University of California, San Diego
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Information to Mammadova-Bach Et Al., “Laminin Α1 Orchestrates VEGFA Functions in the Ecosystem of Colorectal Carcinogenesis”
Supplemental information to Mammadova-Bach et al., “Laminin α1 orchestrates VEGFA functions in the ecosystem of colorectal carcinogenesis” Supplemental material and methods Cloning of the villin-LMα1 vector The plasmid pBS-villin-promoter containing the 3.5 Kb of the murine villin promoter, the first non coding exon, 5.5 kb of the first intron and 15 nucleotides of the second villin exon, was generated by S. Robine (Institut Curie, Paris, France). The EcoRI site in the multi cloning site was destroyed by fill in ligation with T4 polymerase according to the manufacturer`s instructions (New England Biolabs, Ozyme, Saint Quentin en Yvelines, France). Site directed mutagenesis (GeneEditor in vitro Site-Directed Mutagenesis system, Promega, Charbonnières-les-Bains, France) was then used to introduce a BsiWI site before the start codon of the villin coding sequence using the 5’ phosphorylated primer: 5’CCTTCTCCTCTAGGCTCGCGTACGATGACGTCGGACTTGCGG3’. A double strand annealed oligonucleotide, 5’GGCCGGACGCGTGAATTCGTCGACGC3’ and 5’GGCCGCGTCGACGAATTCACGC GTCC3’ containing restriction site for MluI, EcoRI and SalI were inserted in the NotI site (present in the multi cloning site), generating the plasmid pBS-villin-promoter-MES. The SV40 polyA region of the pEGFP plasmid (Clontech, Ozyme, Saint Quentin Yvelines, France) was amplified by PCR using primers 5’GGCGCCTCTAGATCATAATCAGCCATA3’ and 5’GGCGCCCTTAAGATACATTGATGAGTT3’ before subcloning into the pGEMTeasy vector (Promega, Charbonnières-les-Bains, France). After EcoRI digestion, the SV40 polyA fragment was purified with the NucleoSpin Extract II kit (Machery-Nagel, Hoerdt, France) and then subcloned into the EcoRI site of the plasmid pBS-villin-promoter-MES. Site directed mutagenesis was used to introduce a BsiWI site (5’ phosphorylated AGCGCAGGGAGCGGCGGCCGTACGATGCGCGGCAGCGGCACG3’) before the initiation codon and a MluI site (5’ phosphorylated 1 CCCGGGCCTGAGCCCTAAACGCGTGCCAGCCTCTGCCCTTGG3’) after the stop codon in the full length cDNA coding for the mouse LMα1 in the pCIS vector (kindly provided by P. -

Microanatomy of Muscles
Microanatomy of Muscles Anatomy & Physiology Class Three Main Muscle Types Objectives: By the end of this presentation you will have the information to: 1. Describe the 3 main types of muscles. 2. Detail the functions of the muscle system. 3. Correctly label the parts of a myocyte (muscle cell) 4. Identify the levels of organization in a skeletal muscle from organ to myosin. 5. Explain how a muscle contracts utilizing the correct terminology of the sliding filament theory. 6. Contrast and compare cardiac and smooth muscle with skeletal muscle. Major Functions: Muscle System 1. Moving the skeletal system and posture. 2. Passing food through the digestive system & constriction of other internal organs. 3. Production of body heat. 4. Pumping the blood throughout the body. 5. Communication - writing and verbal Specialized Cells (Myocytes) ~ Contractile Cells Can shorten along one or more planes because of specialized cell membrane (sarcolemma) and specialized cytoskeleton. Specialized Structures found in Myocytes Sarcolemma: The cell membrane of a muscle cell Transverse tubule: a tubular invagination of the sarcolemma of skeletal or cardiac muscle fibers that surrounds myofibrils; involved in transmitting the action potential from the sarcolemma to the interior of the myofibril. Sarcoplasmic Reticulum: The special type of smooth endoplasmic Myofibrils: reticulum found in smooth and a contractile fibril of skeletal muscle, composed striated muscle fibers whose function mainly of actin and myosin is to store and release calcium ions. Multiple Nuclei (skeletal) & many mitochondria Skeletal Muscle - Microscopic Anatomy A whole skeletal muscle (such as the biceps brachii) is considered an organ of the muscular system. Each organ consists of skeletal muscle tissue, connective tissue, nerve tissue, and blood or vascular tissue. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

RESEARCH ARTICLE Body Weight-Dependent Troponin T Alternative Splicing Is Evolutionarily Conserved from Insects to Mammals and I
1523 The Journal of Experimental Biology 214, 1523-1532 © 2011. Published by The Company of Biologists Ltd doi:10.1242/jeb.051763 RESEARCH ARTICLE Body weight-dependent troponin T alternative splicing is evolutionarily conserved from insects to mammals and is partially impaired in skeletal muscle of obese rats Rudolf J. Schilder1,*, Scot R. Kimball1, James H. Marden2 and Leonard S. Jefferson1 1Department of Cellular and Molecular Physiology, The Pennsylvania State University College of Medicine, 500 University Drive, Hershey, PA 17033, USA and 2Department of Biology, The Pennsylvania State University, 208 Mueller Lab, University Park, PA 16802, USA *Author for correspondence ([email protected]) Accepted 19 January 2011 SUMMARY Do animals know at a physiological level how much they weigh, and, if so, do they make homeostatic adjustments in response to changes in body weight? Skeletal muscle is a likely tissue for such plasticity, as weight-bearing muscles receive mechanical feedback regarding body weight and consume ATP in order to generate forces sufficient to counteract gravity. Using rats, we examined how variation in body weight affected alternative splicing of fast skeletal muscle troponin T (Tnnt3), a component of the thin filament that regulates the actin–myosin interaction during contraction and modulates force output. In response to normal growth and experimental body weight increases, alternative splicing of Tnnt3 in rat gastrocnemius muscle was adjusted in a quantitative fashion. The response depended on weight per se, as externally attached loads had the same effect as an equal change in actual body weight. Examining the association between Tnnt3 alternative splicing and ATP consumption rate, we found that the Tnnt3 splice form profile had a significant association with nocturnal energy expenditure, independently of effects of weight. -

TNNT3 Rabbit Pab
Leader in Biomolecular Solutions for Life Science TNNT3 Rabbit pAb Catalog No.: A15323 Basic Information Background Catalog No. The binding of Ca(2+) to the trimeric troponin complex initiates the process of muscle A15323 contraction. Increased Ca(2+) concentrations produce a conformational change in the troponin complex that is transmitted to tropomyosin dimers situated along actin Observed MW filaments. The altered conformation permits increased interaction between a myosin 37kDa head and an actin filament which, ultimately, produces a muscle contraction. The troponin complex has protein subunits C, I, and T. Subunit C binds Ca(2+) and subunit I Calculated MW binds to actin and inhibits actin-myosin interaction. Subunit T binds the troponin 29kDa/30kDa/31kDa complex to the tropomyosin complex and is also required for Ca(2+)-mediated activation of actomyosin ATPase activity. There are 3 different troponin T genes that Category encode tissue-specific isoforms of subunit T for fast skeletal-, slow skeletal-, and cardiac-muscle. This gene encodes fast skeletal troponin T protein; also known as Primary antibody troponin T type 3. Alternative splicing results in multiple transcript variants encoding additional distinct troponin T type 3 isoforms. A developmentally regulated switch Applications between fetal/neonatal and adult troponin T type 3 isoforms occurs. Additional splice WB variants have been described but their biological validity has not been established. Mutations in this gene may cause distal arthrogryposis multiplex congenita type 2B Cross-Reactivity (DA2B). Mouse, Rat Recommended Dilutions Immunogen Information WB 1:200 - 1:2000 Gene ID Swiss Prot 7140 P45378 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 147-256 of human TNNT3 (NP_001036246.1). -

Updates on Myopia
Updates on Myopia A Clinical Perspective Marcus Ang Tien Y. Wong Editors Updates on Myopia Marcus Ang • Tien Y. Wong Editors Updates on Myopia A Clinical Perspective Editors Marcus Ang Tien Y. Wong Singapore National Eye Center Singapore National Eye Center Duke-NUS Medical School Duke-NUS Medical School National University of Singapore National University of Singapore Singapore Singapore This book is an open access publication. ISBN 978-981-13-8490-5 ISBN 978-981-13-8491-2 (eBook) https://doi.org/10.1007/978-981-13-8491-2 © The Editor(s) (if applicable) and The Author(s) 2020, corrected publication 2020 Open Access This book is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made. The images or other third party material in this book are included in the book's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the book's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. The use of general descriptive names, registered names, trademarks, service marks, etc. in this publication does not imply, even in the absence of a specifc statement, that such names are exempt from the relevant protective laws and regulations and therefore free for general use. -
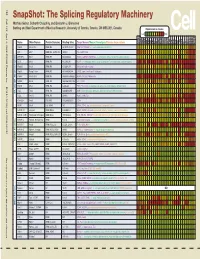
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -

A Missense Mutation in the RSRSP Stretch of Rbm20 Causes Dilated
www.nature.com/scientificreports OPEN A missense mutation in the RSRSP stretch of Rbm20 causes dilated cardiomyopathy and atrial fbrillation in mice Kensuke Ihara1,2*, Tetsuo Sasano2, Yuichi Hiraoka3, Marina Togo‑Ohno4, Yurie Soejima5, Motoji Sawabe5, Megumi Tsuchiya6, Hidesato Ogawa6, Tetsushi Furukawa1 & Hidehito Kuroyanagi4* Dilated cardiomyopathy (DCM) is a fatal heart disease characterized by left ventricular dilatation and cardiac dysfunction. Recent genetic studies on DCM have identifed causative mutations in over 60 genes, including RBM20, which encodes a regulator of heart‑specifc splicing. DCM patients with RBM20 mutations have been reported to present with more severe cardiac phenotypes, including impaired cardiac function, atrial fbrillation (AF), and ventricular arrhythmias leading to sudden cardiac death, compared to those with mutations in the other genes. An RSRSP stretch of RBM20, a hotspot of missense mutations found in patients with idiopathic DCM, functions as a crucial part of its nuclear localization signals. However, the relationship between mutations in the RSRSP stretch and cardiac phenotypes has never been assessed in an animal model. Here, we show that Rbm20 mutant mice harboring a missense mutation S637A in the RSRSP stretch, mimicking that in a DCM patient, demonstrated severe cardiac dysfunction and spontaneous AF and ventricular arrhythmias mimicking the clinical state in patients. In contrast, Rbm20 mutant mice with frame‑shifting deletion demonstrated less severe phenotypes, although loss of RBM20‑dependent alternative splicing was indistinguishable. RBM20S637A protein cannot be localized to the nuclear speckles, but accumulated in cytoplasmic, perinuclear granule‑like structures in cardiomyocytes, which might contribute to the more severe cardiac phenotypes. Dilated cardiomyopathy (DCM) is a fatal cardiac disease characterized by enlargement of the cardiac chambers and impaired systolic function1. -

Individual Protomers of a G Protein-Coupled Receptor Dimer Integrate Distinct Functional Modules
OPEN Citation: Cell Discovery (2015) 1, 15011; doi:10.1038/celldisc.2015.11 © 2015 SIBS, CAS All rights reserved 2056-5968/15 ARTICLE www.nature.com/celldisc Individual protomers of a G protein-coupled receptor dimer integrate distinct functional modules Nathan D Camp1, Kyung-Soon Lee2, Jennifer L Wacker-Mhyre2, Timothy S Kountz2, Ji-Min Park2, Dorathy-Ann Harris2, Marianne Estrada2, Aaron Stewart2, Alejandro Wolf-Yadlin1, Chris Hague2 1Department of Genome Sciences, University of Washington School of Medicine, Seattle, WA, USA; 2Department of Pharmacology, University of Washington School of Medicine, Seattle, WA, USA Recent advances in proteomic technology reveal G-protein-coupled receptors (GPCRs) are organized as large, macromolecular protein complexes in cell membranes, adding a new layer of intricacy to GPCR signaling. We previously reported the α1D-adrenergic receptor (ADRA1D)—a key regulator of cardiovascular, urinary and CNS function—binds the syntrophin family of PDZ domain proteins (SNTA, SNTB1, and SNTB2) through a C-terminal PDZ ligand inter- action, ensuring receptor plasma membrane localization and G-protein coupling. To assess the uniqueness of this novel GPCR complex, 23 human GPCRs containing Type I PDZ ligands were subjected to TAP/MS proteomic analysis. Syntrophins did not interact with any other GPCRs. Unexpectedly, a second PDZ domain protein, scribble (SCRIB), was detected in ADRA1D complexes. Biochemical, proteomic, and dynamic mass redistribution analyses indicate syntrophins and SCRIB compete for the PDZ ligand, simultaneously exist within an ADRA1D multimer, and impart divergent pharmacological properties to the complex. Our results reveal an unprecedented modular dimeric architecture for the ADRA1D in the cell membrane, providing unexpected opportunities for fine-tuning receptor function through novel protein interactions in vivo, and for intervening in signal transduction with small molecules that can stabilize or disrupt unique GPCR:PDZ protein interfaces. -

Key Genes Regulating Skeletal Muscle Development and Growth in Farm Animals
animals Review Key Genes Regulating Skeletal Muscle Development and Growth in Farm Animals Mohammadreza Mohammadabadi 1 , Farhad Bordbar 1,* , Just Jensen 2 , Min Du 3 and Wei Guo 4 1 Department of Animal Science, Faculty of Agriculture, Shahid Bahonar University of Kerman, Kerman 77951, Iran; [email protected] 2 Center for Quantitative Genetics and Genomics, Aarhus University, 8210 Aarhus, Denmark; [email protected] 3 Washington Center for Muscle Biology, Department of Animal Sciences, Washington State University, Pullman, WA 99163, USA; [email protected] 4 Muscle Biology and Animal Biologics, Animal and Dairy Science, University of Wisconsin-Madison, Madison, WI 53558, USA; [email protected] * Correspondence: [email protected] Simple Summary: Skeletal muscle mass is an important economic trait, and muscle development and growth is a crucial factor to supply enough meat for human consumption. Thus, understanding (candidate) genes regulating skeletal muscle development is crucial for understanding molecular genetic regulation of muscle growth and can be benefit the meat industry toward the goal of in- creasing meat yields. During the past years, significant progress has been made for understanding these mechanisms, and thus, we decided to write a comprehensive review covering regulators and (candidate) genes crucial for muscle development and growth in farm animals. Detection of these genes and factors increases our understanding of muscle growth and development and is a great help for breeders to satisfy demands for meat production on a global scale. Citation: Mohammadabadi, M.; Abstract: Farm-animal species play crucial roles in satisfying demands for meat on a global scale, Bordbar, F.; Jensen, J.; Du, M.; Guo, W. -

Muscle Physiology Dr
Muscle Physiology Dr. Ebneshahidi Copyright © 2004 Pearson Education, Inc., publishing as Benjamin Cummings Skeletal Muscle Figure 9.2 (a) Copyright © 2004 Pearson Education, Inc., publishing as Benjamin Cummings Functions of the muscular system . 1. Locomotion . 2. Vasoconstriction and vasodilatation- constriction and dilation of blood vessel Walls are the results of smooth muscle contraction. 3. Peristalsis – wavelike motion along the digestive tract is produced by the Smooth muscle. 4. Cardiac motion . 5. Posture maintenance- contraction of skeletal muscles maintains body posture and muscle tone. 6. Heat generation – about 75% of ATP energy used in muscle contraction is released as heat. Copyright. © 2004 Pearson Education, Inc., publishing as Benjamin Cummings . Striation: only present in skeletal and cardiac muscles. Absent in smooth muscle. Nucleus: smooth and cardiac muscles are uninculcated (one nucleus per cell), skeletal muscle is multinucleated (several nuclei per cell ). Transverse tubule ( T tubule ): well developed in skeletal and cardiac muscles to transport calcium. Absent in smooth muscle. Intercalated disk: specialized intercellular junction that only occurs in cardiac muscle. Control: skeletal muscle is always under voluntary control‚ with some exceptions ( the tongue and pili arrector muscles in the dermis). smooth and cardiac muscles are under involuntary control. Copyright © 2004 Pearson Education, Inc., publishing as Benjamin Cummings Innervation: motor unit . a) a motor nerve and a myofibril from a neuromuscular junction where gap (called synapse) occurs between the two structures. at the end of motor nerve‚ neurotransmitter (i.e. acetylcholine) is stored in synaptic vesicles which will release the neurotransmitter using exocytosis upon the stimulation of a nerve impulse. Across the synapse the surface the of myofibril contains receptors that can bind with the neurotransmitter.