Aryl Hydrocarbon Receptor-Mediated Regulation of Gene Expression During Cardiomyocyte Differentiation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

ATF4 Promotes Angiogenesis and Neuronal Cell Death and Confers Ferroptosis in a Xct-Dependent Manner
OPEN Oncogene (2017) 36, 5593–5608 www.nature.com/onc ORIGINAL ARTICLE ATF4 promotes angiogenesis and neuronal cell death and confers ferroptosis in a xCT-dependent manner D Chen1,2,ZFan1,3, M Rauh4, M Buchfelder5, IY Eyupoglu1,5 and N Savaskan1,5,6 Activating transcription factor 4 (ATF4) is a critical mediator of metabolic and oxidative homeostasis and cell survival. ATF4 is elevated in response to diverse microenvironmental stresses, including starvation, ER stress damages and exposure to toxic factors. Here we show that ATF4 expression fosters the malignancy of primary brain tumors (WHO grade III and IV gliomas) and increases proliferation and tumor angiogenesis. Hence, ATF4 expression promotes cell migration and anchorage-independent cell growth, whereas siRNA-mediated knockdown of ATF4 attenuates these features of malignancy in human gliomas. Further experiments revealed that ATF4-dependent tumor promoting effects are mediated by transcriptional targeting the glutamate antiporter xCT/SCL7A11 (also known as system Xc-). Thus, xCT is elevated as a consequence of ATF4 activation. We further found evidence that ATF4-induced proliferation can be attenuated by pharmacological or genetic xCT inhibition and ferroptosis inducers such as sorafenib, erastin and GPx4 inhibitor RSL3. Further, fostered xCT expression promotes cell survival and growth in ATF4 knockdown cells. Moreover, increased xCT levels ameliorate sorafenib and erastin-induced ferroptosis. Conversely, ATF4 knockdown renders cells susceptible for erastin, sorafenib and RSL3-induced ferroptosis. We further identified that ATF4 promotes tumor-mediated neuronal cell death which can be alleviated by xCT inhibition. Moreover, elevated ATF4 expression in gliomas promotes tumor angiogenesis. Noteworthy, ATF4-induced angiogenesis could be diminished by ferroptosis inducers erastin and by GPx4 inhibitor RSL3. -

Aberrant Methylation Underlies Insulin Gene Expression in Human Insulinoma
ARTICLE https://doi.org/10.1038/s41467-020-18839-1 OPEN Aberrant methylation underlies insulin gene expression in human insulinoma Esra Karakose1,6, Huan Wang 2,6, William Inabnet1, Rajesh V. Thakker 3, Steven Libutti4, Gustavo Fernandez-Ranvier 1, Hyunsuk Suh1, Mark Stevenson 3, Yayoi Kinoshita1, Michael Donovan1, Yevgeniy Antipin1,2, Yan Li5, Xiaoxiao Liu 5, Fulai Jin 5, Peng Wang 1, Andrew Uzilov 1,2, ✉ Carmen Argmann 1, Eric E. Schadt 1,2, Andrew F. Stewart 1,7 , Donald K. Scott 1,7 & Luca Lambertini 1,6 1234567890():,; Human insulinomas are rare, benign, slowly proliferating, insulin-producing beta cell tumors that provide a molecular “recipe” or “roadmap” for pathways that control human beta cell regeneration. An earlier study revealed abnormal methylation in the imprinted p15.5-p15.4 region of chromosome 11, known to be abnormally methylated in another disorder of expanded beta cell mass and function: the focal variant of congenital hyperinsulinism. Here, we compare deep DNA methylome sequencing on 19 human insulinomas, and five sets of normal beta cells. We find a remarkably consistent, abnormal methylation pattern in insu- linomas. The findings suggest that abnormal insulin (INS) promoter methylation and altered transcription factor expression create alternative drivers of INS expression, replacing cano- nical PDX1-driven beta cell specification with a pathological, looping, distal enhancer-based form of transcriptional regulation. Finally, NFaT transcription factors, rather than the cano- nical PDX1 enhancer complex, are predicted to drive INS transactivation. 1 From the Diabetes Obesity and Metabolism Institute, The Department of Surgery, The Department of Pathology, The Department of Genetics and Genomics Sciences and The Institute for Genomics and Multiscale Biology, The Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. -
Nucleoporin 107, 62 and 153 Mediate Kcnq1ot1 Imprinted Domain Regulation in Extraembryonic Endoderm Stem Cells
ARTICLE DOI: 10.1038/s41467-018-05208-2 OPEN Nucleoporin 107, 62 and 153 mediate Kcnq1ot1 imprinted domain regulation in extraembryonic endoderm stem cells Saqib S. Sachani 1,2,3,4, Lauren S. Landschoot1,2, Liyue Zhang1,2, Carlee R. White1,2, William A. MacDonald3,4, Michael C. Golding 5 & Mellissa R.W. Mann 3,4 1234567890():,; Genomic imprinting is a phenomenon that restricts transcription to predominantly one par- ental allele. How this transcriptional duality is regulated is poorly understood. Here we perform an RNA interference screen for epigenetic factors involved in paternal allelic silen- cing at the Kcnq1ot1 imprinted domain in mouse extraembryonic endoderm stem cells. Multiple factors are identified, including nucleoporin 107 (NUP107). To determine NUP107’s role and specificity in Kcnq1ot1 imprinted domain regulation, we deplete Nup107, as well as Nup62, Nup98/96 and Nup153. Nup107, Nup62 and Nup153, but not Nup98/96 depletion, reduce Kcnq1ot1 noncoding RNA volume, displace the Kcnq1ot1 domain from the nuclear periphery, reactivate a subset of normally silent paternal alleles in the domain, alter histone modifications with concomitant changes in KMT2A, EZH2 and EHMT2 occupancy, as well as reduce cohesin interactions at the Kcnq1ot1 imprinting control region. Our results establish an important role for specific nucleoporins in mediating Kcnq1ot1 imprinted domain regulation. 1 Departments of Obstetrics & Gynaecology, and Biochemistry, Western University, Schulich School of Medicine and Dentistry, London, ON N6A 5W9, Canada. 2 Children’s Health Research Institute, London, ON N6C 2V5, Canada. 3 Departments of Obstetrics, Gynecology and Reproductive Sciences, University of Pittsburgh School of Medicine, Pittsburgh, PA 15213, USA. 4 Magee-Womens Research Institute, Pittsburgh, PA 15213, USA. -
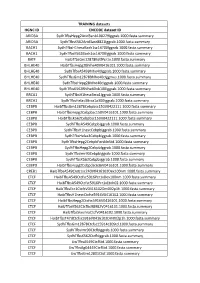
TRAINING Datasets HGNC ID ENCODE Dataset ID ARID3A
TRAINING datasets HGNC ID ENCODE dataset ID ARID3A SydhT+sHepg2Arid3anb100279Iggrab.1000.fasta.summary ARID3A SydhT+sK562Arid3asC8821Iggrab.1000.fasta.summary BACH1 SydhT+sH1hesCBaCh1sC14700Iggrab.1000.fasta.summary BACH1 SydhT+sK562BaCh1sC14700Iggrab.1000.fasta.summary BATF HaibT+sGm12878BaJPCr1x.1000.fasta.summary BHLHE40 HaibT+sHepg2Bhlhe40V0416101.1000.fasta.summary BHLHE40 SydhT+sA549Bhlhe40Iggrab.1000.fasta.summary BHLHE40 SydhT+sGm12878Bhlhe40CIggmus.1000.fasta.summary BHLHE40 SydhT+sHepg2Bhlhe40CIggrab.1000.fasta.summary BHLHE40 SydhT+sK562Bhlhe40nb100Iggrab.1000.fasta.summary BRCA1 SydhT+sH1hesCBrCa1Iggrab.1000.fasta.summary BRCA1 SydhT+sHelas3BrCa1a300Iggrab.1000.fasta.summary CEBPB HaibT+sGm12878CebpbsC150V0422111.1000.fasta.summary CEBPB HaibT+sHepg2CebpbsC150V0416101.1000.fasta.summary CEBPB HaibT+sK562CebpbsC150V0422111.1000.fasta.summary CEBPB SydhT+sA549CebpbIggrab.1000.fasta.summary CEBPB SydhT+sH1hesCCebpbIggrab.1000.fasta.summary CEBPB SydhT+sHelas3CebpbIggrab.1000.fasta.summary CEBPB SydhT+sHepg2CebpbForsklnStd.1000.fasta.summary CEBPB SydhT+sHepg2CebpbIggrab.1000.fasta.summary CEBPB SydhT+sImr90CebpbIggrab.1000.fasta.summary CEBPB SydhT+sK562CebpbIggrab.1000.fasta.summary CEBPD HaibT+sHepg2CebpdsC636V0416101.1000.fasta.summary CREB1 HaibT+sA549Creb1sC240V0416102Dex100nm.1000.fasta.summary CTCF HaibT+sA549CtCfsC5916PCr1xDex100nm.1000.fasta.summary CTCF HaibT+sA549CtCfsC5916PCr1xEtoh02.1000.fasta.summary CTCF HaibT+sECC1CtCfCV0416102Dm002p1h.1000.fasta.summary CTCF HaibT+sH1hesCCtCfsC5916V0416102.1000.fasta.summary -

1 the Neuroprotective Transcription Factor ATF5 Is Decreased And
The neuroprotective transcription factor ATF5 is decreased and sequestered into polyglutamine inclusions in Huntington’s disease Ivó H. Hernández1,2,3, Jesús Torres-Peraza1,2,5, María Santos-Galindo1,2, Eloísa Ramos-Morón4, María R. Fernández-Fernández1,2, María J. Pérez-Álvarez1,2,3, Antonio Miranda-Vizuete4 and José J. Lucas1,2* 1 Centro de Biología Molecular Severo Ochoa (CBM”SO”) CSIC/UAM, Madrid, Spain. 2 Instituto de Salud Carlos III, Networking Research Center on Neurodegenerative Diseases (CIBERNED), Spain. 3 Departamento de Biología, Facultad de Ciencias, Universidad Autónoma de Madrid, Madrid, Spain. 4 Instituto de Biomedicina de Sevilla, Hospital Universitario Virgen del Rocío/CSIC/Universidad de Sevilla, 41013 Sevilla, Spain 5 Present Address: Gerència d’Atenció Primària del Servei de Salut de les Illes Balears (IB-SALUT), Palma, Spain *Corresponding author: José J. Lucas [email protected] http://www.cbm.uam.es/lineas/lucasgroup.htm provided by Digital.CSIC View metadata, citation and similar papers at core.ac.uk CORE brought to you by 1 Abstract Activating transcription factor-5 (ATF5) is a stress-response transcription factor induced upon different cell stressors like fasting, amino-acid limitation, cadmium or arsenite. ATF5 is also induced, and promotes transcription of anti-apoptotic target genes like MCL1, during the unfolded protein response (UPR) triggered by endoplasmic reticulum stress. In the brain, high ATF5 levels are found in gliomas and also in neural progenitor cells, which need to decrease their ATF5 levels for differentiation into mature neurons or glia. This initially led to believe that ATF5 is not expressed in adult neurons. More recently, we reported basal neuronal ATF5 expression in adult mouse brain and its neuroprotective induction during UPR in a mouse model of status epilepticus. -

Roles of the CSE1L-Mediated Nuclear Import Pathway in Epigenetic
Roles of the CSE1L-mediated nuclear import pathway PNAS PLUS in epigenetic silencing Qiang Donga,b,c, Xiang Lia,b,c, Cheng-Zhi Wangb, Shaohua Xuc, Gang Yuanc, Wei Shaoc, Baodong Liud, Yong Zhengb, Hailin Wangd, Xiaoguang Leic,e,f, Zhuqiang Zhangb,1, and Bing Zhua,b,g,1 aGraduate Program, Peking Union Medical College and Chinese Academy of Medical Sciences, 100730 Beijing, China; bNational Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 100101 Beijing, China; cNational Institute of Biological Sciences, 102206 Beijing, China; dThe State Key Laboratory of Environmental Chemistry and Ecotoxicology, Research Center for Eco-Environmental Sciences, Chinese Academy of Sciences, 100085 Beijing, China; eBeijing National Laboratory for Molecular Sciences, Department of Chemical Biology, College of Chemistry and Molecular Engineering, Peking University, Beijing 100871, China; fPeking-Tsinghua Center for Life Sciences, Peking University, 100871 Beijing, China; and gCollege of Life Sciences, University of Chinese Academy of Sciences, 100049 Beijing, China Edited by Arthur D. Riggs, Beckman Research Institute of City of Hope, Duarte, CA, and approved March 21, 2018 (received for review January 17, 2018) Epigenetic silencing can be mediated by various mechanisms, CSE1L, a key player in the nuclear import pathway, as an es- and many regulators remain to be identified. Here, we report a sential factor for maintaining the repression of many methyl- genome-wide siRNA screening to identify regulators essential for ated genes. Mechanistically, CSE1L functions by facilitating maintaining gene repression of a CMV promoter silenced by DNA the nuclear import of certain cargo proteins that are essential methylation. -

Prospective Isolation of NKX2-1–Expressing Human Lung Progenitors Derived from Pluripotent Stem Cells
The Journal of Clinical Investigation RESEARCH ARTICLE Prospective isolation of NKX2-1–expressing human lung progenitors derived from pluripotent stem cells Finn Hawkins,1,2 Philipp Kramer,3 Anjali Jacob,1,2 Ian Driver,4 Dylan C. Thomas,1 Katherine B. McCauley,1,2 Nicholas Skvir,1 Ana M. Crane,3 Anita A. Kurmann,1,5 Anthony N. Hollenberg,5 Sinead Nguyen,1 Brandon G. Wong,6 Ahmad S. Khalil,6,7 Sarah X.L. Huang,3,8 Susan Guttentag,9 Jason R. Rock,4 John M. Shannon,10 Brian R. Davis,3 and Darrell N. Kotton1,2 2 1Center for Regenerative Medicine, and The Pulmonary Center and Department of Medicine, Boston University School of Medicine, Boston, Massachusetts, USA. 3Center for Stem Cell and Regenerative Medicine, Brown Foundation Institute of Molecular Medicine, University of Texas Health Science Center, Houston, Texas, USA. 4Department of Anatomy, UCSF, San Francisco, California, USA. 5Division of Endocrinology, Diabetes and Metabolism, Beth Israel Deaconess Medical Center and Harvard Medical School, Boston, Massachusetts, USA. 6Department of Biomedical Engineering and Biological Design Center, Boston University, Boston, Massachusetts, USA. 7Wyss Institute for Biologically Inspired Engineering, Harvard University, Boston, Massachusetts, USA. 8Columbia Center for Translational Immunology & Columbia Center for Human Development, Columbia University Medical Center, New York, New York, USA. 9Department of Pediatrics, Monroe Carell Jr. Children’s Hospital, Vanderbilt University, Nashville, Tennessee, USA. 10Division of Pulmonary Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA. It has been postulated that during human fetal development, all cells of the lung epithelium derive from embryonic, endodermal, NK2 homeobox 1–expressing (NKX2-1+) precursor cells. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

A Clinicopathological and Molecular Genetic Analysis of Low-Grade Glioma in Adults
A CLINICOPATHOLOGICAL AND MOLECULAR GENETIC ANALYSIS OF LOW-GRADE GLIOMA IN ADULTS Presented by ANUSHREE SINGH MSc A thesis submitted in partial fulfilment of the requirements of the University of Wolverhampton for the degree of Doctor of Philosophy Brain Tumour Research Centre Research Institute in Healthcare Sciences Faculty of Science and Engineering University of Wolverhampton November 2014 i DECLARATION This work or any part thereof has not previously been presented in any form to the University or to any other body whether for the purposes of assessment, publication or for any other purpose (unless otherwise indicated). Save for any express acknowledgments, references and/or bibliographies cited in the work, I confirm that the intellectual content of the work is the result of my own efforts and of no other person. The right of Anushree Singh to be identified as author of this work is asserted in accordance with ss.77 and 78 of the Copyright, Designs and Patents Act 1988. At this date copyright is owned by the author. Signature: Anushree Date: 30th November 2014 ii ABSTRACT The aim of the study was to identify molecular markers that can determine progression of low grade glioma. This was done using various approaches such as IDH1 and IDH2 mutation analysis, MGMT methylation analysis, copy number analysis using array comparative genomic hybridisation and identification of differentially expressed miRNAs using miRNA microarray analysis. IDH1 mutation was present at a frequency of 71% in low grade glioma and was identified as an independent marker for improved OS in a multivariate analysis, which confirms the previous findings in low grade glioma studies. -

Supplemental Materials ZNF281 Enhances Cardiac Reprogramming
Supplemental Materials ZNF281 enhances cardiac reprogramming by modulating cardiac and inflammatory gene expression Huanyu Zhou, Maria Gabriela Morales, Hisayuki Hashimoto, Matthew E. Dickson, Kunhua Song, Wenduo Ye, Min S. Kim, Hanspeter Niederstrasser, Zhaoning Wang, Beibei Chen, Bruce A. Posner, Rhonda Bassel-Duby and Eric N. Olson Supplemental Table 1; related to Figure 1. Supplemental Table 2; related to Figure 1. Supplemental Table 3; related to the “quantitative mRNA measurement” in Materials and Methods section. Supplemental Table 4; related to the “ChIP-seq, gene ontology and pathway analysis” and “RNA-seq” and gene ontology analysis” in Materials and Methods section. Supplemental Figure S1; related to Figure 1. Supplemental Figure S2; related to Figure 2. Supplemental Figure S3; related to Figure 3. Supplemental Figure S4; related to Figure 4. Supplemental Figure S5; related to Figure 6. Supplemental Table S1. Genes included in human retroviral ORF cDNA library. Gene Gene Gene Gene Gene Gene Gene Gene Symbol Symbol Symbol Symbol Symbol Symbol Symbol Symbol AATF BMP8A CEBPE CTNNB1 ESR2 GDF3 HOXA5 IL17D ADIPOQ BRPF1 CEBPG CUX1 ESRRA GDF6 HOXA6 IL17F ADNP BRPF3 CERS1 CX3CL1 ETS1 GIN1 HOXA7 IL18 AEBP1 BUD31 CERS2 CXCL10 ETS2 GLIS3 HOXB1 IL19 AFF4 C17ORF77 CERS4 CXCL11 ETV3 GMEB1 HOXB13 IL1A AHR C1QTNF4 CFL2 CXCL12 ETV7 GPBP1 HOXB5 IL1B AIMP1 C21ORF66 CHIA CXCL13 FAM3B GPER HOXB6 IL1F3 ALS2CR8 CBFA2T2 CIR1 CXCL14 FAM3D GPI HOXB7 IL1F5 ALX1 CBFA2T3 CITED1 CXCL16 FASLG GREM1 HOXB9 IL1F6 ARGFX CBFB CITED2 CXCL3 FBLN1 GREM2 HOXC4 IL1F7 -

CDC2 Mediates Progestin Initiated Endometrial Stromal Cell Proliferation: a PR Signaling to Gene Expression Independently of Its Binding to Chromatin
CDC2 Mediates Progestin Initiated Endometrial Stromal Cell Proliferation: A PR Signaling to Gene Expression Independently of Its Binding to Chromatin Griselda Vallejo1, Alejandro D. La Greca1., Inti C. Tarifa-Reischle1., Ana C. Mestre-Citrinovitz1, Cecilia Ballare´ 2, Miguel Beato2,3, Patricia Saragu¨ eta1* 1 Instituto de Biologı´a y Medicina Experimental, IByME-Conicet, Buenos Aires, Argentina, 2 Centre de Regulacio´ Geno`mica, (CRG), Barcelona, Spain, 3 University Pompeu Fabra (UPF), Barcelona, Spain Abstract Although non-genomic steroid receptor pathways have been studied over the past decade, little is known about the direct gene expression changes that take place as a consequence of their activation. Progesterone controls proliferation of rat endometrial stromal cells during the peri-implantation phase of pregnancy. We showed that picomolar concentration of progestin R5020 mimics this control in UIII endometrial stromal cells via ERK1-2 and AKT activation mediated by interaction of Progesterone Receptor (PR) with Estrogen Receptor beta (ERb) and without transcriptional activity of endogenous PR and ER. Here we identify early downstream targets of cytoplasmic PR signaling and their possible role in endometrial stromal cell proliferation. Microarray analysis of global gene expression changes in UIII cells treated for 45 min with progestin identified 97 up- and 341 down-regulated genes. The most over-represented molecular functions were transcription factors and regulatory factors associated with cell proliferation and cell cycle, a large fraction of which were repressors down-regulated by hormone. Further analysis verified that progestins regulate Ccnd1, JunD, Usf1, Gfi1, Cyr61, and Cdkn1b through PR- mediated activation of ligand-free ER, ERK1-2 or AKT, in the absence of genomic PR binding. -

Association of Gene Ontology Categories with Decay Rate for Hepg2 Experiments These Tables Show Details for All Gene Ontology Categories
Supplementary Table 1: Association of Gene Ontology Categories with Decay Rate for HepG2 Experiments These tables show details for all Gene Ontology categories. Inferences for manual classification scheme shown at the bottom. Those categories used in Figure 1A are highlighted in bold. Standard Deviations are shown in parentheses. P-values less than 1E-20 are indicated with a "0". Rate r (hour^-1) Half-life < 2hr. Decay % GO Number Category Name Probe Sets Group Non-Group Distribution p-value In-Group Non-Group Representation p-value GO:0006350 transcription 1523 0.221 (0.009) 0.127 (0.002) FASTER 0 13.1 (0.4) 4.5 (0.1) OVER 0 GO:0006351 transcription, DNA-dependent 1498 0.220 (0.009) 0.127 (0.002) FASTER 0 13.0 (0.4) 4.5 (0.1) OVER 0 GO:0006355 regulation of transcription, DNA-dependent 1163 0.230 (0.011) 0.128 (0.002) FASTER 5.00E-21 14.2 (0.5) 4.6 (0.1) OVER 0 GO:0006366 transcription from Pol II promoter 845 0.225 (0.012) 0.130 (0.002) FASTER 1.88E-14 13.0 (0.5) 4.8 (0.1) OVER 0 GO:0006139 nucleobase, nucleoside, nucleotide and nucleic acid metabolism3004 0.173 (0.006) 0.127 (0.002) FASTER 1.28E-12 8.4 (0.2) 4.5 (0.1) OVER 0 GO:0006357 regulation of transcription from Pol II promoter 487 0.231 (0.016) 0.132 (0.002) FASTER 6.05E-10 13.5 (0.6) 4.9 (0.1) OVER 0 GO:0008283 cell proliferation 625 0.189 (0.014) 0.132 (0.002) FASTER 1.95E-05 10.1 (0.6) 5.0 (0.1) OVER 1.50E-20 GO:0006513 monoubiquitination 36 0.305 (0.049) 0.134 (0.002) FASTER 2.69E-04 25.4 (4.4) 5.1 (0.1) OVER 2.04E-06 GO:0007050 cell cycle arrest 57 0.311 (0.054) 0.133 (0.002)