A Novel View of the Transcriptome Revealed from Gene Trapping In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

1 Vitamins D2 and D3 Have Overlapping but Different Effects On
medRxiv preprint doi: https://doi.org/10.1101/2020.12.16.20247700; this version posted February 22, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY 4.0 International license . Vitamins D2 and D3 have overlapping but different effects on human gene expression revealed through analysis of blood transcriptomes in a randomised double-blind placebo-controlled food-fortification trial Louise R. Durrant,1, 2, Giselda Buccaa,b, 1,3, Andrew Heskethb, 1, Carla Möller-Leveta, Laura Tripkovica, Huihai Wua, Kathryn H. Harta, John C. Mathersc, Ruan M. Elliotta, Susan A. Lanham-Newa & Colin P. Smitha,b, 3* a Department of Nutritional Sciences, School of Biosciences and Medicine, Faculty of Health and Medical Sciences, University of Surrey, Guildford, UK b School of Pharmacy and Biomolecular Sciences, University of Brighton, Brighton, UK c Human Nutrition Research Centre, Population Health Sciences Institute, Newcastle University, UK 1 These authors contributed equally 2 Current address: Yakult UK Ltd., Anteros, Odyssey Business Park, West End Road, South Ruislip, HA4 6QQ; formerly known as Louise Wilson. 3 Current address: School of Pharmacy and Biomolecular Sciences, University of Brighton, Brighton, UK *For correspondence: Prof Colin P. Smith, School of Pharmacy and Biomolecular Sciences, University of Brighton, Huxley Building, Moulsecoomb, Brighton, BN2 4GJ, UK; Tel. +44 (0)1273 642048. Email: [email protected] Key words: Vitamin D3, vitamin D2, 25-hydroxyvitamin D, human transcriptome, immunity, ethnic differences, Covid-19 Abstract For the first time, we report the influence of vitamin D2 and vitamin D3 on genome- wide gene expression in whole blood from healthy women representing two ethnic groups, white European and South Asian. -

Identification of the Binding Partners for Hspb2 and Cryab Reveals
Brigham Young University BYU ScholarsArchive Theses and Dissertations 2013-12-12 Identification of the Binding arP tners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non- Redundant Roles for Small Heat Shock Proteins Kelsey Murphey Langston Brigham Young University - Provo Follow this and additional works at: https://scholarsarchive.byu.edu/etd Part of the Microbiology Commons BYU ScholarsArchive Citation Langston, Kelsey Murphey, "Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non-Redundant Roles for Small Heat Shock Proteins" (2013). Theses and Dissertations. 3822. https://scholarsarchive.byu.edu/etd/3822 This Thesis is brought to you for free and open access by BYU ScholarsArchive. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of BYU ScholarsArchive. For more information, please contact [email protected], [email protected]. Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non-Redundant Roles for Small Heat Shock Proteins Kelsey Langston A thesis submitted to the faculty of Brigham Young University in partial fulfillment of the requirements for the degree of Master of Science Julianne H. Grose, Chair William R. McCleary Brian Poole Department of Microbiology and Molecular Biology Brigham Young University December 2013 Copyright © 2013 Kelsey Langston All Rights Reserved ABSTRACT Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactors and Non-Redundant Roles for Small Heat Shock Proteins Kelsey Langston Department of Microbiology and Molecular Biology, BYU Master of Science Small Heat Shock Proteins (sHSP) are molecular chaperones that play protective roles in cell survival and have been shown to possess chaperone activity. -

Chromosomal Aberrations in Head and Neck Squamous Cell Carcinomas in Norwegian and Sudanese Populations by Array Comparative Genomic Hybridization
825-843 12/9/08 15:31 Page 825 ONCOLOGY REPORTS 20: 825-843, 2008 825 Chromosomal aberrations in head and neck squamous cell carcinomas in Norwegian and Sudanese populations by array comparative genomic hybridization ERIC ROMAN1,2, LEONARDO A. MEZA-ZEPEDA3, STINE H. KRESSE3, OLA MYKLEBOST3,4, ENDRE N. VASSTRAND2 and SALAH O. IBRAHIM1,2 1Department of Biomedicine, Faculty of Medicine and Dentistry, University of Bergen, Jonas Lies vei 91; 2Department of Oral Sciences - Periodontology, Faculty of Medicine and Dentistry, University of Bergen, Årstadveien 17, 5009 Bergen; 3Department of Tumor Biology, Institute for Cancer Research, Rikshospitalet-Radiumhospitalet Medical Center, Montebello, 0310 Oslo; 4Department of Molecular Biosciences, University of Oslo, Blindernveien 31, 0371 Oslo, Norway Received January 30, 2008; Accepted April 29, 2008 DOI: 10.3892/or_00000080 Abstract. We used microarray-based comparative genomic logical parameters showed little correlation, suggesting an hybridization to explore genome-wide profiles of chromosomal occurrence of gains/losses regardless of ethnic differences and aberrations in 26 samples of head and neck cancers compared clinicopathological status between the patients from the two to their pair-wise normal controls. The samples were obtained countries. Our findings indicate the existence of common from Sudanese (n=11) and Norwegian (n=15) patients. The gene-specific amplifications/deletions in these tumors, findings were correlated with clinicopathological variables. regardless of the source of the samples or attributed We identified the amplification of 41 common chromosomal carcinogenic risk factors. regions (harboring 149 candidate genes) and the deletion of 22 (28 candidate genes). Predominant chromosomal alterations Introduction that were observed included high-level amplification at 1q21 (harboring the S100A gene family) and 11q22 (including Head and neck squamous cell carcinoma (HNSCC), including several MMP family members). -

Meta-Analysis of Nasopharyngeal Carcinoma
BMC Genomics BioMed Central Research article Open Access Meta-analysis of nasopharyngeal carcinoma microarray data explores mechanism of EBV-regulated neoplastic transformation Xia Chen†1,2, Shuang Liang†1, WenLing Zheng1,3, ZhiJun Liao1, Tao Shang1 and WenLi Ma*1 Address: 1Institute of Genetic Engineering, Southern Medical University, Guangzhou, PR China, 2Xiangya Pingkuang associated hospital, Pingxiang, Jiangxi, PR China and 3Southern Genomics Research Center, Guangzhou, Guangdong, PR China Email: Xia Chen - [email protected]; Shuang Liang - [email protected]; WenLing Zheng - [email protected]; ZhiJun Liao - [email protected]; Tao Shang - [email protected]; WenLi Ma* - [email protected] * Corresponding author †Equal contributors Published: 7 July 2008 Received: 16 February 2008 Accepted: 7 July 2008 BMC Genomics 2008, 9:322 doi:10.1186/1471-2164-9-322 This article is available from: http://www.biomedcentral.com/1471-2164/9/322 © 2008 Chen et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: Epstein-Barr virus (EBV) presumably plays an important role in the pathogenesis of nasopharyngeal carcinoma (NPC), but the molecular mechanism of EBV-dependent neoplastic transformation is not well understood. The combination of bioinformatics with evidences from biological experiments paved a new way to gain more insights into the molecular mechanism of cancer. Results: We profiled gene expression using a meta-analysis approach. Two sets of meta-genes were obtained. Meta-A genes were identified by finding those commonly activated/deactivated upon EBV infection/reactivation. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The Inactive X Chromosome Is Epigenetically Unstable and Transcriptionally Labile in Breast Cancer
Supplemental Information The inactive X chromosome is epigenetically unstable and transcriptionally labile in breast cancer Ronan Chaligné1,2,3,8, Tatiana Popova1,4, Marco-Antonio Mendoza-Parra5, Mohamed-Ashick M. Saleem5 , David Gentien1,6, Kristen Ban1,2,3,8, Tristan Piolot1,7, Olivier Leroy1,7, Odette Mariani6, Hinrich Gronemeyer*5, Anne Vincent-Salomon*1,4,6,8, Marc-Henri Stern*1,4,6 and Edith Heard*1,2,3,8 Extended Experimental Procedures Cell Culture Human Mammary Epithelial Cells (HMEC, Invitrogen) were grown in serum-free medium (HuMEC, Invitrogen). WI- 38, ZR-75-1, SK-BR-3 and MDA-MB-436 cells were grown in Dulbecco’s modified Eagle’s medium (DMEM; Invitrogen) containing 10% fetal bovine serum (FBS). DNA Methylation analysis. We bisulfite-treated 2 µg of genomic DNA using Epitect bisulfite kit (Qiagen). Bisulfite converted DNA was amplified with bisulfite primers listed in Table S3. All primers incorporated a T7 promoter tag, and PCR conditions are available upon request. We analyzed PCR products by MALDI-TOF mass spectrometry after in vitro transcription and specific cleavage (EpiTYPER by Sequenom®). For each amplicon, we analyzed two independent DNA samples and several CG sites in the CpG Island. Design of primers and selection of best promoter region to assess (approx. 500 bp) were done by a combination of UCSC Genome Browser (http://genome.ucsc.edu) and MethPrimer (http://www.urogene.org). All the primers used are listed (Table S3). NB: MAGEC2 CpG analysis have been done with a combination of two CpG island identified in the gene core. Analysis of RNA allelic expression profiles (based on Human SNP Array 6.0) DNA and RNA hybridizations were normalized by Genotyping console. -

Anti-ARMCX1 Monoclonal Antibody, Clone FQS23965 (DCABH-6907) This Product Is for Research Use Only and Is Not Intended for Diagnostic Use
Anti-ARMCX1 monoclonal antibody, clone FQS23965 (DCABH-6907) This product is for research use only and is not intended for diagnostic use. PRODUCT INFORMATION Product Overview Rabbit monoclonal to ARMCX1 - N-terminal Antigen Description This gene encodes a member of the ALEX family of proteins and may play a role in tumor suppression. The encoded protein contains a potential N-terminal transmembrane domain and two Armadillo (arm) repeats. Other proteins containing the arm repeat are involved in development, maintenance of tissue integrity, and tumorigenesis. This gene is closely localized with other family members, including ALEX2 and ALEX3, on the X chromosome. Immunogen Synthetic peptide (the amino acid sequence is considered to be commercially sensitive) within Human ARMCX1 aa 1-150 (N terminal). The exact sequence is proprietary.Database link: Q9P291 Isotype IgG Source/Host Rabbit Species Reactivity Human Clone FQS23965 Purity Tissue culture supernatant Conjugate Unconjugated Applications WB, ICC/IF Positive Control Human fetal heart or fetal brain lysate, HeLa cells. Format Liquid Size 100 μl Buffer pH: 7.2; Preservative: 0.01% Sodium azide; Constituents: 50% Glycerol, 0.05% BSA, PBS Preservative 0.01% Sodium Azide Storage Store at +4°C short term (1-2 weeks). Upon delivery aliquot. Store at -20°C long term. Avoid freeze / thaw cycle. Ship Shipped at 4°C. 45-1 Ramsey Road, Shirley, NY 11967, USA Email: [email protected] Tel: 1-631-624-4882 Fax: 1-631-938-8221 1 © Creative Diagnostics All Rights Reserved GENE INFORMATION -

Analysis of the Dynamics of Limb Transcriptomes During Mouse Development Istvan Gyurján1, Bernhard Sonderegger1, Felix Naef1,2 and Denis Duboule1,3*
Gyurján et al. BMC Developmental Biology 2011, 11:47 http://www.biomedcentral.com/1471-213X/11/47 RESEARCH ARTICLE Open Access Analysis of the dynamics of limb transcriptomes during mouse development Istvan Gyurján1, Bernhard Sonderegger1, Felix Naef1,2 and Denis Duboule1,3* Abstract Background: The development of vertebrate limbs has been a traditional system to study fundamental processes at work during ontogenesis, such as the establishment of spatial cellular coordinates, the effect of diffusible morphogenetic molecules or the translation between gene activity and morphogenesis. In addition, limbs are amongst the first targets of malformations in human and they display a huge realm of evolutionary variations within tetrapods, which make them a paradigm to study the regulatory genome. Results: As a reference resource for future biochemical and genetic analyses, we used genome-wide tiling arrays to establish the transcriptomes of mouse limb buds at three different stages, during which major developmental events take place. We compare the three time-points and discuss some aspects of these datasets, for instance related to transcriptome dynamics or to the potential association between active genes and the distribution of intergenic transcriptional activity. Conclusions: These datasets provide a valuable resource, either for research projects involving gene expression and regulation in developing mouse limbs, or as examples of tissue-specific, genome-wide transcriptional activities. Background regulators of the dorso-ventral (DV) patterning are Limb development has fascinated biologists for a cen- Wnt7a, expressed in dorsal ectoderm, and Engrailed,in tury, mostly because of the importance of these struc- ventral ectoderm, as well as Lmxb1,agenetranscribed tures in the evolution of land vertebrates and due to in dorsal mesenchyme. -
Download Validation Data
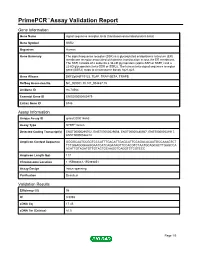
PrimePCR™Assay Validation Report Gene Information Gene Name signal sequence receptor, beta (translocon-associated protein beta) Gene Symbol SSR2 Organism Human Gene Summary The signal sequence receptor (SSR) is a glycosylated endoplasmic reticulum (ER) membrane receptor associated with protein translocation across the ER membrane. The SSR consists of 2 subunits a 34-kD glycoprotein (alpha-SSR or SSR1) and a 22-kD glycoprotein (beta-SSR or SSR2). The human beta-signal sequence receptor gene (SSR2) maps to chromosome bands 1q21-q23. Gene Aliases DKFZp686F19123, TLAP, TRAP-BETA, TRAPB RefSeq Accession No. NC_000001.10, NT_004487.19 UniGene ID Hs.74564 Ensembl Gene ID ENSG00000163479 Entrez Gene ID 6746 Assay Information Unique Assay ID qHsaCID0014663 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000295702, ENST00000529008, ENST00000480567, ENST00000531917, ENST00000526212 Amplicon Context Sequence GGGGCAATCCGGTCCCATTTGACATTGAGCATTCCAGACACAATGCCAAAGTCT TCTGGAGGGAAGGAATCATCAGATAGTTCCACGTCTAATGCAGCACTTGAGCCA ACATTGTAGATGTTGTACTGCAAGGTCAGGTCTCGTCCC Amplicon Length (bp) 117 Chromosome Location 1:155988061-155989851 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 98 R2 0.9998 cDNA Cq 17.45 cDNA Tm (Celsius) 81.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report SSR2, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference -

Variation in Protein Coding Genes Identifies Information
bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 1 1 2 3 4 5 Variation in protein coding genes identifies information flow as a contributor to 6 animal complexity 7 8 Jack Dean, Daniela Lopes Cardoso and Colin Sharpe* 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Institute of Biological and Biomedical Sciences 25 School of Biological Science 26 University of Portsmouth, 27 Portsmouth, UK 28 PO16 7YH 29 30 * Author for correspondence 31 [email protected] 32 33 Orcid numbers: 34 DLC: 0000-0003-2683-1745 35 CS: 0000-0002-5022-0840 36 37 38 39 40 41 42 43 44 45 46 47 48 49 Abstract bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 2 1 Across the metazoans there is a trend towards greater organismal complexity. How 2 complexity is generated, however, is uncertain. Since C.elegans and humans have 3 approximately the same number of genes, the explanation will depend on how genes are 4 used, rather than their absolute number. -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

This Electronic Thesis Or Dissertation Has Been Downloaded from Explore Bristol Research
This electronic thesis or dissertation has been downloaded from Explore Bristol Research, http://research-information.bristol.ac.uk Author: Al Ahdal, Hadil Title: Investigating the role of miR-21 in adult neurogenesis General rights Access to the thesis is subject to the Creative Commons Attribution - NonCommercial-No Derivatives 4.0 International Public License. A copy of this may be found at https://creativecommons.org/licenses/by-nc-nd/4.0/legalcode This license sets out your rights and the restrictions that apply to your access to the thesis so it is important you read this before proceeding. Take down policy Some pages of this thesis may have been removed for copyright restrictions prior to having it been deposited in Explore Bristol Research. However, if you have discovered material within the thesis that you consider to be unlawful e.g. breaches of copyright (either yours or that of a third party) or any other law, including but not limited to those relating to patent, trademark, confidentiality, data protection, obscenity, defamation, libel, then please contact [email protected] and include the following information in your message: •Your contact details •Bibliographic details for the item, including a URL •An outline nature of the complaint Your claim will be investigated and, where appropriate, the item in question will be removed from public view as soon as possible. Investigating the role of miR-21 in adult neurogenesis Hadil Mohammad Al Ahdal Faculty of Health Sciences Bristol Medical School A dissertation submitted to the University of Bristol in accordance with the requirements for award of the degree of Doctor of Philosophy in the Faculty of Health Sciences, Bristol Medical School 64,598 words Abstract MicroRNAs (miRNAs) are a class of small non-coding RNAs that act as post- transcriptional regulators and play important roles in neurodegenerative diseases and brain disorders (Nelson et al.