Microorganisms-08-01166-V3.Pdf
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Ever-Expanding Pseudomonas Genus: Description of 43
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 14 July 2021 doi:10.20944/preprints202107.0335.v1 Article The Ever-Expanding Pseudomonas Genus: Description of 43 New Species and Partition of the Pseudomonas Putida Group Léa Girard1+, Cédric Lood1,2+, Monica Höfte3, Peter Vandamme4, Hassan Rokni-Zadeh5, Vera van Noort1,6, Rob Lavigne2*, René De Mot1,* 1 Centre of Microbial and Plant Genetics, Faculty of Bioscience Engineering, KU Leuven, Kasteelpark Aren- berg 20, 3001 Leuven, Belgium; [email protected] (L.G.), [email protected] (C.L.), [email protected] (V.v.N.) 2 Department of Biosystems, Laboratory of Gene Technology, KU Leuven, Kasteelpark Arenberg 21, 3001 Leuven, Belgium; [email protected] 3 Department of Plants and Crops, Laboratory of Phytopathology, Faculty of Bioscience Engineering, Ghent University, Ghent, Belgium 4 Laboratory of Microbiology, Department of Biochemistry and Microbiology, Faculty of Sciences, Ghent University, K. L. Ledeganckstraat 35, 9000 Ghent, Belgium; [email protected] 5 Zanjan Pharmaceutical Biotechnology Research Center, Zanjan University of Medical Sciences, 45139-56184 Zanjan, Iran; [email protected] 6 Institute of Biology, Leiden University, Sylviusweg 72, 2333 Leiden, The Netherlands + The authors contributed equally to this work. * Correspondence: [email protected], +3216379524; [email protected] ; Tel.: +3216329681 Abstract: The genus Pseudomonas hosts an extensive genetic diversity and is one of the largest genera among Gram-negative bacteria. Type strains of Pseudomonas are well-known to represent only a small fraction of this diversity and the number of available Pseudomonas genome sequences is increasing rapidly. Consequently, new Pseudomonas species are regularly reported and the number of species within the genus is in constant evolution. -

(12) United States Patent (10) Patent No.: US 7476,532 B2 Schneider Et Al
USOO7476532B2 (12) United States Patent (10) Patent No.: US 7476,532 B2 Schneider et al. (45) Date of Patent: Jan. 13, 2009 (54) MANNITOL INDUCED PROMOTER Makrides, S.C., "Strategies for achieving high-level expression of SYSTEMIS IN BACTERAL, HOST CELLS genes in Escherichia coli,” Microbiol. Rev. 60(3):512-538 (Sep. 1996). (75) Inventors: J. Carrie Schneider, San Diego, CA Sánchez-Romero, J., and De Lorenzo, V., "Genetic engineering of nonpathogenic Pseudomonas strains as biocatalysts for industrial (US); Bettina Rosner, San Diego, CA and environmental process.” in Manual of Industrial Microbiology (US) and Biotechnology, Demain, A, and Davies, J., eds. (ASM Press, Washington, D.C., 1999), pp. 460-474. (73) Assignee: Dow Global Technologies Inc., Schneider J.C., et al., “Auxotrophic markers pyrF and proC can Midland, MI (US) replace antibiotic markers on protein production plasmids in high cell-density Pseudomonas fluorescens fermentation.” Biotechnol. (*) Notice: Subject to any disclaimer, the term of this Prog., 21(2):343-8 (Mar.-Apr. 2005). patent is extended or adjusted under 35 Schweizer, H.P.. "Vectors to express foreign genes and techniques to U.S.C. 154(b) by 0 days. monitor gene expression in Pseudomonads. Curr: Opin. Biotechnol., 12(5):439-445 (Oct. 2001). (21) Appl. No.: 11/447,553 Slater, R., and Williams, R. “The expression of foreign DNA in bacteria.” in Molecular Biology and Biotechnology, Walker, J., and (22) Filed: Jun. 6, 2006 Rapley, R., eds. (The Royal Society of Chemistry, Cambridge, UK, 2000), pp. 125-154. (65) Prior Publication Data Stevens, R.C., “Design of high-throughput methods of protein pro duction for structural biology.” Structure, 8(9):R177-R185 (Sep. -

Characterisation of Pseudomonas Spp. Isolated from Foods
07.QXD 9-03-2007 15:08 Pagina 39 Annals of Microbiology, 57 (1) 39-47 (2007) Characterisation of Pseudomonas spp. isolated from foods Laura FRANZETTI*, Mauro SCARPELLINI Dipartimento di Scienze e Tecnologie Alimentari e Microbiologiche, sezione Microbiologia Agraria Alimentare Ecologica, Università degli Studi di Milano, Via Celoria 2, 20133 Milano, Italy Received 30 June 2006 / Accepted 27 December 2006 Abstract - Putative Pseudomonas spp. (102 isolates) from different foods were first characterised by API 20NE and then tested for some enzymatic activities (lipase and lecithinase production, starch hydrolysis and proteolytic activity). However subsequent molecular tests did not always confirm the results obtained, thus highlighting the limits of API 20NE. Instead RFLP ITS1 and the sequencing of 16S rRNA gene grouped the isolates into 6 clusters: Pseudomonas fluorescens (cluster I), Pseudomonas fragi (cluster II and V) Pseudomonas migulae (cluster III), Pseudomonas aeruginosa (cluster IV) and Pseudomonas chicorii (cluster VI). The pectinolytic activity was typical of species isolated from vegetable products, especially Pseudomonas fluorescens. Instead Pseudomonas fragi, predominantly isolated from meat was characterised by proteolytic and lipolytic activities. Key words: Pseudomonas fluorescens, enzymatic activity, ITS1. INTRODUCTION the most frequently found species, however the species dis- tribution within the food ecosystem remains relatively The genus Pseudomonas is the most heterogeneous and unknown (Arnaut-Rollier et al., 1999). The principal micro- ecologically significant group of known bacteria, and bial population of many vegetables in the field consists of includes Gram-negative motile aerobic rods that are wide- species of the genus Pseudomonas, especially the fluores- spread throughout nature and characterised by elevated cent forms. -

Control of Phytopathogenic Microorganisms with Pseudomonas Sp. and Substances and Compositions Derived Therefrom
(19) TZZ Z_Z_T (11) EP 2 820 140 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: A01N 63/02 (2006.01) A01N 37/06 (2006.01) 10.01.2018 Bulletin 2018/02 A01N 37/36 (2006.01) A01N 43/08 (2006.01) C12P 1/04 (2006.01) (21) Application number: 13754767.5 (86) International application number: (22) Date of filing: 27.02.2013 PCT/US2013/028112 (87) International publication number: WO 2013/130680 (06.09.2013 Gazette 2013/36) (54) CONTROL OF PHYTOPATHOGENIC MICROORGANISMS WITH PSEUDOMONAS SP. AND SUBSTANCES AND COMPOSITIONS DERIVED THEREFROM BEKÄMPFUNG VON PHYTOPATHOGENEN MIKROORGANISMEN MIT PSEUDOMONAS SP. SOWIE DARAUS HERGESTELLTE SUBSTANZEN UND ZUSAMMENSETZUNGEN RÉGULATION DE MICRO-ORGANISMES PHYTOPATHOGÈNES PAR PSEUDOMONAS SP. ET DES SUBSTANCES ET DES COMPOSITIONS OBTENUES À PARTIR DE CELLE-CI (84) Designated Contracting States: • O. COUILLEROT ET AL: "Pseudomonas AL AT BE BG CH CY CZ DE DK EE ES FI FR GB fluorescens and closely-related fluorescent GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO pseudomonads as biocontrol agents of PL PT RO RS SE SI SK SM TR soil-borne phytopathogens", LETTERS IN APPLIED MICROBIOLOGY, vol. 48, no. 5, 1 May (30) Priority: 28.02.2012 US 201261604507 P 2009 (2009-05-01), pages 505-512, XP55202836, 30.07.2012 US 201261670624 P ISSN: 0266-8254, DOI: 10.1111/j.1472-765X.2009.02566.x (43) Date of publication of application: • GUANPENG GAO ET AL: "Effect of Biocontrol 07.01.2015 Bulletin 2015/02 Agent Pseudomonas fluorescens 2P24 on Soil Fungal Community in Cucumber Rhizosphere (73) Proprietor: Marrone Bio Innovations, Inc. -
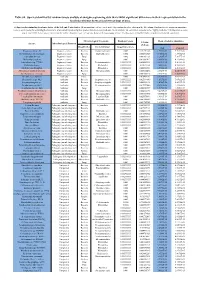
Table S8. Species Identified by Random Forests Analysis of Shotgun Sequencing Data That Exhibit Significant Differences In
Table S8. Species identified by random forests analysis of shotgun sequencing data that exhibit significant differences in their representation in the fecal microbiomes between each two groups of mice. (a) Species discriminating fecal microbiota of the Soil and Control mice. Mean importance of species identified by random forest are shown in the 5th column. Random forests assigns an importance score to each species by estimating the increase in error caused by removing that species from the set of predictors. In our analysis, we considered a species to be “highly predictive” if its importance score was at least 0.001. T-test was performed for the relative abundances of each species between the two groups of mice. P-values were at least 0.05 to be considered statistically significant. Microbiological Taxonomy Random Forests Mean of relative abundance P-Value Species Microbiological Function (T-Test) Classification Bacterial Order Importance Score Soil Control Rhodococcus sp. 2G Engineered strain Bacteria Corynebacteriales 0.002 5.73791E-05 1.9325E-05 9.3737E-06 Herminiimonas arsenitoxidans Engineered strain Bacteria Burkholderiales 0.002 0.005112829 7.1580E-05 1.3995E-05 Aspergillus ibericus Engineered strain Fungi 0.002 0.001061181 9.2368E-05 7.3057E-05 Dichomitus squalens Engineered strain Fungi 0.002 0.018887472 8.0887E-05 4.1254E-05 Acinetobacter sp. TTH0-4 Engineered strain Bacteria Pseudomonadales 0.001333333 0.025523638 2.2311E-05 8.2612E-06 Rhizobium tropici Engineered strain Bacteria Rhizobiales 0.001333333 0.02079554 7.0081E-05 4.2000E-05 Methylocystis bryophila Engineered strain Bacteria Rhizobiales 0.001333333 0.006513543 3.5401E-05 2.2044E-05 Alteromonas naphthalenivorans Engineered strain Bacteria Alteromonadales 0.001 0.000660472 2.0747E-05 4.6463E-05 Saccharomyces cerevisiae Engineered strain Fungi 0.001 0.002980726 3.9901E-05 7.3043E-05 Bacillus phage Belinda Antibiotic Phage 0.002 0.016409765 6.8789E-07 6.0681E-08 Streptomyces sp. -

High- Quality Draft Genome Sequences of Pseudomonas Monteilii DSM
RESEARCH ARTICLE Peña et al., Access Microbiology 2019;1 DOI 10.1099/acmi.0.000067 High- quality draft genome sequences of Pseudomonas monteilii DSM 14164T, Pseudomonas mosselii DSM 17497T, Pseudomonas plecoglossicida DSM 15088T, Pseudomonas taiwanensis DSM 21245T and Pseudomonas vranovensis DSM 16006T: taxonomic considerations Arantxa Peña1, Antonio Busquets1, Margarita Gomila1, Magdalena Mulet1, Rosa M. Gomila2, Elena Garcia- Valdes1,3, T. B. K. Reddy4, Marcel Huntemann4, Neha Varghese4, Natalia Ivanova4, I- Min Chen4, Markus Göker5, Tanja Woyke4, Hans- Peter Klenk6, Nikos Kyrpides4 and Jorge Lalucat1,3,* Abstract Pseudomonas is the bacterial genus of Gram- negative bacteria with the highest number of recognized species. It is divided phylogenetically into three lineages and at least 11 groups of species. The Pseudomonas putida group of species is one of the most versatile and best studied. It comprises 15 species with validly published names. As a part of the Genomic Encyclopedia of Bacteria and Archaea (GEBA) project, we present the genome sequences of the type strains of five species included in this group: Pseudomonas monteilii (DSM 14164T), Pseudomonas mosselii (DSM 17497T), Pseudomonas plecoglossicida (DSM 15088T), Pseudomonas taiwanensis (DSM 21245T) and Pseudomonas vranovensis (DSM 16006T). These strains represent species of envi- ronmental and also of clinical interest due to their pathogenic properties against humans and animals. Some strains of these species promote plant growth or act as plant pathogens. Their genome sizes are among the largest in the group, ranging from 5.3 to 6.3 Mbp. In addition, the genome sequences of the type strains in the Pseudomonas taxonomy were analysed via genome- wide taxonomic comparisons of ANIb, gANI and GGDC values among 130 Pseudomonas strains classified within the group. -

Draft Genome Sequence of Pseudomonas Moraviensis Strain Devor Implicates Metabolic Versatility and Bioremediation Potential
Genomics Data 9 (2016) 154–159 Contents lists available at ScienceDirect Genomics Data journal homepage: www.elsevier.com/locate/gdata Draft genome sequence of Pseudomonas moraviensis strain Devor implicates metabolic versatility and bioremediation potential Neil T. Miller a,DannyFullera,M.B.Cougera, Mark Bagazinski a,PhilipBoynea, Robert C. Devor a, Radwa A. Hanafy a, Connie Budd a, Donald P. French b, Wouter D. Hoff a, Noha Youssef a,⁎ a Department of Microbiology and Molecular Genetics, Oklahoma State University, Stillwater, OK, United States b Department of Integrative Biology, Oklahoma State University, Stillwater, OK, United States article info abstract Article history: Pseudomonas moraviensis is a predominant member of soil environments. We here report on the genomic Received 19 July 2016 analysis of Pseudomonas moraviensis strain Devor that was isolated from a gate at Oklahoma State University, Accepted 2 August 2016 Stillwater, OK, USA. The partial genome of Pseudomonas moraviensis strain Devor consists of 6016489 bp of Available online 4 August 2016 DNA with 5290 protein-coding genes and 66 RNA genes. This is the first detailed analysis of a P. moraviensis genome. Genomic analysis revealed metabolic versatility with genes involved in the metabolism and transport Keywords: of fructose, xylose, mannose and all amino acids with the exception of tryptophan and valine, implying that Pseudomonas moraviensis Draft genome sequence the organism is a versatile heterotroph. The genome of P. moraviensis strain Devor was rich in transporters Detailed analysis and, based on COG analysis, did not cluster closely with P. moraviensis R28-S genome, the only previous report Student Initiated Microbial Discovery (SIMD) of a P. -

Comprehensive List of Names of Plant Pathogenic Bacteria, 1980-2007
001_JPP_Letter_551 16-11-2010 14:12 Pagina 551 Journal of Plant Pathology (2010), 92 (3), 551-592 Edizioni ETS Pisa, 2010 551 LETTER TO THE EDITOR COMPREHENSIVE LIST OF NAMES OF PLANT PATHOGENIC BACTERIA, 1980-2007 C.T. Bull1, S.H. De Boer2, T.P. Denny3, G. Firrao4, M. Fischer-Le Saux5, G.S. Saddler6, M. Scortichini7, D.E. Stead8 and Y. Takikawa9 1United States Department of Agriculture, 1636 E. Alisal Street, Salinas, CA 93905, USA 2Canadian Food Inspection Agency, 93 Mount Edward Road, Charlottetown, PE C1A 5T1, Canada 3University of Georgia, Plant Pathology Department, Plant Science Building, Athens, GA 30602-7274, USA 4Dipartimento di Biologia Applicata alla Difesa delle Piante, Università degli Studi, Via Scienze 208, 33100 Udine, Italy 5UMR de Pathologie Végétale, INRA, BP 60057, 49071 Beaucouzé Cedex, France 6Science and Advice for Scottish Agriculture, Roddinglaw Road, Edinburgh EH12 9FJ, UK 7CRA. Centro di Ricerca per la Frutticoltura, Via di Fioranello 52, 00134 Roma, Italy 8Food and Environment Research Agency, Department for Environment, Food and Rural Affairs, Sand Hutton, York, YO41 1LZ, UK 9Faculty of Agriculture, Shizuoka University, 836 Ohya, Shizuoka 422-8529, Japan SUMMARY INTRODUCTION The names of all plant pathogenic bacteria which The nomenclature of bacterial plant pathogens, like have been effectively and validly published in terms of that of many other life forms, is constantly changing in the International Code of Nomenclature of Bacteria and response to new insights and our understanding of rela- the Standards for Naming Pathovars are listed to pro- tionships among bacteria. For example, the taxonomy vide an authoritative register of names for use by au- of the family Enterobacteriaceae has been extensively re- thors, journal editors and others who require access to vised since the publication of the previous comprehen- currently correct nomenclature. -

Metagenomic and Metatranscriptomic Analyses of Lake Vostok Accretion Ice
METAGENOMIC AND METATRANSCRIPTOMIC ANALYSES OF LAKE VOSTOK ACCRETION ICE Yury M. Shtarkman A Dissertation Submitted to the Graduate College of Bowling Green State University in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY December 2015 Committee: Scott O. Rogers, Advisor Rober W. Midden Graduate Faculty Representative Vipaporn Phuntumart Paul F. Morris Robert Michael McKay © 2015 Yury M Shtarkman All Rights Reserved iii ABSTRACT Scott O. Rogers, Advisor Lake Vostok (Antarctica) is the 4th deepest lake on Earth, the 6th largest by volume, and 16th largest by area, being similar in area to Ladoga Lake (Russia) and Lake Ontario (North America). However, it is a subglacial lake, constantly covered by more than 3,800 m of glacial ice, and has been covered for at least 15 million years. As the glacier slowly traverses the lake, water from the lake freezes (i.e., accretes) to the bottom of the glacier, such that on the far side of the lake a 230 m thick layer of accretion ice collects. This essentially samples various parts of the lake surface water as the glacier moves across the lake. As the glacier enters the lake, it passes over a shallow embayment. The embayment accretion ice is characterized by its silty inclusions and relatively high concentrations of several ions. It then passes over a peninsula (or island) and into the main basin. The main basin accretion ice is clear with almost no inclusions and low ion content. Metagenomic/metatranscriptomic analysis has been performed on two accretion ice samples; one from the shallow embayment and the other from part of the main lake basin. -

Pseudomonas Moraviensis Subsp. Stanleyae, a Bacterial Endophyte Of
Journal of Applied Microbiology ISSN 1364-5072 ORIGINAL ARTICLE Pseudomonas moraviensis subsp. stanleyae, a bacterial endophyte of hyperaccumulator Stanleya pinnata,is capable of efficient selenite reduction to elemental selenium under aerobic conditions L.C. Staicu1,2,3, C.J. Ackerson4, P. Cornelis5,L.Ye5, R.L. Berendsen6, W.J. Hunter7, S.D. Noblitt4, C.S. Henry4, J.J. Cappa2, R.L. Montenieri7, A.O. Wong4, L. Musilova8, M. Sura-de Jong8, E.D. van Hullebusch3, P.N.L. Lens2, R.J.B. Reynolds1 and E.A.H. Pilon-Smits1 1 Biology Department, Colorado State University, Fort Collins, CO, USA 2 UNESCO-IHE Institute for Water Education, Delft, The Netherlands 3 Universite Paris-Est, Laboratoire Geomat eriaux et Environnement, UPEM, Marne-la-Vallee, Cedex 2, France 4 Chemistry Department, Colorado State University, Fort Collins, CO, USA 5 VIB Department of Structural Biology, Department of Bioengineering Sciences, Research Group Microbiology, Vrije Universiteit, Brussels, Belgium 6 Plant–Microbe Interactions, Department of Biology, Faculty of Science, Utrecht University, Utrecht, The Netherlands 7 USDA–ARS, Fort Collins, CO, USA 8 Biochemistry and Microbiology Department, Institute of Chemical Technology in Prague, Prague, Czech Republic Keywords Abstract aerobic selenite reduction, elemental selenium nanoparticles, microchip capillary Aims: To identify bacteria with high selenium tolerance and reduction capacity electrophoresis, multi-locus sequence analysis, for bioremediation of wastewater and nanoselenium particle production. Pseudomonas moraviensis, Stanleya pinnata. Methods and Results: A bacterial endophyte was isolated from the selenium hyperaccumulator Stanleya pinnata (Brassicaceae) growing on seleniferous soils in Correspondence Colorado, USA. Based on fatty acid methyl ester analysis and multi-locus sequence Elizabeth Pilon-Smits, Biology Department, analysis (MLSA) using 16S rRNA, gyrB, rpoB and rpoD genes, the isolate was Colorado State University, 80523, Fort Collins, Á CO, USA. -

Pseudomonas Species P
INTERNATIONALJOURNAL OF SYSTEMATICBACTERIOLOGY, Apr. 1985, p. 169-184 Vol. 35, No. 2 QQ20-7713/85/020169-16$02.OO/O Copyright 0 1985, International Union of Microbiological Societies Ribosomal Ribonucleic Acid Cistron Similarities of Phytopathogenic Pseudomonas Species P. DE VOS, M. GOOR, M. GILLIS, AND J. DE LEY* Laboratorium voor Microbiologie en microhiele Genetica, Rijksuniversiteit, B-9000 Gent, Belgium Deoxyribonucleic acid (DNA)-ribosomal ribonucleic acid (rRNA) hybridizations were carried out between 23s 14C- or 3H-labeled rRNAs from type strains Pseudomonas jluorescens ATCC 13525, Pseudomonas acidovorans ATCC 15668, Pseudomonas solanacearum ATCC 11696, and Xanthomonas campestris NCPPB 528 and DNA from one or more strains of several taxonomically assigned or unassigned phytopathogenic Pseudomonas species. The strains were assigned to one of the following rRNA branches: (i) the P. Jluorescens rRNA branch (the authentic pseudomonads), containing also Pseudomonas agarici, Pseudomonas amygdali, Pseudomonas asplenii, Pseudomonas caricapapayae, Pseudomonas cichorii, Pseudomonas corrugata, Pseu- domonas fuscovaginae, Pseudomonas marginalis (P. marginalis pathovar marginalis, P. marginalis pathovar alfalfae, and P. marginalis pathovar pastinacae) , Pseudomonas meliae, the Pseudomonas syringae group (including “Pseudomonas aceris, ” “Pseudomonas antirrhini,” “Pseudomonas apii,’ ’ “Pseudomonas berber- idis,” “Pseudomonas cannabina,” ‘LPseudomonascoronafaciens”, “Pseudomonas eriobotryae,” “Pseudomo- nas lapsa,’) “Pseudomonas maculicola,” -

Lipopeptide Families at the Interface Between Pathogenic and Beneficial Pseudomonas-Plant Interactions
CRITICAL REVIEWS IN MICROBIOLOGY https://doi.org/10.1080/1040841X.2020.1794790 REVIEW ARTICLE Lipopeptide families at the interface between pathogenic and beneficial Pseudomonas-plant interactions Lea Girarda , Monica Hofte€ b and Rene De Mota aCentre of Microbial and Plant Genetics, Faculty of Bioscience Engineering, KU Leuven, Heverlee-Leuven, Belgium; bDepartment of Plants and Crops, Laboratory of Phytopathology, Faculty of Bioscience Engineering, Ghent University, Ghent, Belgium ABSTRACT ARTICLE HISTORY Lipopeptides (LPs) are a prominent class of molecules among the steadily growing spectrum of Received 10 February 2020 specialized metabolites retrieved from Pseudomonas, in particular soil-dwelling and plant-associ- Revised 27 May 2020 ated isolates. Among the multiple LP families, pioneering research focussed on phytotoxic and Accepted 4 June 2020 antimicrobial cyclic lipopeptides (CLPs) of the ubiquitous plant pathogen Pseudomonas syringae Published online 10 August (syringomycin and syringopeptin). Their non-ribosomal peptide synthetases (NRPSs) are 2020 embedded in biosynthetic gene clusters (BGCs) that are tightly co-clustered on a pathogenicity KEYWORDS island. Other members of the P. syringae group (Pseudomonas cichorii) and some species of the Mycin; peptin; factin; non- Pseudomonas asplenii group and Pseudomonas fluorescens complex have adopted these biosyn- ribosomal peptide thetic strategies to co-produce their own mycin and peptin variants, in some strains supple- synthetase; biocontrol mented with an analogue of the P. syringae linear LP (LLP), syringafactin. This capacity is not confined to phytopathogens but also occurs in some biocontrol strains, which indicates that these LP families not solely function as general virulence factors. We address this issue by scruti- nizing the structural diversity and bioactivities of LPs from the mycin, peptin, and factin families in a phylogenetic and evolutionary perspective.