An Integration of Genome- Wide Association Study and Gene
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

S1 Supplemental Materials Supplemental Methods Supplemental Figure 1. Immune Phenotype of Mcd19 Targeted CAR T and Dose Titratio
Supplemental Materials Supplemental Methods Supplemental Figure 1. Immune phenotype of mCD19 targeted CAR T and dose titration of in vivo efficacy. Supplemental Figure 2. Gene expression of fluorescent-protein tagged CAR T cells. Supplemental Figure 3. Fluorescent protein tagged CAR T cells function similarly to non-tagged counterparts. Supplemental Figure 4. Transduction efficiency and immune phenotype of mCD19 targeted CAR T cells for survival study (Figure 2D). Supplemental Figure 5. Transduction efficiency and immune phenotype of CAR T cells used in irradiated CAR T study (Fig. 3B-C). Supplemental Figure 6. Differential gene expression of CD4+ m19-humBBz CAR T cells. Supplemental Figure 7. CAR expression and CD4/CD8 subsets of human CD19 targeted CAR T cells for Figure 5E-G. Supplemental Figure 8. Transduction efficiency and immune phenotype of mCD19 targeted wild type (WT) and TRAF1-/- CAR T cells used for in vivo study (Figure 6D). Supplemental Figure 9. Mutated m19-musBBz CAR T cells have increased NF-κB signaling, improved cytokine production, anti-apoptosis, and in vivo function. Supplemental Figure 10. TRAF and CAR co-expression in human CD19-targeted CAR T cells. Supplemental Figure 11. TRAF2 over-expressed h19BBz CAR T cells show similar in vivo efficacy to h19BBz CAR T cells in an aggressive leukemia model. S1 Supplemental Table 1. Probesets increased in m19z and m1928z vs m19-musBBz CAR T cells. Supplemental Table 2. Probesets increased in m19-musBBz vs m19z and m1928z CAR T cells. Supplemental Table 3. Probesets differentially expressed in m19z vs m19-musBBz CAR T cells. Supplemental Table 4. Probesets differentially expressed in m1928z vs m19-musBBz CAR T cells. -

Variation in Protein Coding Genes Identifies Information
bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 1 1 2 3 4 5 Variation in protein coding genes identifies information flow as a contributor to 6 animal complexity 7 8 Jack Dean, Daniela Lopes Cardoso and Colin Sharpe* 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Institute of Biological and Biomedical Sciences 25 School of Biological Science 26 University of Portsmouth, 27 Portsmouth, UK 28 PO16 7YH 29 30 * Author for correspondence 31 [email protected] 32 33 Orcid numbers: 34 DLC: 0000-0003-2683-1745 35 CS: 0000-0002-5022-0840 36 37 38 39 40 41 42 43 44 45 46 47 48 49 Abstract bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 2 1 Across the metazoans there is a trend towards greater organismal complexity. How 2 complexity is generated, however, is uncertain. Since C.elegans and humans have 3 approximately the same number of genes, the explanation will depend on how genes are 4 used, rather than their absolute number. -

Genome-Wide Meta-Analysis of Common Variant Differences Between Men and Women
Human Molecular Genetics, 2012, Vol. 21, No. 21 4805–4815 doi:10.1093/hmg/dds304 Advance Access published on July 27, 2012 Genome-wide meta-analysis of common variant differences between men and women Vesna Boraska1,2,∗, Ana Jeroncˇic´ 3, Vincenza Colonna1,5, Lorraine Southam1, Dale R. Nyholt 6, Nigel William Rayner1,7,8, John R.B. Perry7,9,10, Daniela Toniolo11, Eva Albrecht12, Wei Ang13, StefaniaBandinelli14, MajaBarbalic15, IneˆsBarroso1,16, Jacques S.Beckmann18,19, ReinerBiffar20, Dorret Boomsma22, Harry Campbell23, Tanguy Corre11, Jeanette Erdmann24,25,To˜ nu Esko27,28,29, Krista Fischer27, Nora Franceschini30, Timothy M. Frayling9, Giorgia Girotto31, Juan R. Gonzalez32,33, Tamara B. Harris34, Andrew C. Heath35, Iris M. Heid36,37, Wolfgang Hoffmann21, Albert Hofman38,40, Momoko Horikoshi7,8, Jing Hua Zhao17, Anne U. Jackson41,42, Jouke-Jan Hottenga22, Antti Jula43, Mika Ka¨ho¨ nen44,46, Kay-Tee Khaw47, Lambertus A. Kiemeney48, Norman Klopp52, Zolta´n Kutalik18,54, Vasiliki Lagou7,8, Downloaded from Lenore J.Launer34, TerhoLehtima¨ki45,46, Mathieu Lemire55, Marja-LiisaLokki56, Christina Loley26, Jian’an Luan17, Massimo Mangino10, Irene Mateo Leach58, Sarah E. Medland6, Evelin Mihailov28, Grant W. Montgomery6, Gerjan Navis59, John Newnham13, Markku S. Nieminen61, Aarno Palotie1,57,62,63, Kalliope Panoutsopoulou1, Annette Peters53, Nicola Pirastu31, http://hmg.oxfordjournals.org/ Ozren Polasˇek4, Karola Rehnstro¨ m1,57, Samuli Ripatti57, Graham R.S. Ritchie1,64, Fernando Rivadeneira38,39,40, Antonietta Robino31, Nilesh J. Samani65, So-Youn Shin1, Juha Sinisalo61, Johannes H. Smit66, Nicole Soranzo1,10, Lisette Stolk39,40, Dorine W. Swinkels49, Toshiko Tanaka68, Alexander Teumer69, Anke To¨ njes70,71, Michela Traglia11, Jaakko Tuomilehto72,73,74,75, Armand Valsesia18,54,76, Wiek H. -

Distinct Gene Expression Patterns in a Tamoxifen-Sensitive Human Mammary Carcinoma Xenograft and Its Tamoxifen-Resistant Subline Maca 3366/TAM
Molecular Cancer Therapeutics 151 Distinct gene expression patterns in a tamoxifen-sensitive human mammary carcinoma xenograft and its tamoxifen-resistant subline MaCa 3366/TAM Michael Becker,1 Anette Sommer,2 of acquired tamoxifen resistance. Finally, genes whose Jo¨rn R. Kra¨tzschmar,2 Henrik Seidel,2 expression profiles are distinctly changed between the Hans-Dieter Pohlenz,2 and Iduna Fichtner1 two xenograft lines will be further evaluated as potential targets for diagnostic or therapeutic approaches of 1 Max-Delbrueck-Center for Molecular Medicine, Experimental tamoxifen-resistant breast cancer. [Mol Cancer Ther Pharmacology, and 2Research Laboratories, Schering AG, Berlin, Germany 2005;4(1):151–68] Abstract Introduction The reasons why human mammary tumors become Breast cancer is the most common type of cancer in women resistant to tamoxifen therapy are mainly unknown. of the Western world. Due to advances in early detection Changes in gene expression may occur as cells acquire and treatment, breast cancer survival rates have increased resistance to antiestrogens. We therefore undertook a com- markedly over the past decades. After surgery, estrogen parative gene expression analysis of tamoxifen-sensitive receptor a (ERa)–positive breast cancer is usually treated and tamoxifen-resistant human breast cancer in vivo with endocrine therapy. Tamoxifen, a nonsteroidal anti- models using Affymetrix oligonucleotide arrays to analyze estrogen, also termed selective estrogen receptor modula- differential gene expression. Total RNAs from the tamox- tor, is the first-line therapy for premenopausal and, until ifen-sensitive patient-derived mammary carcinoma xeno- recently, also for postmenopausal hormone receptor– graft MaCa 3366 and the tamoxifen-resistant model MaCa positive women (1). -

The Orchestra of Lipid-Transfer Proteins at the Crossroads Between Metabolism and Signaling
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Progress in Lipid Research 61 (2016) 30–39 Contents lists available at ScienceDirect Progress in Lipid Research journal homepage: www.elsevier.com/locate/plipres Review The orchestra of lipid-transfer proteins at the crossroads between metabolism and signaling Antonella Chiapparino a, Kenji Maeda a,1, Denes Turei a,b, Julio Saez-Rodriguez b,2,Anne-ClaudeGavina,c,⁎ a European Molecular Biology Laboratory (EMBL), Structural and Computational Biology Unit, Meyerhofstrasse 1, D-69117 Heidelberg, Germany b European Molecular Biology Laboratory (EMBL), European Bioinformatics Institute (EBI), Cambridge CB10 1SD, UK c European Molecular Biology Laboratory (EMBL), Molecular Medicine Partnership Unit (MMPU), Meyerhofstrasse 1, D-69117 Heidelberg, Germany article info abstract Article history: Within the eukaryotic cell, more than 1000 species of lipids define a series of membranes essential for cell func- Received 29 September 2015 tion. Tightly controlled systems of lipid transport underlie the proper spatiotemporal distribution of membrane Accepted 15 October 2015 lipids, the coordination of spatially separated lipid metabolic pathways, and lipid signaling mediated by soluble Available online 1 December 2015 proteins that may be localized some distance away from membranes. Alongside the well-established vesicular transport of lipids, non-vesicular transport mediated by a group of proteins referred to as lipid-transfer proteins Keywords: (LTPs) is emerging as a key mechanism of lipid transport in a broad range of biological processes. More than a Signaling lipid Biological membranes hundred LTPs exist in humans and these can be divided into at least ten protein families. -

Goat Anti-ORP1 Antibody Peptide-Affinity Purified Goat Antibody Catalog # Af1758a
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 Goat Anti-ORP1 Antibody Peptide-affinity purified goat antibody Catalog # AF1758a Specification Goat Anti-ORP1 Antibody - Product Information Application WB Primary Accession Q9BXW6 Other Accession NP_542164, 114876, 64291 (mouse), 259221 (rat) Reactivity Human Predicted Mouse, Rat, Dog, Cow Host Goat Clonality Polyclonal Concentration 100ug/200ul Isotype IgG Calculated MW 108470 AF1758a staining (0.5 µg/ml) of Human Goat Anti-ORP1 Antibody - Additional Information Muscle lysate (RIPA buffer, 35 µg total protein per lane). Primary incubated for 1 hour. Gene ID 114876 Detected by western blot using chemiluminescence. Other Names Oxysterol-binding protein-related protein 1, ORP-1, OSBP-related protein 1, OSBPL1A, Goat Anti-ORP1 Antibody - Background ORP1, OSBP8, OSBPL1, OSBPL1B This gene encodes a member of the Format oxysterol-binding protein (OSBP) family, a 0.5 mg IgG/ml in Tris saline (20mM Tris group of intracellular lipid receptors. Most pH7.3, 150mM NaCl), 0.02% sodium azide, members contain an N-terminal pleckstrin with 0.5% bovine serum albumin homology domain and a highly conserved Storage C-terminal OSBP-like sterol-binding domain, Maintain refrigerated at 2-8°C for up to 6 although some members contain only the months. For long term storage store at sterol-binding domain. Transcript variants -20°C in small aliquots to prevent derived from alternative promoter usage freeze-thaw cycles. and/or alternative splicing exist; they encode different isoforms. Precautions Goat Anti-ORP1 Antibody is for research use Goat Anti-ORP1 Antibody - References only and not for use in diagnostic or therapeutic procedures. -

Genomic Landscape of Paediatric Adrenocortical Tumours
ARTICLE Received 5 Aug 2014 | Accepted 16 Jan 2015 | Published 6 Mar 2015 DOI: 10.1038/ncomms7302 Genomic landscape of paediatric adrenocortical tumours Emilia M. Pinto1,*, Xiang Chen2,*, John Easton2, David Finkelstein2, Zhifa Liu3, Stanley Pounds3, Carlos Rodriguez-Galindo4, Troy C. Lund5, Elaine R. Mardis6,7,8, Richard K. Wilson6,7,9, Kristy Boggs2, Donald Yergeau2, Jinjun Cheng2, Heather L. Mulder2, Jayanthi Manne2, Jesse Jenkins10, Maria J. Mastellaro11, Bonald C. Figueiredo12, Michael A. Dyer13, Alberto Pappo14, Jinghui Zhang2, James R. Downing10, Raul C. Ribeiro14,* & Gerard P. Zambetti1,* Paediatric adrenocortical carcinoma is a rare malignancy with poor prognosis. Here we analyse 37 adrenocortical tumours (ACTs) by whole-genome, whole-exome and/or transcriptome sequencing. Most cases (91%) show loss of heterozygosity (LOH) of chromosome 11p, with uniform selection against the maternal chromosome. IGF2 on chromosome 11p is overexpressed in 100% of the tumours. TP53 mutations and chromosome 17 LOH with selection against wild-type TP53 are observed in 28 ACTs (76%). Chromosomes 11p and 17 undergo copy-neutral LOH early during tumorigenesis, suggesting tumour-driver events. Additional genetic alterations include recurrent somatic mutations in ATRX and CTNNB1 and integration of human herpesvirus-6 in chromosome 11p. A dismal outcome is predicted by concomitant TP53 and ATRX mutations and associated genomic abnormalities, including massive structural variations and frequent background mutations. Collectively, these findings demonstrate the nature, timing and potential prognostic significance of key genetic alterations in paediatric ACT and outline a hypothetical model of paediatric adrenocortical tumorigenesis. 1 Department of Biochemistry, St Jude Children’s Research Hospital, Memphis, Tennessee 38105, USA. 2 Department of Computational Biology and Bioinformatics, St Jude Children’s Research Hospital, Memphis, Tennessee 38105, USA. -

Letters to the Editor SHOX Point Mutations in Dyschondrosteosis
J Med Genet 2001;38:323–351 323 Letters to the Editor J Med Genet: first published as 10.1136/jmg.38.5.323 on 1 May 2001. Downloaded from SHOX point mutations in dyschondrosteosis Céline Huber, Veronica Cusin, Martine Le Merrer, Michèle Mathieu, Véronique Sulmont, Nathalie Dagoneau, Arnold Munnich, Valérie Cormier-Daire Dyschondrosteosis (DCS) has been recently Table 2 SHOX point mutations identified in DCS patients ascribed to mutations of the SHOX gene on the pseudoautosomal region of the X and Y chro- Sequence change Exon Protein eVect mosomes.12 Most cases are accounted for by Family 4 106–107 del CG 2 77X large scale deletions3–7 and only two point Family 5 334 C→T 3 Q112X Family 6 445 G→T 3 E149X mutations have been hitherto identified in exon Family 7 481 ins GT 3 181X 4 (R195 X and Y199X12). Here, we show that Family 8 517 C→T 4 R173C point mutations in various regions of the 1 SHOX gene also play an important role in the one nonsense mutation ). Studying eight addi- pathogenesis of the disease. tional DCS families, we describe here three A total of 22 aVected subjects belonging to deletions and five point mutations of SHOX. Taken together, these data suggest that SHOX eight families were included in the study. deletion is the most frequent common disease Inclusion criteria for aVected status were short causing mechanism in DCS (10/16, 62.5%) stature (2 SD below normal) with short and that point mutations also account for a sig- forelimbs and distal radioulnar deformity on nificant fraction of our patients (6/16, 37.5%). -
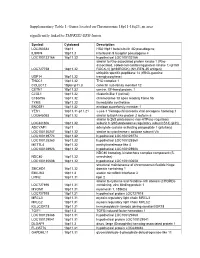
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -
Imbalanced Mitochondrial Function Provokes Heterotaxy Via Aberrant Ciliogenesis
Imbalanced mitochondrial function provokes heterotaxy via aberrant ciliogenesis Martin D. Burkhalter, … , Stephanie M. Ware, Melanie Philipp J Clin Invest. 2019;129(7):2841-2855. https://doi.org/10.1172/JCI98890. Research Article Cardiology Development Graphical abstract Find the latest version: https://jci.me/98890/pdf The Journal of Clinical Investigation RESEARCH ARTICLE Imbalanced mitochondrial function provokes heterotaxy via aberrant ciliogenesis Martin D. Burkhalter,1,2 Arthi Sridhar,3 Pedro Sampaio,4 Raquel Jacinto,4 Martina S. Burczyk,2 Cornelia Donow,2 Max Angenendt,2 Competence Network for Congenital Heart Defects Investigators,5 Maja Hempel,6 Paul Walther,7 Petra Pennekamp,8 Heymut Omran,8 Susana S. Lopes,4 Stephanie M. Ware,3 and Melanie Philipp1,2 1 Department of Experimental and Clinical Pharmacology and Pharmacogenomics, University of Tübingen, Tübingen, Germany. 2Institute of Biochemistry and Molecular Biology, Ulm University, Ulm, Germany. 3Department of Pediatrics, Indiana University School of Medicine, Indianapolis, Indiana, USA. 4CEDOC Chronic Diseases Research Center, NOVA Medical School, Faculdade de Ciências Médicas, Universidade Nova de Lisboa, Lisboa, Portugal. 5Competence Network for Congenital Heart Defects, National Register for Congenital Heart Defects, Deutsches Zentrum für Herz-Kreislaufforschung (DZHK), Berlin, Germany. 6Institute of Human Genetics, University Medical Center Hamburg-Eppendorf, Hamburg, Germany. 7Central Facility for Electron Microscopy, Ulm University, Ulm, Germany. 8Department of General Pediatrics, University Hospital Muenster, Muenster, Germany. About 1% of all newborns are affected by congenital heart disease (CHD). Recent findings identify aberrantly functioning cilia as a possible source for CHD. Faulty cilia also prevent the development of proper left-right asymmetry and cause heterotaxy, the incorrect placement of visceral organs. -

Gnomad Lof Supplement
1 gnomAD supplement gnomAD supplement 1 Data processing 4 Alignment and read processing 4 Variant Calling 4 Coverage information 5 Data processing 5 Sample QC 7 Hard filters 7 Supplementary Table 1 | Sample counts before and after hard and release filters 8 Supplementary Table 2 | Counts by data type and hard filter 9 Platform imputation for exomes 9 Supplementary Table 3 | Exome platform assignments 10 Supplementary Table 4 | Confusion matrix for exome samples with Known platform labels 11 Relatedness filters 11 Supplementary Table 5 | Pair counts by degree of relatedness 12 Supplementary Table 6 | Sample counts by relatedness status 13 Population and subpopulation inference 13 Supplementary Figure 1 | Continental ancestry principal components. 14 Supplementary Table 7 | Population and subpopulation counts 16 Population- and platform-specific filters 16 Supplementary Table 8 | Summary of outliers per population and platform grouping 17 Finalizing samples in the gnomAD v2.1 release 18 Supplementary Table 9 | Sample counts by filtering stage 18 Supplementary Table 10 | Sample counts for genomes and exomes in gnomAD subsets 19 Variant QC 20 Hard filters 20 Random Forest model 20 Features 21 Supplementary Table 11 | Features used in final random forest model 21 Training 22 Supplementary Table 12 | Random forest training examples 22 Evaluation and threshold selection 22 Final variant counts 24 Supplementary Table 13 | Variant counts by filtering status 25 Comparison of whole-exome and whole-genome coverage in coding regions 25 Variant annotation 30 Frequency and context annotation 30 2 Functional annotation 31 Supplementary Table 14 | Variants observed by category in 125,748 exomes 32 Supplementary Figure 5 | Percent observed by methylation. -
Notes and References
Notes and References Caveat: The Dangers of Behavioral Biology The contents of this book are known to be dangerous. I do not mean that in the sense that all ideas are potentially dangerous. Specifically, ideas about the biological basis of behavior have encouraged political tendencies and movements later regretted by all decent people and condemned in school histories. Why, then, purvey such ideas? Because some ideas in behavioral biology are true—among them, to the best of my knowledge, the ones in this book—and the truth is essential to wise action. But that does not mean that these ideas cannot be distorted, nor that evil acts cannot arise from them. I doubt, in fact, that what I say can prevent such distortion. Political and social movements arise from worldly causes, and then seize whatever congenial ideas are at hand. Nonethe- less, I am not comfortable in the company of scientists who are content to search for the truth and let the consequences accumulate as they may. I therefore recount here a few pas- sages in the dismal, indeed shameful history of the abuse of behavioral biology, in some of which scientists were willing participants. The first episode is recounted in William Stanton’s The Leopard’s Spots: Scientific Atti- tudes Toward Race in America, 1815–59 (Chicago: University of Chicago, 1960). Such names as Samuel George Morton, George Robins Gliddon, and Josiah Clark Nott mean little to present-day students of anthropology, but in the difficult decades between the death of Jeffer- son and the Civil War, they founded the American School of Anthropology.