Geochemical Influences on Nonenzymatic Oligomerization Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Targets and Biological Functions of Camp Signaling in Arabidopsis
biomolecules Article Molecular Targets and Biological Functions of cAMP Signaling in Arabidopsis Ruqiang Xu 1,2,*, Yanhui Guo 1, Song Peng 1, Jinrui Liu 1, Panyu Li 1, Wenjing Jia 1 and Junheng Zhao 1 1 School of Agricultural Sciences, Zhengzhou University, Zhengzhou 450001, China; [email protected] (Y.G.); [email protected] (S.P.); [email protected] (J.L.); [email protected] (P.L.); [email protected] (W.J.); [email protected] (J.Z.) 2 Zhengzhou Research Base, State Key Laboratory of Cotton Biology, Zhengzhou University, Zhengzhou 450001, China * Correspondence: [email protected]; Tel.: +86-0371-6778-5095 Abstract: Cyclic AMP (cAMP) is a pivotal signaling molecule existing in almost all living organisms. However, the mechanism of cAMP signaling in plants remains very poorly understood. Here, we employ the engineered activity of soluble adenylate cyclase to induce cellular cAMP elevation in Arabidopsis thaliana plants and identify 427 cAMP-responsive genes (CRGs) through RNA-seq analysis. Induction of cellular cAMP elevation inhibits seed germination, disturbs phytohormone contents, promotes leaf senescence, impairs ethylene response, and compromises salt stress tolerance and pathogen resistance. A set of 62 transcription factors are among the CRGs, supporting a prominent role of cAMP in transcriptional regulation. The CRGs are significantly overrepresented in the pathways of plant hormone signal transduction, MAPK signaling, and diterpenoid biosynthesis, but + they are also implicated in lipid, sugar, K , nitrate signaling, and beyond. Our results provide a basic framework of cAMP signaling for the community to explore. The regulatory roles of cAMP signaling Citation: Xu, R.; Guo, Y.; Peng, S.; in plant plasticity are discussed. -

Mechanisms Whereby Extracellular Adenosine 3',5'- Monophosphate Inhibits Phosphate Transport in Cultured Opossum Kidney Cells and in Rat Kidney
Mechanisms whereby extracellular adenosine 3',5'- monophosphate inhibits phosphate transport in cultured opossum kidney cells and in rat kidney. Physiological implication. G Friedlander, … , C Coureau, C Amiel J Clin Invest. 1992;90(3):848-858. https://doi.org/10.1172/JCI115960. Research Article The mechanism of phosphaturia induced by cAMP infusion and the physiological role of extracellular cAMP in modulation of renal phosphate transport were examined. In cultured opossum kidney cells, extracellular cAMP (10-1,000 microM) inhibited Na-dependent phosphate uptake in a time- and concentration-dependent manner. The effect of cAMP was reproduced by ATP, AMP, and adenosine, and was blunted by phosphodiesterase inhibitors or by dipyridamole which inhibits adenosine uptake. [3H]cAMP was degraded extracellularly into AMP and adenosine, and radioactivity accumulated in the cells as labeled adenosine and, subsequently, as adenine nucleotides including cAMP. Radioactivity accumulation was decreased by dipyridamole and by inhibitors of phosphodiesterases and ecto-5'-nucleotidase, assessing the existence of stepwise hydrolysis of extracellular cAMP and intracellular processing of taken up adenosine. In vivo, dipyridamole abolished the phosphaturia induced by exogenous cAMP infusion in acutely parathyroidectomized (APTX) rats, decreased phosphate excretion in intact rats, and blunted phosphaturia induced by PTH infusion in APTX rats. These results indicate that luminal degradation of cAMP into adenosine, followed by cellular uptake of the nucleoside by tubular cells, is a key event which accounts for the phosphaturic effect of exogenous cAMP and for the part of the phosphaturic effect of PTH which is mediated by cAMP added to the tubular lumen under the influence of the hormone. -

Alternative Biochemistries for Alien Life: Basic Concepts and Requirements for the Design of a Robust Biocontainment System in Genetic Isolation
G C A T T A C G G C A T genes Review Alternative Biochemistries for Alien Life: Basic Concepts and Requirements for the Design of a Robust Biocontainment System in Genetic Isolation Christian Diwo 1 and Nediljko Budisa 1,2,* 1 Institut für Chemie, Technische Universität Berlin Müller-Breslau-Straße 10, 10623 Berlin, Germany; [email protected] 2 Department of Chemistry, University of Manitoba, 144 Dysart Rd, 360 Parker Building, Winnipeg, MB R3T 2N2, Canada * Correspondence: [email protected] or [email protected]; Tel.: +49-30-314-28821 or +1-204-474-9178 Received: 27 November 2018; Accepted: 21 December 2018; Published: 28 December 2018 Abstract: The universal genetic code, which is the foundation of cellular organization for almost all organisms, has fostered the exchange of genetic information from very different paths of evolution. The result of this communication network of potentially beneficial traits can be observed as modern biodiversity. Today, the genetic modification techniques of synthetic biology allow for the design of specialized organisms and their employment as tools, creating an artificial biodiversity based on the same universal genetic code. As there is no natural barrier towards the proliferation of genetic information which confers an advantage for a certain species, the naturally evolved genetic pool could be irreversibly altered if modified genetic information is exchanged. We argue that an alien genetic code which is incompatible with nature is likely to assure the inhibition of all mechanisms of genetic information transfer in an open environment. The two conceivable routes to synthetic life are either de novo cellular design or the successive alienation of a complex biological organism through laboratory evolution. -

Monophosphate- Binding Protein Complex with Subsequent Poly(A) RNA Synthesis in Embryonic Chick Cartilage
Specific nuclear binding of adenosine 3',5'-monophosphate- binding protein complex with subsequent poly(A) RNA synthesis in embryonic chick cartilage. W M Burch Jr, H E Lebovitz J Clin Invest. 1980;66(3):532-542. https://doi.org/10.1172/JCI109885. Research Article We used embryonic chick pelvic cartilage as a model to study the mechanism by which cyclic AMP increases RNA synthesis. Isolated nuclei were incubated with [32P]-8-azidoadenosine 3,5'-monophosphate ([32P]N3cAMP) with no resultant specific nuclear binding. However, in the presence of cytosol proteins, nuclear binding of [32P]N3cAMP was demonstrable that was specific, time dependent, and dependent on a heat-labile cytosol factor. The possible biological significance of the nuclear binding of the cyclic AMP-protein complex was identified by incubating isolating nuclei with either cyclic AMP or cytosol cyclic AMP-binding proteins prepared by batch elution DEAE cellulose chromatography (DEAE peak cytosol protein), or both, in the presence of cold nucleotides and [3H]uridine 5'-triphosphate. Poly(A) RNA production occurred only in nuclei incubated with cyclic AMP and the DEAE peak cytosol protein preparation. Actinomycin D inhibited the incorporation of [3H]uridine 5'-monophosphate into poly(A) RNA. The newly synthesized poly(A) RNA had a sedimentation constant of 23S. Characterization of the cytosol cyclic AMP binding proteins using [32P]N3-cAMP with photoaffinity labeling three major cAMP-binding complexes (41,000, 51,000, and 55,000 daltons). The 51,000 and 55,000 dalton cyclic AMP binding proteins were further purified by DNA-cellulose chromatography. In the presence of cyclic AMP they stimulated poly(A) RNA synthesis in isolated nuclei. -

Challenges on Cyclic Nucleotide Phosphodiesterases Imaging with Positron Emission Tomography: Novel Radioligands and (Pre-)Clinical Insights Since 2016
International Journal of Molecular Sciences Review Challenges on Cyclic Nucleotide Phosphodiesterases Imaging with Positron Emission Tomography: Novel Radioligands and (Pre-)Clinical Insights since 2016 Susann Schröder 1,2,* , Matthias Scheunemann 2, Barbara Wenzel 2 and Peter Brust 2 1 Department of Research and Development, ROTOP Pharmaka Ltd., 01328 Dresden, Germany 2 Department of Neuroradiopharmaceuticals, Institute of Radiopharmaceutical Cancer Research, Research Site Leipzig, Helmholtz-Zentrum Dresden-Rossendorf (HZDR), 04318 Leipzig, Germany; [email protected] (M.S.); [email protected] (B.W.); [email protected] (P.B.) * Correspondence: [email protected]; Tel.: +49-341-234-179-4631 Abstract: Cyclic nucleotide phosphodiesterases (PDEs) represent one of the key targets in the research field of intracellular signaling related to the second messenger molecules cyclic adenosine monophosphate (cAMP) and/or cyclic guanosine monophosphate (cGMP). Hence, non-invasive imaging of this enzyme class by positron emission tomography (PET) using appropriate isoform- selective PDE radioligands is gaining importance. This methodology enables the in vivo diagnosis and staging of numerous diseases associated with altered PDE density or activity in the periphery and the central nervous system as well as the translational evaluation of novel PDE inhibitors as therapeutics. In this follow-up review, we summarize the efforts in the development of novel PDE radioligands and highlight (pre-)clinical insights from PET studies using already known PDE Citation: Schröder, S.; Scheunemann, radioligands since 2016. M.; Wenzel, B.; Brust, P. Challenges on Cyclic Nucleotide Keywords: positron emission tomography; cyclic nucleotide phosphodiesterases; PDE inhibitors; Phosphodiesterases Imaging with PDE radioligands; radiochemistry; imaging; recent (pre-)clinical insights Positron Emission Tomography: Novel Radioligands and (Pre-)Clinical Insights since 2016. -

Oxidative Stress and the Guanosine Nucleotide Triphosphate Pool: Implications for a Biomarker and Mechanism of Impaired Cell Function
University of Montana ScholarWorks at University of Montana Graduate Student Theses, Dissertations, & Professional Papers Graduate School 2008 OXIDATIVE STRESS AND THE GUANOSINE NUCLEOTIDE TRIPHOSPHATE POOL: IMPLICATIONS FOR A BIOMARKER AND MECHANISM OF IMPAIRED CELL FUNCTION Celeste Maree Bolin The University of Montana Follow this and additional works at: https://scholarworks.umt.edu/etd Let us know how access to this document benefits ou.y Recommended Citation Bolin, Celeste Maree, "OXIDATIVE STRESS AND THE GUANOSINE NUCLEOTIDE TRIPHOSPHATE POOL: IMPLICATIONS FOR A BIOMARKER AND MECHANISM OF IMPAIRED CELL FUNCTION" (2008). Graduate Student Theses, Dissertations, & Professional Papers. 728. https://scholarworks.umt.edu/etd/728 This Dissertation is brought to you for free and open access by the Graduate School at ScholarWorks at University of Montana. It has been accepted for inclusion in Graduate Student Theses, Dissertations, & Professional Papers by an authorized administrator of ScholarWorks at University of Montana. For more information, please contact [email protected]. OXIDATIVE STRESS AND THE GUANOSINE NUCLEOTIDE TRIPHOSPHATE POOL: IMPLICATIONS FOR A BIOMARKER AND MECHANISM OF IMPAIRED CELL FUNCTION By Celeste Maree Bolin B.A. Chemistry, Whitman College, Walla Walla, WA 2001 Dissertation presented in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Toxicology The University of Montana Missoula, Montana Spring 2008 Approved by: Dr. David A. Strobel, Dean Graduate School Dr. Fernando Cardozo-Pelaez, -

RSC Article Template
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics This journal is © The Owner Societies 2013 Supplimentary Information Figure Legends: S1: Raman spectra of RNA building blocks and related compounds. All spectra are the average of 3 Raman spectra and have been baseline corrected, buffer subtracted and scaled to an effective 5 concentration of 50 mM. (a) Adenine related molecules: adenine (black), adenosine (red) and adenosine monophosphate (blue). The measured concentrations were 47.81 mM, 46.40 mM and 56.75 mM, respectively. (b) Cytosine related molecules: cytosine (black), cytidine (red) and cytidine monophosphate (blue). The measured concentrations were 47.70 mM, 56.74 mM and 55.38 mM, respectively. (c) Guanine related molecules: guanine (black), guanosine (red) and guanosine 10 monophosphate (blue). The measured concentrations were 46.19 mM, 63.90 mM and 46.67 mM, respectively. (d) Uracil related molecules: uracil (black), uridine (red) and uridine monophosphate (blue). The measured concentrations were 48.09 mM, 48.32 mM and 67.64 mM, respectively. (e) Ribose and phosphate molecules: ribose (black), ribose phosphate (red) and sodium phosphate (blue). The measured concentrations were 287.08 mM, 434.58 mM and 47.75 mM, respectively. 15 S2: Second derivative spectra of the measured RNA Raman spectra; SL-HIV1 (black), SL-APTR (red) and SL-SARS (blue). The positions of peaks discussed in the text are noted. Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics This journal is © The -
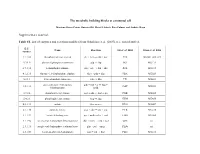
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

Guanosine-Based Nucleotides, the Sons of a Lesser God in the Purinergic Signal Scenario of Excitable Tissues
International Journal of Molecular Sciences Review Guanosine-Based Nucleotides, the Sons of a Lesser God in the Purinergic Signal Scenario of Excitable Tissues 1,2, 2,3, 1,2 1,2, Rosa Mancinelli y, Giorgio Fanò-Illic y, Tiziana Pietrangelo and Stefania Fulle * 1 Department of Neuroscience Imaging and Clinical Sciences, University “G. d’Annunzio” of Chieti-Pescara, 66100 Chieti, Italy; [email protected] (R.M.); [email protected] (T.P.) 2 Interuniversity Institute of Miology (IIM), 66100 Chieti, Italy; [email protected] 3 Libera Università di Alcatraz, Santa Cristina di Gubbio, 06024 Gubbio, Italy * Correspondence: [email protected] Both authors contributed equally to this work. y Received: 30 January 2020; Accepted: 25 February 2020; Published: 26 February 2020 Abstract: Purines are nitrogen compounds consisting mainly of a nitrogen base of adenine (ABP) or guanine (GBP) and their derivatives: nucleosides (nitrogen bases plus ribose) and nucleotides (nitrogen bases plus ribose and phosphate). These compounds are very common in nature, especially in a phosphorylated form. There is increasing evidence that purines are involved in the development of different organs such as the heart, skeletal muscle and brain. When brain development is complete, some purinergic mechanisms may be silenced, but may be reactivated in the adult brain/muscle, suggesting a role for purines in regeneration and self-repair. Thus, it is possible that guanosine-50-triphosphate (GTP) also acts as regulator during the adult phase. However, regarding GBP, no specific receptor has been cloned for GTP or its metabolites, although specific binding sites with distinct GTP affinity characteristics have been found in both muscle and neural cell lines. -

Developmental Disorder Associated with Increased Cellular Nucleotidase Activity (Purine-Pyrimidine Metabolism͞uridine͞brain Diseases)
Proc. Natl. Acad. Sci. USA Vol. 94, pp. 11601–11606, October 1997 Medical Sciences Developmental disorder associated with increased cellular nucleotidase activity (purine-pyrimidine metabolismyuridineybrain diseases) THEODORE PAGE*†,ALICE YU‡,JOHN FONTANESI‡, AND WILLIAM L. NYHAN‡ Departments of *Neurosciences and ‡Pediatrics, University of California at San Diego, La Jolla, CA 92093 Communicated by J. Edwin Seegmiller, University of California at San Diego, La Jolla, CA, August 7, 1997 (received for review June 26, 1997) ABSTRACT Four unrelated patients are described with a represent defects of purine metabolism, although no specific syndrome that included developmental delay, seizures, ataxia, enzyme abnormality has been identified in these cases (6). In recurrent infections, severe language deficit, and an unusual none of these disorders has it been possible to delineate the behavioral phenotype characterized by hyperactivity, short mechanism through which the enzyme deficiency produces the attention span, and poor social interaction. These manifesta- neurological or behavioral abnormalities. Therapeutic strate- tions appeared within the first few years of life. Each patient gies designed to treat the behavioral and neurological abnor- displayed abnormalities on EEG. No unusual metabolites were malities of these disorders by replacing the supposed deficient found in plasma or urine, and metabolic testing was normal metabolites have not been successful in any case. except for persistent hypouricosuria. Investigation of purine This report describes four unrelated patients in whom and pyrimidine metabolism in cultured fibroblasts derived developmental delay, seizures, ataxia, recurrent infections, from these patients showed normal incorporation of purine speech deficit, and an unusual behavioral phenotype were bases into nucleotides but decreased incorporation of uridine. -

Nucleobases Thin Films Deposited on Nanostructured Transparent Conductive Electrodes for Optoelectronic Applications
www.nature.com/scientificreports OPEN Nucleobases thin flms deposited on nanostructured transparent conductive electrodes for optoelectronic applications C. Breazu1*, M. Socol1, N. Preda1, O. Rasoga1, A. Costas1, G. Socol2, G. Petre1,3 & A. Stanculescu1* Environmentally-friendly bio-organic materials have become the centre of recent developments in organic electronics, while a suitable interfacial modifcation is a prerequisite for future applications. In the context of researches on low cost and biodegradable resource for optoelectronics applications, the infuence of a 2D nanostructured transparent conductive electrode on the morphological, structural, optical and electrical properties of nucleobases (adenine, guanine, cytosine, thymine and uracil) thin flms obtained by thermal evaporation was analysed. The 2D array of nanostructures has been developed in a polymeric layer on glass substrate using a high throughput and low cost technique, UV-Nanoimprint Lithography. The indium tin oxide electrode was grown on both nanostructured and fat substrate and the properties of the heterostructures built on these two types of electrodes were analysed by comparison. We report that the organic-electrode interface modifcation by nano- patterning afects both the optical (transmission and emission) properties by multiple refections on the walls of nanostructures and the electrical properties by the efect on the organic/electrode contact area and charge carrier pathway through electrodes. These results encourage the potential application of the nucleobases thin flms deposited on nanostructured conductive electrode in green optoelectronic devices. Te use of natural or nature-inspired materials in organic electronics is a dynamic emerging research feld which aims to replace the synthesized materials with natural (bio) ones in organic electronics1–3. -

Nucleobase-Containing Compounds Evoke Behavioural, Olfactory, and Transcriptional Responses in Model Fishes
Correction-Nucleobase-containing compounds evoke behavioural, olfactory, and transcriptional responses in model fishes Shamchuk, A. L., Blunt, B. J., Lyons, D. D., Wang, M. Q., Gasheva, A., Lewis, C. R., Tomlin, K., Starr Hazard, E., Hardiman, G., & Tierney, K. B. (2018). Correction-Nucleobase-containing compounds evoke behavioural, olfactory, and transcriptional responses in model fishes. FACETS, 3(1). https://doi.org/10.1139/facets-2017-0101 Published in: FACETS Document Version: Publisher's PDF, also known as Version of record Queen's University Belfast - Research Portal: Link to publication record in Queen's University Belfast Research Portal Publisher rights Copyright 2018 the authors. This is an open access article published under a Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution and reproduction in any medium, provided the author and source are cited. 4.0), which permits unrestricted use, General rights Copyright for the publications made accessible via the Queen's University Belfast Research Portal is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The Research Portal is Queen's institutional repository that provides access to Queen's research output. Every effort has been made to ensure that content in the Research Portal does not infringe any person's rights, or applicable UK laws. If you discover content in the Research Portal that you believe breaches copyright or violates any law, please contact [email protected].