Control of Tumourigenesis by the Scribble/Dlg/Lgl Polarity Module
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic and Neurodevelopmental Spectrum Of
Cognitive and behavioural genetics J Med Genet: first published as 10.1136/jmedgenet-2015-103451 on 17 March 2016. Downloaded from ORIGINAL ARTICLE Genetic and neurodevelopmental spectrum of SYNGAP1-associated intellectual disability and epilepsy Cyril Mignot,1,2,3 Celina von Stülpnagel,4,5 Caroline Nava,1,6 Dorothée Ville,7 Damien Sanlaville,8,9,10 Gaetan Lesca,8,9,10 Agnès Rastetter,6 Benoit Gachet,6 Yannick Marie,6 G Christoph Korenke,11 Ingo Borggraefe,12 Dorota Hoffmann-Zacharska,13 Elżbieta Szczepanik,14 Mariola Rudzka-Dybała,14 Uluç Yiş,15 Hande Çağlayan,16 Arnaud Isapof,17 Isabelle Marey,1 Eleni Panagiotakaki,18 Christian Korff,19 Eva Rossier,20 Angelika Riess,21 Stefanie Beck-Woedl,21 Anita Rauch,22 Christiane Zweier,23 Juliane Hoyer,23 André Reis,23 Mikhail Mironov,24 Maria Bobylova,24 Konstantin Mukhin,24 Laura Hernandez-Hernandez,25 Bridget Maher,25 Sanjay Sisodiya,25 Marius Kuhn,26 Dieter Glaeser,26 Sarah Weckhuysen,6,27 Candace T Myers,28 Heather C Mefford,28 Konstanze Hörtnagel,29 Saskia Biskup,29 EuroEPINOMICS-RES MAE working group, Johannes R Lemke,30 Delphine Héron,1,2,3,4 Gerhard Kluger,4,5 Christel Depienne1,6 ▸ Additional material is ABSTRACT INTRODUCTION published online only. To view Objective We aimed to delineate the neurodevelopmental The human SYNGAP1 gene on chromosome please visit the journal online (http://dx.doi.org/10.1136/ spectrum associated with SYNGAP1 mutations and to 6p21.3 encodes the synaptic RAS-GTPase-activating jmedgenet-2015-103451). investigate genotype–phenotype correlations. protein 1, a protein of the post-synaptic density Methods We sequenced the exome or screened the exons (PSD) of glutamatergic neurons.12SYNGAP1 inter- For numbered affiliations see end of article. -
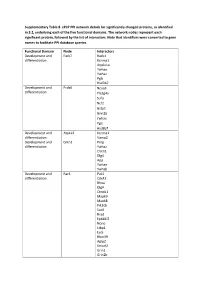
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Artificial Intelligence-Based Computational Framework for Drug
Tsuji et al. Alzheimer’s Research & Therapy (2021) 13:92 https://doi.org/10.1186/s13195-021-00826-3 RESEARCH Open Access Artificial intelligence-based computational framework for drug-target prioritization and inference of novel repositionable drugs for Alzheimer’s disease Shingo Tsuji1*† , Takeshi Hase2,3,4,5†, Ayako Yachie-Kinoshita2,4, Taiko Nishino2,SamikGhosh2, Masataka Kikuchi6, Kazuro Shimokawa7, Hiroyuki Aburatani1, Hiroaki Kitano2 and Hiroshi Tanaka3 Abstract Background: Identifying novel therapeutic targets is crucial for the successful development of drugs. However, the cost to experimentally identify therapeutic targets is huge and only approximately 400 genes are targets for FDA-approved drugs. As a result, it is inevitable to develop powerful computational tools that can identify potential novel therapeutic targets. Fortunately, the human protein-protein interaction network (PIN) could be a useful resource to achieve this objective. Methods: In this study, we developed a deep learning-based computational framework that extracts low-dimensional representations of high-dimensional PIN data. Our computational framework uses latent features and state-of-the-art machine learning techniques to infer potential drug target genes. Results: We applied our computational framework to prioritize novel putative target genes for Alzheimer’s disease and successfully identified key genes that may serve as novel therapeutic targets (e.g., DLG4, EGFR, RAC1, SYK, PTK2B, SOCS1). Furthermore, based on these putative targets, we could infer repositionable -

47421 SAP102 (A7R8L) Rabbit Mab
Revision 1 C 0 2 - t SAP102 (A7R8L) Rabbit mAb a e r o t S Orders: 877-616-CELL (2355) [email protected] 1 Support: 877-678-TECH (8324) 2 4 Web: [email protected] 7 www.cellsignal.com 4 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source/Isotype: UniProt ID: Entrez-Gene Id: WB, IP H M R Endogenous 102 Rabbit Q92796 1741 Product Usage Information 1. Cuthbert, P.C. et al. (2007) J Neurosci 27, 2673-82. 2. Makino, K. et al. (1997) Oncogene 14, 2425-33. Application Dilution 3. Masuko, N. et al. (1999) J Biol Chem 274, 5782-90. 4. Lau, L.F. et al. (1996) J Biol Chem 271, 21622-8. Western Blotting 1:1000 5. Yan, J. et al. (2005) Mol Psychiatry 10, 329-32. Immunoprecipitation 1:50 Storage Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA, 50% glycerol and less than 0.02% sodium azide. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity SAP102 (A7R8L) Rabbit mAb recognizes endogenous levels of total SAP102 protein. Species Reactivity: Human, Mouse, Rat Source / Purification Monoclonal antibody is produced by immunizing animals with a synthetic peptide corresponding to residues near the carboxy terminus of human SAP102 protein. Background Synapse-associated protein 102 (SAP102, DLG3) belongs to the membrane-associated guanylate kinase (MAGUK) protein family and is a homolog of the Drosophila disc large (dlg) tumor suppressor protein. SAP102 consists of three PDZ domains, a Src homology 3 (SH3) domain, and a guanylate kinase (GK) domain (1). -

LGI1–ADAM22–MAGUK Configures Transsynaptic Nanoalignment for Synaptic Transmission and Epilepsy Prevention
LGI1–ADAM22–MAGUK configures transsynaptic nanoalignment for synaptic transmission and epilepsy prevention Yuko Fukataa,b,1, Xiumin Chenc,d,1, Satomi Chikenb,e, Yoko Hiranoa,f, Atsushi Yamagatag, Hiroki Inahashia, Makoto Sanboh, Hiromi Sanob,e, Teppei Gotoh, Masumi Hirabayashib,h, Hans-Christian Kornaui,j, Harald Prüssi,k, Atsushi Nambub,e, Shuya Fukail, Roger A. Nicollc,d,2, and Masaki Fukataa,b,2 aDivision of Membrane Physiology, Department of Molecular and Cellular Physiology, National Institute for Physiological Sciences, National Institutes of Natural Sciences, Aichi 444-8787, Japan; bDepartment of Physiological Sciences, School of Life Science, SOKENDAI (The Graduate University for Advanced Studies), Aichi 444-8585, Japan; cDepartment of Cellular and Molecular Pharmacology, University of California, San Francisco, CA 94158; dDepartment of Physiology, University of California, San Francisco, CA 94158; eDivision of System Neurophysiology, Department of System Neuroscience, National Institute for Physiological Sciences, National Institutes of Natural Sciences, Aichi 444-8585, Japan; fDepartment of Pediatrics, Graduate School of Medicine, The University of Tokyo, Tokyo 113-8655, Japan; gLaboratory for Protein Functional and Structural Biology, RIKEN Center for Biosystems Dynamics Research, Kanagawa 230-0045, Japan; hCenter for Genetic Analysis of Behavior, National Institute for Physiological Sciences, National Institutes of Natural Sciences, Okazaki 444-8787, Japan; iGerman Center for Neurodegenerative Diseases (DZNE), 10117 Berlin, Germany; jNeuroscience Research Center, Cluster NeuroCure, Charité-Universitätsmedizin Berlin, 10117 Berlin, Germany; kDepartment of Neurology and Experimental Neurology, Charité-Universitätsmedizin Berlin, 10117 Berlin, Germany; and lDepartment of Chemistry, Graduate School of Science, Kyoto University, 606-8502 Kyoto, Japan Contributed by Roger A. Nicoll, December 1, 2020 (sent for review October 29, 2020; reviewed by David S. -

Meta-Analytical Biomarker Search of EST Expression Data Reveals Three
Wu et al. BMC Genomics 2012, 13(Suppl 7):S12 http://www.biomedcentral.com/1471-2164/13/S7/S12 PROCEEDINGS Open Access Meta-analytical biomarker search of EST expression data reveals three differentially expressed candidates Timothy H Wu1, Lichieh J Chu2, Jian-Chiao Wang3, Ting-Wen Chen2,4, Yin-Jing Tien5, Wen-Chang Lin1,6, Wailap V Ng1,3,7* From Asia Pacific Bioinformatics Network (APBioNet) Eleventh International Conference on Bioinformatics (InCoB2012) Bangkok, Thailand. 3-5 October 2012 Abstract Background: Researches have been conducted for the identification of differentially expressed genes (DEGs) by generating and mining of cDNA expressed sequence tags (ESTs) for more than a decade. Although the availability of public databases make possible the comprehensive mining of DEGs among the ESTs from multiple tissue types, existing studies usually employed statistics suitable only for two categories. Multi-class test has been developed to enable the finding of tissue specific genes, but subsequent search for cancer genes involves separate two-category test only on the ESTs of the tissue of interest. This constricts the amount of data used. On the other hand, simple pooling of cancer and normal genes from multiple tissue types runs the risk of Simpson’s paradox. Here we presented a different approach which searched for multi-cancer DEG candidates by analyzing all pertinent ESTs in all categories and narrowing down the cancer biomarker candidates via integrative analysis with microarray data and selection of secretory and membrane protein genes as well as incorporation of network analysis. Finally, the differential expression patterns of three selected cancer biomarker candidates were confirmed by real-time qPCR analysis. -

Perkinelmer Genomics to Request the Saliva Swab Collection Kit for Patients That Cannot Provide a Blood Sample As Whole Blood Is the Preferred Sample
Autism and Intellectual Disability TRIO Panel Test Code TR002 Test Summary This test analyzes 2429 genes that have been associated with Autism and Intellectual Disability and/or disorders associated with Autism and Intellectual Disability with the analysis being performed as a TRIO Turn-Around-Time (TAT)* 3 - 5 weeks Acceptable Sample Types Whole Blood (EDTA) (Preferred sample type) DNA, Isolated Dried Blood Spots Saliva Acceptable Billing Types Self (patient) Payment Institutional Billing Commercial Insurance Indications for Testing Comprehensive test for patients with intellectual disability or global developmental delays (Moeschler et al 2014 PMID: 25157020). Comprehensive test for individuals with multiple congenital anomalies (Miller et al. 2010 PMID 20466091). Patients with autism/autism spectrum disorders (ASDs). Suspected autosomal recessive condition due to close familial relations Previously negative karyotyping and/or chromosomal microarray results. Test Description This panel analyzes 2429 genes that have been associated with Autism and ID and/or disorders associated with Autism and ID. Both sequencing and deletion/duplication (CNV) analysis will be performed on the coding regions of all genes included (unless otherwise marked). All analysis is performed utilizing Next Generation Sequencing (NGS) technology. CNV analysis is designed to detect the majority of deletions and duplications of three exons or greater in size. Smaller CNV events may also be detected and reported, but additional follow-up testing is recommended if a smaller CNV is suspected. All variants are classified according to ACMG guidelines. Condition Description Autism Spectrum Disorder (ASD) refers to a group of developmental disabilities that are typically associated with challenges of varying severity in the areas of social interaction, communication, and repetitive/restricted behaviors. -

Genetic and Neurodevelopmental Spectrum Of
Downloaded from http://jmg.bmj.com/ on November 23, 2016 - Published by group.bmj.com JMG Online First, published on March 17, 2016 as 10.1136/jmedgenet-2015-103451 Cognitive and behavioural genetics ORIGINAL ARTICLE Genetic and neurodevelopmental spectrum of SYNGAP1-associated intellectual disability and epilepsy Cyril Mignot,1,2,3 Celina von Stülpnagel,4,5 Caroline Nava,1,6 Dorothée Ville,7 Damien Sanlaville,8,9,10 Gaetan Lesca,8,9,10 Agnès Rastetter,6 Benoit Gachet,6 Yannick Marie,6 G Christoph Korenke,11 Ingo Borggraefe,12 Dorota Hoffmann-Zacharska,13 Elżbieta Szczepanik,14 Mariola Rudzka-Dybała,14 Uluç Yiş,15 Hande Çağlayan,16 Arnaud Isapof,17 Isabelle Marey,1 Eleni Panagiotakaki,18 Christian Korff,19 Eva Rossier,20 Angelika Riess,21 Stefanie Beck-Woedl,21 Anita Rauch,22 Christiane Zweier,23 Juliane Hoyer,23 André Reis,23 Mikhail Mironov,24 Maria Bobylova,24 Konstantin Mukhin,24 Laura Hernandez-Hernandez,25 Bridget Maher,25 Sanjay Sisodiya,25 Marius Kuhn,26 Dieter Glaeser,26 Sarah Wechuysen,6,27 Candace T Myers,28 Heather C Mefford,28 Konstanze Hörtnagel,29 Saskia Biskup,29 EuroEPINOMICS-RES MAE working group, Johannes R Lemke,30 Delphine Héron,1,2,3,4 Gerhard Kluger,4,5 Christel Depienne1,6 ▸ Additional material is ABSTRACT INTRODUCTION published online only. To view Objective We aimed to delineate the neurodevelopmental The human SYNGAP1 gene on chromosome please visit the journal online (http://dx.doi.org/10.1136/ spectrum associated with SYNGAP1 mutations and to 6p21.3 encodes the synaptic RAS-GTPase-activating jmedgenet-2015-103451). investigate genotype–phenotype correlations. -

X-Linked Mental Retardation
REVIEWS X-LINKED MENTAL RETARDATION H.-Hilger Ropers* and Ben C. J. Hamel‡ Abstract | Genetic factors have an important role in the aetiology of mental retardation. However, their contribution is often underestimated because in developed countries, severely affected patients are mainly sporadic cases and familial cases are rare. X-chromosomal mental retardation is the exception to this rule, and this is one of the reasons why research into the genetic and molecular causes of mental retardation has focused almost entirely on the X-chromosome. Here, we review the remarkable recent progress in this field, its promise for understanding neural function, learning and memory, and the implications of this research for health care. Mental retardation is one of the main reasons for refer- subject (by Chelly and Mandel7) was published. Pro- ral in paediatric, child-neurological and clinical genetic gress has been particularly spectacular for so-called practice. Often, however, despite extensive investiga- non-syndromic or ‘pure’ forms of XLMR, consisting of tions, an aetiological diagnosis cannot be made, leaving numerous disorders that are clinically indistinguishable families without accurate genetic counselling or repro- because cognitive impairment is their only manifestation. ductive options, such as prenatal diagnosis. The preva- In this review, we describe the strategies used to iden- lence of mental retardation in developed countries is tify all genes that have been implicated in XLMR so far, thought to be on the order of 2–3% (REF.1), although emphasizing the crucial importance of large cohorts of estimates vary widely, particularly for mild mental retar- well-characterized families. We provide a brief summary dation. -

De Novo CNV Analysis Implicates Specific Abnormalities of Postsynaptic Signalling Complexes in the Pathogenesis of Schizophrenia
Molecular Psychiatry (2012) 17, 142–153 OPEN & 2012 Macmillan Publishers Limited All rights reserved 1359-4184/12 www.nature.com/mp IMMEDIATE COMMUNICATION De novo CNV analysis implicates specific abnormalities of postsynaptic signalling complexes in the pathogenesis of schizophrenia G Kirov1, AJ Pocklington1, P Holmans1, D Ivanov1, M Ikeda2, D Ruderfer3,4,5, J Moran5, K Chambert5, D Toncheva6, L Georgieva1, D Grozeva1, M Fjodorova1, R Wollerton1, E Rees1, I Nikolov1, LN van de Lagemaat7,A` Baye´s7, E Fernandez8, PI Olason9,YBo¨ttcher9, NH Komiyama7, MO Collins10, J Choudhary10, K Stefansson9, H Stefansson9, SGN Grant7, S Purcell3,4,5, P Sklar3,4,5, MC O’Donovan1 and MJ Owen1 1Department of Psychological Medicine and Neurology, MRC Centre for Neuropsychiatric Genetics and Genomics, School of Medicine, Neuroscience and Mental Health Research Institute, Cardiff University, Cardiff, UK; 2Department of Psychiatry, School of Medicine, Fujita Health University, Toyoake, Aichi, Japan; 3Department of Psychiatry, Psychiatric and Neurodevelopmental Genetics Unit, Massachusetts General Hospital and Harvard Medical School, Boston, MA, USA; 4Center for Human Genetics Research, Massachusetts General Hospital, Boston, MA, USA; 5Stanley Center for Psychiatric Research, Broad Institute, Cambridge, MA, USA; 6University Hospital Maichin Dom, Sofia, Bulgaria; 7Genes to Cognition Program, School of Molecular and Clinical Medicine, Edinburgh University, Edinburgh, UK; 8VIB Department of Molecular and Developmental Genetics, KU Leuven Medical School, Leuven, Belgium; 9deCODE Genetics, Reykjavı´k, Iceland and 10The Wellcome Trust Sanger Institute, Hinxton, Cambridgeshire, UK A small number of rare, recurrent genomic copy number variants (CNVs) are known to substantially increase susceptibility to schizophrenia. As a consequence of the low fecundity in people with schizophrenia and other neurodevelopmental phenotypes to which these CNVs contribute, CNVs with large effects on risk are likely to be rapidly removed from the population by natural selection. -

X-Linked Mental Retardation: a Clinical Guide
JMG Online First, published on August 23, 2005 as 10.1136/jmg.2005.033043 J Med Genet: first published as 10.1136/jmg.2005.033043 on 23 August 2005. Downloaded from X-linked Mental Retardation: a clinical guide F Lucy Raymond University Lecturer and Honorary Consultant in Medical Genetics Running title: XLMR review http://jmg.bmj.com/ Cambridge Institute of Medical Research, Department of Medical Genetics, on September 30, 2021 by guest. Protected copyright. University of Cambridge, Addenbrookes Hospital Cambridge CB2 2XY, UK T: 44 (0) 1223 762609 F: 44 (0)1223 331206 Correspondence: [email protected] Key words: Mental Retardation; X chromosome; recurrence risks; X-linked; XLMR XLMR review 1 Copyright Article author (or their employer) 2005. Produced by BMJ Publishing Group Ltd under licence. J Med Genet: first published as 10.1136/jmg.2005.033043 on 23 August 2005. Downloaded from Abstract Mental retardation is more common in males than females in the population and the predominant cause of this is the presence of mutations in any one of 24 genes on the X chromosome. The prevalence of each gene as a cause of mental retardation is low and less common than Fragile X syndrome. Expansions in FMR1 are still the most common cause of X-linked mental retardation. Systematic screening of all other X- linked genes in X-linked families with mental retardation is currently not feasible in a clinical setting. This review discusses the phenotypes of genes that cause syndromic and non-syndromic mental retardation, as these may be the focus of more targeted mutation analysis: NLGN3, NLGN4, RPS6KA3(RSK2), OPHN1, ATRX, SLC6A8, ARX, SYN1, AGTR2, MECP2, PQBP1, SMCX and SLC16A2. -

Autocrine IFN Signaling Inducing Profibrotic Fibroblast Responses By
Downloaded from http://www.jimmunol.org/ by guest on September 23, 2021 Inducing is online at: average * The Journal of Immunology , 11 of which you can access for free at: 2013; 191:2956-2966; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication August 2013; doi: 10.4049/jimmunol.1300376 http://www.jimmunol.org/content/191/6/2956 A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Autocrine IFN Signaling Feng Fang, Kohtaro Ooka, Xiaoyong Sun, Ruchi Shah, Swati Bhattacharyya, Jun Wei and John Varga J Immunol cites 49 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130037 6.DC1 This article http://www.jimmunol.org/content/191/6/2956.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 23, 2021. The Journal of Immunology A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Inducing Autocrine IFN Signaling Feng Fang,* Kohtaro Ooka,* Xiaoyong Sun,† Ruchi Shah,* Swati Bhattacharyya,* Jun Wei,* and John Varga* Activation of TLR3 by exogenous microbial ligands or endogenous injury-associated ligands leads to production of type I IFN.