We inhibitor of AURKB (ZM447439) do not address the function of −/− report that Aurkc females are subfertile, and that AURKB func- AURKB because AURKC is also inhibited (13–17). Strategies tion declines as development progresses based on increasing se- using dominant-negative versions of AURKC are also difficult to verity of cytokinesis failure and arrested embryonic development. interpret, because the mutant may also compete with AURKB Furthermore, we find that neither Aurkb nor Aurkc is expressed (18). Overexpression studies have similar limitations because after the one-cell stage, and that AURKC is more stable during both kinases interact with inner centromere protein (INCENP) maturation than AURKB using fluorescently tagged reporter and these studies did not report expression levels of AURKB proteins. -

Globozoospermia Syndrome: an Update
Received: 16 July 2019 | Revised: 17 September 2019 | Accepted: 21 September 2019 DOI: 10.1111/and.13459 INVITED REVIEW Globozoospermia syndrome: An update Farzaneh Fesahat1 | Ralf Henkel2,3 | Ashok Agarwal3 1Reproductive Immunology Research Center, Shahid Sadoughi University of Abstract Medical Sciences, Yazd, Iran Among the factors involved in male infertility, there is a rare morphology disorder 2 Department of Medical called "globozoospermia" that is classified into total globozoospermia and partial Bioscience, University of the Western Cape, Bellville, South Africa globozoospermia (type I and type II, respectively). This syndrome is primarily char‐ 3American Center for Reproductive acterised by the presence of round‐headed spermatozoa with cytoskeleton defects Medicine, Cleveland Clinic, Cleveland, OH, USA around the nucleus and no acrosome. Current data support the negative correlation between globozoospermia and conventional intracytoplasmic sperm injection (ICSI) Correspondence Ashok Agarwal, American Center for outcomes, revealing the need for the management of patients undergoing assisted Reproductive Medicine, Cleveland Clinic, reproduction technology (ART) through more effective treatment techniques. This Cleveland, OH, USA. Email: [email protected] review highlights the most important characteristics of globozoospermia such as sperm parameters, DNA/chromatin integrity and sperm DNA fragmentation (SDF), as well as genetic features based on the latest knowledge. Additionally, we looked into current progress on fertilisation potential and possible treatment strategies for patients presenting with globozoospermia. KEYWORDS DNA fragmentation, globozoospermia, human, intracytoplasmic sperm injection, morphology, spermatozoa 1 | INTRODUCTION causes primary male infertility (Singh, 1992). Contrary, men with type II globozoospermia have both normal and round‐headed sperm Among the factors involved in male infertility, there is a rare mor‐ cells with large CDs, which impair motility. -

Genetic Variations in AURORA Cell Cycle Kinases Are Associated With
www.nature.com/scientificreports OPEN Genetic variations in AURORA cell cycle kinases are associated with glioblastoma multiforme Aner Mesic1, Marija Rogar2, Petra Hudler2*, Nurija Bilalovic3, Izet Eminovic1 & Radovan Komel2 Glioblastoma multiforme (GBM) is the most frequent type of primary astrocytomas. We examined the association between single nucleotide polymorphisms (SNPs) in Aurora kinase A (AURKA), Aurora kinase B (AURKB), Aurora kinase C (AURKC) and Polo-like kinase 1 (PLK1) mitotic checkpoint genes and GBM risk by qPCR genotyping. In silico analysis was performed to evaluate efects of polymorphic biological sequences on protein binding motifs. Chi-square and Fisher statistics revealed a signifcant diference in genotypes frequencies between GBM patients and controls for AURKB rs2289590 variant (p = 0.038). Association with decreased GBM risk was demonstrated for AURKB rs2289590 AC genotype (OR = 0.54; 95% CI = 0.33–0.88; p = 0.015). Furthermore, AURKC rs11084490 CG genotype was associated with lower GBM risk (OR = 0.57; 95% CI = 0.34–0.95; p = 0.031). Bioinformatic analysis of rs2289590 polymorphic region identifed additional binding site for the Yin-Yang 1 (YY1) transcription factor in the presence of C allele. Our results indicated that rs2289590 in AURKB and rs11084490 in AURKC were associated with a reduced GBM risk. The present study was performed on a less numerous but ethnically homogeneous population. Hence, future investigations in larger and multiethnic groups are needed to strengthen these results. Glioblastoma multiforme (GBM) represents the most common and lethal form of primary brain tumor with an annual incidence of 5.26 per 100,000 people 1,2 and it stands for more than 60% of all brain tumors in adults3. -

Promising Therapy in Lung Cancer: Spotlight on Aurora Kinases
cancers Review Promising Therapy in Lung Cancer: Spotlight on Aurora Kinases Domenico Galetta 1,* and Lourdes Cortes-Dericks 2 1 Division of Thoracic Surgery, European Institute of Oncology, IRCCS, 20141 Milan, Italy 2 Department of Biology, University of Hamburg, 20146 Hamburg, Germany; [email protected] * Correspondence: [email protected] Received: 4 September 2020; Accepted: 12 November 2020; Published: 14 November 2020 Simple Summary: Lung cancer has remained one of the major causes of death worldwide. Thus, a more effective treatment approach is essential, such as the inhibition of specific cancer-promoting molecules. Aurora kinases regulate the process of mitosis—a process of cell division that is necessary for normal cell proliferation. Dysfunction of these kinases can contribute to cancer formation. In this review, we present studies indicating the implication of Aurora kinases in tumor formation, drug resistance, and disease prognosis. The effectivity of using Aurora kinase inhibitors in the pre-clinical and clinical investigations has proven their therapeutic potential in the setting of lung cancer. This work may provide further information to broaden the development of anticancer drugs and, thus, improve the conventional lung cancer management. Abstract: Despite tremendous efforts to improve the treatment of lung cancer, prognosis still remains poor; hence, the search for efficacious therapeutic option remains a prime concern in lung cancer research. Cell cycle regulation including mitosis has emerged as an important target for cancer management. Novel pharmacological agents blocking the activities of regulatory molecules that control the functional aspects of mitosis such as Aurora kinases are now being investigated. The Aurora kinases, Aurora-A (AURKA), and Aurora B (AURKB) are overexpressed in many tumor entities such as lung cancer that correlate with poor survival, whereby their inhibition, in most cases, enhances the efficacy of chemo-and radiotherapies, indicating their implication in cancer therapy. -
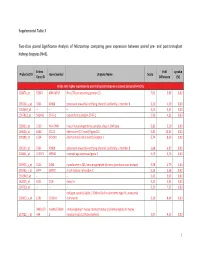
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at