Maintenance of the Marginal Zone B Cell Compartment Specifically Requires the RNA-Binding Protein ZFP36L1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mechanical Forces Induce an Asthma Gene Signature in Healthy Airway Epithelial Cells Ayşe Kılıç1,10, Asher Ameli1,2,10, Jin-Ah Park3,10, Alvin T
www.nature.com/scientificreports OPEN Mechanical forces induce an asthma gene signature in healthy airway epithelial cells Ayşe Kılıç1,10, Asher Ameli1,2,10, Jin-Ah Park3,10, Alvin T. Kho4, Kelan Tantisira1, Marc Santolini 1,5, Feixiong Cheng6,7,8, Jennifer A. Mitchel3, Maureen McGill3, Michael J. O’Sullivan3, Margherita De Marzio1,3, Amitabh Sharma1, Scott H. Randell9, Jefrey M. Drazen3, Jefrey J. Fredberg3 & Scott T. Weiss1,3* Bronchospasm compresses the bronchial epithelium, and this compressive stress has been implicated in asthma pathogenesis. However, the molecular mechanisms by which this compressive stress alters pathways relevant to disease are not well understood. Using air-liquid interface cultures of primary human bronchial epithelial cells derived from non-asthmatic donors and asthmatic donors, we applied a compressive stress and then used a network approach to map resulting changes in the molecular interactome. In cells from non-asthmatic donors, compression by itself was sufcient to induce infammatory, late repair, and fbrotic pathways. Remarkably, this molecular profle of non-asthmatic cells after compression recapitulated the profle of asthmatic cells before compression. Together, these results show that even in the absence of any infammatory stimulus, mechanical compression alone is sufcient to induce an asthma-like molecular signature. Bronchial epithelial cells (BECs) form a physical barrier that protects pulmonary airways from inhaled irritants and invading pathogens1,2. Moreover, environmental stimuli such as allergens, pollutants and viruses can induce constriction of the airways3 and thereby expose the bronchial epithelium to compressive mechanical stress. In BECs, this compressive stress induces structural, biophysical, as well as molecular changes4,5, that interact with nearby mesenchyme6 to cause epithelial layer unjamming1, shedding of soluble factors, production of matrix proteins, and activation matrix modifying enzymes, which then act to coordinate infammatory and remodeling processes4,7–10. -

Microarray Analysis of Differentially Expressed Lncrnas with Associated Co-Expression and Cerna Networks in Coronary Heart Disease
Volume 3- Issue 1: 2018 DOI: 10.26717/BJSTR.2018.03.000886 Siying Wu. Biomed J Sci & Tech Res ISSN: 2574-1241 Research Article Open Access Microarray Analysis of Differentially Expressed LncRNAs with Associated Co-Expression and CeRNA Networks in Coronary Heart Disease Yi Sun1,a, Shuna Huang1,a, Qing Huang1,a, Guiqing Wu2, Qishuang Ruan2, Shaowei Lin1, Tingxing Zhang3, Huangyuan Li4*, Siying Wu1* 1Department of Epidemiology and Health Statistics, the Key Laboratory of Environment and Health among Universities and Colleges in Fujian, School of Public Health, Fujian Medical University, Minhou County, Fuzhou, China 2 Department of Orthopedics, Fujian Medical University Union Hospital 3Department of Cardiology, The First Affiliated Hospital of Fujian Medical University, China 4 Department of Preventive Medicine, Fujian Provincial Key Laboratory of Environmental Factors and Cancer, School of Public Health, Fujian Medical University, Minhou County, Fuzhou, China a These authors contributed equally to this work Received: February 28, 2018; Published: March 26, 2018 *Corresponding author: Siying Wu, Department of Epidemiology and Health Statistics, the Key Laboratory of Environment and Health among Universities and Colleges in Fujian, School of Public Health, Fujian Medical University, Minhou County, Fuzhou, China, Tel: ; Email: Huangyuan Li, Department of Preventive Medicine, Fujian Provincial Key Laboratory of Environmental Factors and Cancer, School of Public Health, Fujian Medical University, Minhou County, Fuzhou, China, Tel: ; Email: Abstract Objectives: coronary heart disease (CHD) and to construct a lncRNA/microRNA (miRNA)/messenger RNA (mRNA) network for mechanism exploration. The present study aims to explore the expression profiles and biological functions of long-chain noncoding RNA (lncRNA) in Methods: miRNAs, and mRNAs were evaluated using microarray. -
![CD98 [19] Among Others [5][23]](https://docslib.b-cdn.net/cover/4111/cd98-19-among-others-5-23-1304111.webp)
CD98 [19] Among Others [5][23]
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.15.439921; this version posted April 18, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Physiological Substrates and Ontogeny-Specific Expression of the Ubiquitin Ligases 2 MARCH1 and MARCH8 3 4 Patrick Schriek1, Haiyin Liu1, Alan C. Ching1, Pauline Huang1, Nishma Gupta1, Kayla R. 5 Wilson1, MinHsuang Tsai1, Yuting Yan2, Christophe F. Macri1, Laura F. Dagley3,4, Giuseppe 6 Infusini3,4, Andrew I. Webb3,4, Hamish McWilliam1,2, Satoshi Ishido5, Justine D. Mintern1 and 7 Jose A. Villadangos1,2 8 9 1Department of Biochemistry and Pharmacology, Bio21 Molecular Science and Biotechnology 10 Institute, The University of Melbourne, Parkville, VIC 3010, Australia. 11 2Department of Microbiology and Immunology, Peter Doherty Institute for Infection and 12 Immunity, The University of Melbourne, Parkville, VIC 3010, Australia. 13 3Advanced Technology and Biology Division, The Walter and Eliza Hall Institute of Medical 14 Research, Parkville, VIC 3052, Australia. 15 4Department of Medical Biology, University of Melbourne, Parkville, VIC 3010, Australia. 16 5Department of Microbiology, Hyogo College of Medicine, 1-1 Mukogawa-cho, Nishinomiya 17 17 663-8501, Japan 18 19 20 21 Correspondence to Justine D. Mintern ([email protected]) or 22 Jose A. Villadangos ([email protected]) 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.04.15.439921; this version posted April 18, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

35Th International Society for Animal Genetics Conference 7
35th INTERNATIONAL SOCIETY FOR ANIMAL GENETICS CONFERENCE 7. 23.16 – 7.27. 2016 Salt Lake City, Utah ABSTRACT BOOK https://www.asas.org/meetings/isag2016 INVITED SPEAKERS S0100 – S0124 https://www.asas.org/meetings/isag2016 epigenetic modifications, such as DNA methylation, and measuring different proteins and cellular metab- INVITED SPEAKERS: FUNCTIONAL olites. These advancements provide unprecedented ANNOTATION OF ANIMAL opportunities to uncover the genetic architecture GENOMES (FAANG) ASAS-ISAG underlying phenotypic variation. In this context, the JOINT SYMPOSIUM main challenge is to decipher the flow of biological information that lies between the genotypes and phe- notypes under study. In other words, the new challenge S0100 Important lessons from complex genomes. is to integrate multiple sources of molecular infor- T. R. Gingeras* (Cold Spring Harbor Laboratory, mation (i.e., multiple layers of omics data to reveal Functional Genomics, Cold Spring Harbor, NY) the causal biological networks that underlie complex traits). It is important to note that knowledge regarding The ~3 billion base pairs of the human DNA rep- causal relationships among genes and phenotypes can resent a storage devise encoding information for be used to predict the behavior of complex systems, as hundreds of thousands of processes that can go on well as optimize management practices and selection within and outside a human cell. This information is strategies. Here, we describe a multi-step procedure revealed in the RNAs that are composed of 12 billion for inferring causal gene-phenotype networks underly- nucleotides, considering the strandedness and allelic ing complex phenotypes integrating multi-omics data. content of each of the diploid copies of the genome. -

Clinically Annotated Breast, Ovarian and Pancreatic Cancer
www.nature.com/scientificreports OPEN MetaGxData: Clinically Annotated Breast, Ovarian and Pancreatic Cancer Datasets and their Use in Received: 19 November 2018 Accepted: 31 May 2019 Generating a Multi-Cancer Gene Published: xx xx xxxx Signature Deena M. A. Gendoo 1, Michael Zon2,4, Vandana Sandhu2, Venkata S. K. Manem2,3,5, Natchar Ratanasirigulchai2, Gregory M. Chen2, Levi Waldron 6 & Benjamin Haibe- Kains2,3,7,8,9 A wealth of transcriptomic and clinical data on solid tumours are under-utilized due to unharmonized data storage and format. We have developed the MetaGxData package compendium, which includes manually-curated and standardized clinical, pathological, survival, and treatment metadata across breast, ovarian, and pancreatic cancer data. MetaGxData is the largest compendium of curated transcriptomic data for these cancer types to date, spanning 86 datasets and encompassing 15,249 samples. Open access to standardized metadata across cancer types promotes use of their transcriptomic and clinical data in a variety of cross-tumour analyses, including identifcation of common biomarkers, and assessing the validity of prognostic signatures. Here, we demonstrate that MetaGxData is a fexible framework that facilitates meta-analyses by using it to identify common prognostic genes in ovarian and breast cancer. Furthermore, we use the data compendium to create the frst gene signature that is prognostic in a meta-analysis across 3 cancer types. These fndings demonstrate the potential of MetaGxData to serve as an important resource in oncology research, and provide a foundation for future development of cancer-specifc compendia. Ovarian, breast and pancreatic cancers are among the leading causes of cancer deaths among women, and recent studies have identifed biological and molecular commonalities between them1–4. -

GJA1 Depletion Causes Ciliary Defects by Affecting Rab11 Trafficking to the Ciliary Base
bioRxiv preprint doi: https://doi.org/10.1101/2020.11.13.381764; this version posted November 16, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Title 2 GJA1 Depletion Causes Ciliary Defects by Affecting Rab11 Trafficking to the Ciliary Base. 3 4 Author names and affiliations 5 Dong Gil Jang1, Keun Yeong Kwon1, Yeong Cheon Kweon1, Byung-gyu Kim2, Kyungjae Myung2, 6 Hyun-Shik Lee3, Chan Young Park1, Taejoon Kwon2,4,* and Tae Joo Park1,2,* 7 1 Department of Biological Sciences, College of Information-Bio Convergence Engineering, Ulsan 8 National Institute of Science and Technology, Ulsan 44919, Republic of Korea. 9 2 Center for Genomic Integrity, Institute for Basic Science, Ulsan 44919, Republic of Korea 10 3 KNU-Center for Nonlinear Dynamics, CMRI, School of Life Sciences, BK21 Plus KNU Creative 11 Bio Research Group, College of Natural Sciences, Kyungpook National University, Daegu 41566, 12 Republic of Korea 13 4 Department of Biomedical Engineering, College of Information-Bio Convergence Engineering, 14 Ulsan National Institute of Science and Technology, Ulsan 44919, Republic of Korea. 15 * Corresponding author 16 17 18 19 20 21 22 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.11.13.381764; this version posted November 16, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Transdifferentiation of Human Mesenchymal Stem Cells
Transdifferentiation of Human Mesenchymal Stem Cells Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Julius-Maximilians-Universität Würzburg vorgelegt von Tatjana Schilling aus San Miguel de Tucuman, Argentinien Würzburg, 2007 Eingereicht am: Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Martin J. Müller Gutachter: PD Dr. Norbert Schütze Gutachter: Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am: Hiermit erkläre ich ehrenwörtlich, dass ich die vorliegende Dissertation selbstständig angefertigt und keine anderen als die von mir angegebenen Hilfsmittel und Quellen verwendet habe. Des Weiteren erkläre ich, dass diese Arbeit weder in gleicher noch in ähnlicher Form in einem Prüfungsverfahren vorgelegen hat und ich noch keinen Promotionsversuch unternommen habe. Gerbrunn, 4. Mai 2007 Tatjana Schilling Table of contents i Table of contents 1 Summary ........................................................................................................................ 1 1.1 Summary.................................................................................................................... 1 1.2 Zusammenfassung..................................................................................................... 2 2 Introduction.................................................................................................................... 4 2.1 Osteoporosis and the fatty degeneration of the bone marrow..................................... 4 2.2 Adipose and bone -

Chromatin Occupancy and Target Genes of the Haematopoietic Master Transcription Factor MYB Roza B
www.nature.com/scientificreports OPEN Chromatin occupancy and target genes of the haematopoietic master transcription factor MYB Roza B. Lemma1,2,8, Marit Ledsaak1,3,8, Bettina M. Fuglerud1,4,5, Geir Kjetil Sandve6, Ragnhild Eskeland1,3,7 & Odd S. Gabrielsen1* The transcription factor MYB is a master regulator in haematopoietic progenitor cells and a pioneer factor afecting diferentiation and proliferation of these cells. Leukaemic transformation may be promoted by high MYB levels. Despite much accumulated molecular knowledge of MYB, we still lack a comprehensive understanding of its target genes and its chromatin action. In the present work, we performed a ChIP-seq analysis of MYB in K562 cells accompanied by detailed bioinformatics analyses. We found that MYB occupies both promoters and enhancers. Five clusters (C1–C5) were found when we classifed MYB peaks according to epigenetic profles. C1 was enriched for promoters and C2 dominated by enhancers. C2-linked genes were connected to hematopoietic specifc functions and had GATA factor motifs as second in frequency. C1 had in addition to MYB-motifs a signifcant frequency of ETS-related motifs. Combining ChIP-seq data with RNA-seq data allowed us to identify direct MYB target genes. We also compared ChIP-seq data with digital genomic footprinting. MYB is occupying nearly a third of the super-enhancers in K562. Finally, we concluded that MYB cooperates with a subset of the other highly expressed TFs in this cell line, as expected for a master regulator. Te transcription factor c-Myb (approved human symbol MYB), encoded by the MYB proto-oncogene, is highly expressed in haematopoietic progenitor cells and plays a key role in regulating the expression of genes involved in diferentiation and proliferation of myeloid and lymphoid progenitors 1–5. -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

Investigating Transcriptome-Wide Sex Dimorphism by Multi-Level Analysis of Single-Cell RNA Sequencing Data in Ten Mouse Cell Types Tianyuan Lu1,2 and Jessica C
Lu and Mar Biology of Sex Differences (2020) 11:61 https://doi.org/10.1186/s13293-020-00335-2 RESEARCH Open Access Investigating transcriptome-wide sex dimorphism by multi-level analysis of single-cell RNA sequencing data in ten mouse cell types Tianyuan Lu1,2 and Jessica C. Mar1* Abstract Background: It is a long established fact that sex is an important factor that influences the transcriptional regulatory processes of an organism. However, understanding sex-based differences in gene expression has been limited because existing studies typically sequence and analyze bulk tissue from female or male individuals. Such analyses average cell-specific gene expression levels where cell-to-cell variation can easily be concealed. We therefore sought to utilize data generated by the rapidly developing single cell RNA sequencing (scRNA-seq) technology to explore sex dimorphism and its functional consequences at the single cell level. Methods: Our study included scRNA-seq data of ten well-defined cell types from the brain and heart of female and male young adult mice in the publicly available tissue atlas dataset, Tabula Muris. We combined standard differential expression analysis with the identification of differential distributions in single cell transcriptomes to test for sex-based gene expression differences in each cell type. The marker genes that had sex-specific inter-cellular changes in gene expression formed the basis for further characterization of the cellular functions that were differentially regulated between the female and male cells. We also inferred activities of transcription factor-driven gene regulatory networks by leveraging knowledge of multidimensional protein-to-genome and protein-to-protein interactions and analyzed pathways that were potential modulators of sex differentiation and dimorphism. -
Downloaded from the Mouse Lysosome Gene Database, Mlgdb
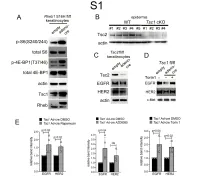
1 Supplemental Figure Legends 2 3 Supplemental Figure S1: Epidermal-specific mTORC1 gain-of-function models show 4 increased mTORC1 activation and down-regulate EGFR and HER2 protein expression in a 5 mTORC1-sensitive manner. (A) Immunoblotting of Rheb1 S16H flox/flox keratinocyte cultures 6 infected with empty or adenoviral cre recombinase for markers of mTORC1 (p-S6, p-4E-BP1) 7 activity. (B) Tsc1 cKO epidermal lysates also show decreased expression of TSC2 by 8 immunoblotting of the same experiment as in Figure 2A. (C) Immunoblotting of Tsc2 flox/flox 9 keratinocyte cultures infected with empty or adenoviral cre recombinase showing decreased EGFR 10 and HER2 protein expression. (D) Expression of EGFR and HER2 was decreased in Tsc1 cre 11 keratinocytes compared to empty controls, and up-regulated in response to Torin1 (1µM, 24 hrs), 12 by immunoblot analyses. Immunoblots are contemporaneous and parallel from the same biological 13 replicate and represent the same experiment as depicted in Figure 7B. (E) Densitometry 14 quantification of representative immunoblot experiments shown in Figures 2E and S1D (r≥3; error 15 bars represent STDEV; p-values by Student’s T-test). 16 17 18 19 20 21 22 23 Supplemental Figure S2: EGFR and HER2 transcription are unchanged with epidermal/ 24 keratinocyte Tsc1 or Rptor loss. Egfr and Her2 mRNA levels in (A) Tsc1 cKO epidermal lysates, 25 (B) Tsc1 cKO keratinocyte lysates and(C) Tsc1 cre keratinocyte lysates are minimally altered 26 compared to their respective controls. (r≥3; error bars represent STDEV; p-values by Student’s T- 27 test). -

Ncomms7336.Pdf
ARTICLE Received 18 Jul 2014 | Accepted 21 Jan 2015 | Published 19 Mar 2015 DOI: 10.1038/ncomms7336 OPEN Recurrent chromosomal gains and heterogeneous driver mutations characterise papillary renal cancer evolution Michal Kovac1,2,*, Carolina Navas3,*, Stuart Horswell4,*, Max Salm4,*, Chiara Bardella1,*, Andrew Rowan3, Mark Stares3, Francesc Castro-Giner1, Rosalie Fisher3, Elza C. de Bruin5, Monika Kovacova6, Maggie Gorman1, Seiko Makino1, Jennet Williams1, Emma Jaeger1, Angela Jones1, Kimberley Howarth1, James Larkin7, Lisa Pickering7, Martin Gore7, David L. Nicol8,9, Steven Hazell10, Gordon Stamp11, Tim O’Brien12, Ben Challacombe12, Nik Matthews13, Benjamin Phillimore13, Sharmin Begum13, Adam Rabinowitz13, Ignacio Varela14, Ashish Chandra15, Catherine Horsfield15, Alexander Polson15, Maxine Tran16, Rupesh Bhatt17, Luigi Terracciano18, Serenella Eppenberger-Castori18, Andrew Protheroe19, Eamonn Maher20, Mona El Bahrawy21, Stewart Fleming22, Peter Ratcliffe23, Karl Heinimann2, Charles Swanton3,5 & Ian Tomlinson1,24 Papillary renal cell carcinoma (pRCC) is an important subtype of kidney cancer with a problematic pathological classification and highly variable clinical behaviour. Here we sequence the genomes or exomes of 31 pRCCs, and in four tumours, multi-region sequencing is undertaken. We identify BAP1, SETD2, ARID2 and Nrf2 pathway genes (KEAP1, NHE2L2 and CUL3) as probable drivers, together with at least eight other possible drivers. However, only B10% of tumours harbour detectable pathogenic changes in any one driver gene, and where present, the mutations are often predicted to be present within cancer sub-clones. We specifically detect parallel evolution of multiple SETD2 mutations within different sub-regions of the same tumour. By contrast, large copy number gains of chromosomes 7, 12, 16 and 17 are usually early, monoclonal changes in pRCC evolution.