Analysis of Chromatin-State Plasticity Identifies Cell-Type–Specific Regulators of H3k27me3 Patterns
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Single Nucleotide Variants in Metastasis-Related Genes Are
View metadata, citation and similar papers at core.ac.uk brought to you by CORE HHS Public Access provided by CDC Stacks Author manuscript Author ManuscriptAuthor Manuscript Author Mol Carcinog Manuscript Author . Author manuscript; Manuscript Author available in PMC 2018 March 01. Published in final edited form as: Mol Carcinog. 2017 March ; 56(3): 1000–1009. doi:10.1002/mc.22565. Single nucleotide variants in metastasis-related genes are associated with breast cancer risk, by lymph node involvement and estrogen receptor status, in women with European and African ancestry Michelle R. Roberts1,2,3, Lara E. Sucheston-Campbell4, Gary R. Zirpoli5, Michael Higgins6, Jo L. Freudenheim3, Elisa V. Bandera7, Christine B. Ambrosone2, and Song Yao2 1Channing Division of Network Medicine, Brigham and Women’s Hospital and Harvard Medical School, Boston, MA 2Department of Cancer Prevention and Control, Roswell Park Cancer Institute, Buffalo, NY 3Department of Epidemiology and Environmental Health, University at Buffalo, Buffalo, NY 4Division of Pharmacy Practice and Science, The Ohio State University, Columbus, OH 5Department of Neurology, Massachusetts General Hospital and Harvard Medical School, Boston, MA 6Department of Molecular and Cellular Biology, Roswell Park Cancer Institute, Buffalo, NY 7Rutgers Cancer Institute of New Jersey, New Brunswick, NJ Abstract Background—Single nucleotide polymorphisms (SNPs) in pathways influencing lymph node (LN) metastasis and estrogen receptor (ER) status in breast cancer may partially explain inter- patient variability in prognosis. We examined 154 SNPs in 12 metastasis-related genes for associations with breast cancer risk, stratified by LN and ER status, in European-American (EA) and African-American (AA) women. Methods—2,671 women enrolled in the Women’s Circle of Health Study were genotyped. -

A Gene Regulatory Network Armature for T Lymphocyte Specification
Supporting Information Georgescu et al. 10.1073/pnas.0806501105 SI Text orientation in the multidimensional space, while the relative magnitude of gene expression variation is still visible on the 2D Computational Methods pls projection. All of the 2D visualizations included use the Partial Least Squares (PLS) Analysis of Normal Developmental Gene above transformations. Expression Change. PLS regression analysis was performed to find The lengths of the DN-stage vectors in the pls principal axis a hyper plane of reduced dimensionality relating the expression space indicate the extent to which these axes are explanatory of variation of the studied genes to the developmental stages DN1, the corresponding DN state. Because in the full multidimen- DN2, DN3A, DN3B, and DN4. PLS begins by characterizing the sional space this length is the same for all five DN stages, the five developmental stages DN1–DN4 as orthogonal axes in length of these vectors in the 2D projection shows the relative five-dimensional space. Each of the assayed genes is represented extent to which the DN stages are represented in the projection. as a point in this space, such that the gene’s coordinate along Long vectors indicate DN stages well represented in the projec- each axis corresponds to its expression level at the corresponding tion. In the multidimensional space, the cosine of the angle developmental stage. Genes with similar expression changes between the vectors corresponding to two variables (genes during development will tend to form clusters in this space, and/or DN stages) gives the correlation between the two vari- whereas genes with distinctly different expression profiles will ables. -

Transcriptome Dynamics and Potential Roles of Sox6 in the Postnatal Heart
RESEARCH ARTICLE Transcriptome Dynamics and Potential Roles of Sox6 in the Postnatal Heart Chung-Il An1☯*, Yasunori Ichihashi2☯¤a¤b*, Jie Peng3, Neelima R. Sinha2, Nobuko Hagiwara1* 1 Division of Cardiovascular Medicine, Department of Internal Medicine, University of California Davis, Davis, California, United States of America, 2 Department of Plant Biology, University of California Davis, Davis, California, United States of America, 3 Department of Statistics, University of California Davis, Davis, California, United States of America ☯ These authors contributed equally to this work. a11111 ¤a Current address: RIKEN Center for Sustainable Resource Science, Yokohama, Kanagawa, Japan ¤b Current address: JST, PRESTO, Kawaguchi, Saitama, Japan * [email protected] (CA); [email protected] (YI); [email protected] (NH) Abstract OPEN ACCESS The postnatal heart undergoes highly coordinated developmental processes culminating in Citation: An C-I, Ichihashi Y, Peng J, Sinha NR, the complex physiologic properties of the adult heart. The molecular mechanisms of postna- Hagiwara N (2016) Transcriptome Dynamics and tal heart development remain largely unexplored despite their important clinical implications. Potential Roles of Sox6 in the Postnatal Heart. To gain an integrated view of the dynamic changes in gene expression during postnatal PLoS ONE 11(11): e0166574. doi:10.1371/journal. heart development at the organ level, time-series transcriptome analyses of the postnatal pone.0166574 hearts of neonatal through adult mice (P1, P7, P14, P30, and P60) were performed using a Editor: Katherine Yutzey, Cincinnati Children's newly developed bioinformatics pipeline. We identified functional gene clusters by principal Hospital Medical Center, UNITED STATES component analysis with self-organizing map clustering which revealed organized, discrete Received: July 16, 2016 gene expression patterns corresponding to biological functions associated with the neona- Accepted: October 31, 2016 tal, juvenile and adult stages of postnatal heart development. -

Prox1regulates the Subtype-Specific Development of Caudal Ganglionic
The Journal of Neuroscience, September 16, 2015 • 35(37):12869–12889 • 12869 Development/Plasticity/Repair Prox1 Regulates the Subtype-Specific Development of Caudal Ganglionic Eminence-Derived GABAergic Cortical Interneurons X Goichi Miyoshi,1 Allison Young,1 Timothy Petros,1 Theofanis Karayannis,1 Melissa McKenzie Chang,1 Alfonso Lavado,2 Tomohiko Iwano,3 Miho Nakajima,4 Hiroki Taniguchi,5 Z. Josh Huang,5 XNathaniel Heintz,4 Guillermo Oliver,2 Fumio Matsuzaki,3 Robert P. Machold,1 and Gord Fishell1 1Department of Neuroscience and Physiology, NYU Neuroscience Institute, Smilow Research Center, New York University School of Medicine, New York, New York 10016, 2Department of Genetics & Tumor Cell Biology, St. Jude Children’s Research Hospital, Memphis, Tennessee 38105, 3Laboratory for Cell Asymmetry, RIKEN Center for Developmental Biology, Kobe 650-0047, Japan, 4Laboratory of Molecular Biology, Howard Hughes Medical Institute, GENSAT Project, The Rockefeller University, New York, New York 10065, and 5Cold Spring Harbor Laboratory, Cold Spring Harbor, New York 11724 Neurogliaform (RELNϩ) and bipolar (VIPϩ) GABAergic interneurons of the mammalian cerebral cortex provide critical inhibition locally within the superficial layers. While these subtypes are known to originate from the embryonic caudal ganglionic eminence (CGE), the specific genetic programs that direct their positioning, maturation, and integration into the cortical network have not been eluci- dated. Here, we report that in mice expression of the transcription factor Prox1 is selectively maintained in postmitotic CGE-derived cortical interneuron precursors and that loss of Prox1 impairs the integration of these cells into superficial layers. Moreover, Prox1 differentially regulates the postnatal maturation of each specific subtype originating from the CGE (RELN, Calb2/VIP, and VIP). -
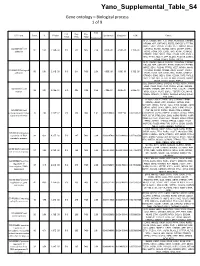
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

Accompanies CD8 T Cell Effector Function Global DNA Methylation
Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer, Benjamin G. Barwick, Benjamin A. Youngblood, Rafi Ahmed and Jeremy M. Boss This information is current as of October 1, 2021. J Immunol 2013; 191:3419-3429; Prepublished online 16 August 2013; doi: 10.4049/jimmunol.1301395 http://www.jimmunol.org/content/191/6/3419 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130139 Material 5.DC1 References This article cites 81 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/191/6/3419.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer,* Benjamin G. Barwick,* Benjamin A. Youngblood,*,† Rafi Ahmed,*,† and Jeremy M. -

Supplemental Materials and Methods
Supplemental Material for: Transcriptional silencing of γ-globin by BCL11A involves long-range interactions and cooperation with SOX6 Jian Xu, Vijay G. Sankaran, Min Ni, Tobias F. Menne, Rishi V. Puram, Woojin Kim, Stuart H. Orkin* *To whom correspondence should be addressed. E-mail: [email protected] Supplemental Materials and Methods Flow cytometry Cells at various stages of differentiation were analyzed by flow cytometry using FACSCalibur (BD Biosciences, San Jose, CA). Live cells were identified and gated by exclusion of 7-amino-actinomycin D (7-AAD; BD Pharmingen). The cells were analyzed for expression of cell surface receptors with antibodies specific for CD34, CD45, CD71, CD235, and CD36 conjugated to phycoerythrin (PE), fluorescein isothiocyanate (FITC), or allophycocyanin (APC; BD Pharmingen). Data were analyzed using FlowJo software (Ashland, OR). Cytology Cytocentrifuge preparations were stained with May-Grunwald-Giemsa as previously described (Sankaran et al. 2008). Real-time RT-PCR Real-time quantitative RT-PCR was performed using the iQ SYBR Green Supermix (Bio- Rad). The following primers were used for real-time RT-PCR: human and mouse BCL11A-XL (forward, 5’-ATGCGAGCTGTGCAACTATG-3’; reverse, 5’- GTAAACGTCCTTCCCCACCT-3’), human and mouse BCL11A-L (forward, 5’- CAGCTCAAAAGAGGGCAGAC-3’; reverse, 5’-GAGCTTCCATCCGAAAACTG-3’), and human BCL11A exon 1 and 2 (common between all known isoforms; forward, 5’- AACCCCAGCACTTAAGCAAA-3’; reverse, 5’-GGAGGTCATGATCCCCTTCT-3’). Supplemental Figure Legends Supplemental Figure 1. Expression of BCL11A isoforms in human and mouse erythroid cells. (A) Schematic diagram of human BCL11A isoforms (Liu et al. 2006). The antibodies used for ChIP experiments and their corresponding epitopes are indicated. Locations of primers used for RT-PCR analysis of all BCL11A isoforms (forward and reverse primers indicated by arrowheads), XL and L isoforms are indicated. -

Metastasis-Associated Protein 2 Represses NF-Ƙb to Reduce Lung Tumor Growth And
Author Manuscript Published OnlineFirst on August 14, 2020; DOI: 10.1158/0008-5472.CAN-20-1158 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. 1 Metastasis-associated protein 2 represses NF-ƙB to reduce lung tumor growth and 2 inflammation 3 Nefertiti El-Nikhely1#, Annika Karger1, Poonam Sarode1, Indrabahadur Singh1§, Andreas 4 Weigert2, Astrid Wietelmann1, Thorsten Stiewe3, Reinhard Dammann4, Ludger Fink5, 5 Friedrich Grimminger6, Guillermo Barreto1,7, Werner Seeger1,6,8, Soni S. Pullamsetti1,6, Ulf R. 6 Rapp1, Rajkumar Savai1,6,8* 7 Affiliations: 1Max Planck Institute for Heart and Lung Research, Member of the German 8 Center for Lung Research (DZL), Member of the Cardio-Pulmonary Institute (CPI), Bad 9 Nauheim, Germany; 2Institute of Biochemistry I, Goethe University Frankfurt, Frankfurt, 10 Germany; 3Institute of Molecular Oncology, Member of the DZL, Philipps-University 11 Marburg, Marburg, Germany; 4Institute for Genetics; member of the German Center for 12 Lung Research (DZL), Justus-Liebig-University, Giessen, Germany; 5Institute of Pathology and 13 Cytology, UEGP, Wetzlar, Germany; 6Department of Internal Medicine, Member of the DZL, 14 Member of CPI, Justus Liebig University, Giessen, Germany; 7Brain and Lung Epigenetics 15 (BLUE), Glycobiology, Cell Growth and Tissue Repair Research Unit (Gly-CRRET), Université 16 Paris-Est Créteil (UPEC), Créteil, France; 8Institute for Lung Health (ILH), Justus Liebig 17 University, Giessen, Germany; #present address: Department of Biotechnology, Institute of 18 Graduate Studies and Research, Alexandria University, Egypt; §present address: Emmy 19 Noether Research Group Epigenetic Machineries and Cancer, Division of Chronic 20 Inflammation and Cancer, German Cancer Research Center (DKFZ), Heidelberg, Germany. -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -

Transcriptional Silencing of G-Globin by BCL11A Involves Long-Range Interactions and Cooperation with SOX6
Downloaded from genesdev.cshlp.org on September 25, 2021 - Published by Cold Spring Harbor Laboratory Press Transcriptional silencing of g-globin by BCL11A involves long-range interactions and cooperation with SOX6 Jian Xu,1,2,3 Vijay G. Sankaran,1,2,4 Min Ni,5 Tobias F. Menne,1,2 Rishi V. Puram,4 Woojin Kim,1,2 and Stuart H. Orkin1,2,3,6 1Division of Hematology/Oncology, Children’s Hospital Boston, Boston, Massachusetts 02115, USA; 2Department of Pediatric Oncology, Dana-Farber Cancer Institute, Harvard Stem Cell Institute, Harvard Medical School, Boston, Massachusetts 02115, USA; 3Howard Hughes Medical Institute, Boston, Massachusetts 02115, USA; 4Harvard-MIT Division of Health Sciences and Technology, Boston, Massachusetts 02115, USA; 5Division of Molecular and Cellular Oncology, Department of Medical Oncology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, Massachusetts 02115, USA The developmental switch from human fetal (g) to adult (b) hemoglobin represents a clinically important example of developmental gene regulation. The transcription factor BCL11A is a central mediator of g-globin silencing and hemoglobin switching. Here we determine chromatin occupancy of BCL11A at the human b-globin locus and other genomic regions in vivo by high-resolution chromatin immunoprecipitation (ChIP)–chip analysis. BCL11A binds the upstream locus control region (LCR), e-globin, and the intergenic regions between g-globin and d-globin genes. A chromosome conformation capture (3C) assay shows that BCL11A reconfigures the b-globin cluster by modulating chromosomal loop formation. We also show that BCL11A and the HMG-box-containing transcription factor SOX6 interact physically and functionally during erythroid maturation. BCL11A and SOX6 co-occupy the human b-globin cluster along with GATA1, and cooperate in silencing g-globin transcription in adult human erythroid progenitors. -

1 Unique Protein Interaction Networks Define the Chromatin Remodeling
bioRxiv preprint doi: https://doi.org/10.1101/2021.01.27.428018; this version posted January 28, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Unique Protein Interaction Networks Define The Chromatin Remodeling Module of The NuRD Complex Mehdi Sharifi Tabar1,2, Caroline Giardina1, Yue Julie Feng1, Habib Francis1, Hakimeh Moghaddas Sani3, Jason K. K. Low3, Joel P. Mackay3, Charles G. Bailey1,2,4, John E.J. Rasko1,2,5,6 1Gene and Stem Cell Therapy Program Centenary Institute, The University of Sydney, Camperdown, NSW, 2050, Australia 2Faculty of Medicine & Health, The University of Sydney, Sydney, NSW, 2006, Australia 3School of Life & Environmental Sciences, The University of Sydney, Sydney, NSW, 2006, Australia 4Cancer and Gene Regulation Laboratory Centenary Institute, The University of Sydney, Camperdown, NSW, 2050, Australia 5Cell and Molecular Therapies, Royal Prince Alfred Hospital, Camperdown, NSW, 2050, Australia 6Corresponding author 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.01.27.428018; this version posted January 28, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The combination of four proteins and their paralogues including MBD2/3, GATAD2A/B, CDK2AP1, and CHD3/4/5, which we refer to as the MGCC module, form the chromatin remodeling module of the Nucleosome Remodeling and Deacetylase (NuRD) complex, a gene repressor complex. Specific paralogues of the MGCC subunits such as MBD2 and CHD4 are amongst the key repressors of adult-stage fetal globin and provide important targets for molecular therapies in beta (β)-thalassemia. -

Snail/PRMT5/Nurd Complex Contributes to DNA Hypermethylation in Cervical Cancer by TET1 Inhibition
Cell Death & Differentiation (2021) 28:2818–2836 https://doi.org/10.1038/s41418-021-00786-z ARTICLE Snail/PRMT5/NuRD complex contributes to DNA hypermethylation in cervical cancer by TET1 inhibition 1,2 2 1 3 4 4 1 Jie Gao ● Ruiqiong Liu ● Dandan Feng ● Wei Huang ● Miaomiao Huo ● Jingyao Zhang ● Shuai Leng ● 1 3 3 3 3 4 1,4 Yang Yang ● Tianshu Yang ● Xin Yin ● Xu Teng ● Hefen Yu ● Baowen Yuan ● Yan Wang Received: 15 September 2020 / Revised: 8 April 2021 / Accepted: 15 April 2021 / Published online: 5 May 2021 © The Author(s) 2021. This article is published with open access Abstract The biological function of PRMT5 remains poorly understood in cervical cancer metastasis. Here, we report that PRMT5 physically associates with the transcription factor Snail and the NuRD(MTA1) complex to form a transcriptional-repressive complex that catalyzes the symmetrical histone dimethylation and deacetylation. This study shows that the Snail/PRMT5/ NuRD(MTA1) complex targets genes, such as TET1 and E-cadherin, which are critical for epithelial-mesenchymal transition (EMT). This complex also affects the conversion of 5mC to 5hmC. This study demonstrates that the Snail/ PRMT5/NuRD(MTA1) complex promotes the invasion and metastasis of cervical cancer in vitro and in vivo. This study 1234567890();,: 1234567890();,: also shows that PRMT5 expression is upregulated in cervical cancer and various human cancers, and the PRMT5 inhibitor EPZ015666 suppresses EMT and the invasion potential of cervical cancer cells by disinhibiting the expression of TET1 and increasing 5hmC, suggesting that PRMT5 is a potential target for cancer therapy. Introduction These authors contributed equally: Jie Gao, Ruiqiong Liu, Protein arginine methyltransferase 5 (PRMT5) is a type II Dandan Feng protein arginine methyltransferase (PRMT) that has been Edited by R Johnstone reported to catalyze the symmetrical dimethylation of argi- nine [1].