Artificial Intelligence Based Computational Framework for Drug
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

PTK2B (Human) Recombinant Protein
PTK2B (Human) Recombinant PKB, PTK, PYK2, RAFTK Protein Gene Summary: This gene encodes a cytoplasmic protein tyrosine kinase which is involved in Catalog Number: P5800 calcium-induced regulation of ion channels and Regulation Status: For research use only (RUO) activation of the map kinase signaling pathway. The encoded protein may represent an important signaling Product Description: Human PTK2B (NP_775267.1, 1 intermediate between neuropeptide-activated receptors a.a. - 967 a.a.) full-length recombinant protein with GST or neurotransmitters that increase calcium flux and the tag expressed in Baculovirus infected Sf21 cells. downstream signals that regulate neuronal activity. The encoded protein undergoes rapid tyrosine Host: Insect phosphorylation and activation in response to increases in the intracellular calcium concentration, nicotinic Theoretical MW (kDa): 138 acetylcholine receptor activation, membrane depolarization, or protein kinase C activation. This Applications: Func, SDS-PAGE protein has been shown to bind CRK-associated (See our web site product page for detailed applications substrate, nephrocystin, GTPase regulator associated information) with FAK, and the SH2 domain of GRB2. The encoded protein is a member of the FAK subfamily of protein Protocols: See our web site at tyrosine kinases but lacks significant sequence similarity http://www.abnova.com/support/protocols.asp or product to kinases from other subfamilies. Four transcript page for detailed protocols variants encoding two different isoforms have been Form: Liquid found for this gene. [provided by RefSeq] Preparation Method: Baculovirus infected insect cell (Sf21) expression system Purification: Glutathione sepharose chromatography Purity: 74 % by SDS-PAGE/CBB staining Activity: The activity was measured by off-chip mobility shift assay. -

Supplementary Table S1. Control Capsules (CC) and Study Capsules (CR) Composition
Supplementary Table S1. Control Capsules (CC) and Study Capsules (CR) composition CC Capsules Composition mg Bioactive compound Sunflower oil 1.100 Gelatin 318,019 Glycerin 118,750 Clorophill E-140 1,923 CAPSULE TOTAL WEIGHT 1.538,692 CR Capsules Composition mg Bioactive compound Shark liver oil 20% 750 150 mg alkylglycerols alkylglycerols (Gustav Heess) Rosemary Antioxidant 11,25 mg Diterpene Phenols extract, 25 % Diterpene 45 Phenols, Type Nº 027.020 (Rosmarinus officinalis L.) Glyceril-monoestearate 30 Sunflower oil 263 Soy lecithin VEROLEC 56 12 Gelatin 318,019 Glycerin 118,750 Clorophill E-140 1,923 CAPSULE TOTAL WEIGHT 1.538,692 Supplementary Table S2. Table of the selected genes and pathways analyzed in the study Pathway Gen Gene name Inflammation, IL1B Interleukin 1, Beta Immunomodulation TNF (TNFA) Tumor Necrosis Factor MAPK1 Mitogen-Activated Protein Kinase 1 PTK2B Protein Tyrosine Kinase 2 Beta STAT3 Signal Transducer Activator Of Transcription 3 JAK1 Janus Kinase 1 JAK3 Janus Kinase 3 NFKB Nuclear Factor Of Kappa Light Polypeptide Gene Enhancer In B-Cells 1 NLRP3 NLR Family, Pyrin Domain Containing 3 CCL2 (MCP-1) Chemokine (C-C Motif) Ligand 2 CXCR1 Chemokine (C-X-C Motif) Receptor 1 CSF2 Colony Stimulating Factor 2 (Granulocyte-Macrophage) CCL5(RANTES) Chemokine (C-C Motif) Ligand 5 CCR5 Chemokine (C-C Motif) Receptor 5 (Gene/Pseudogene) PLCG1 Phospholipase C, Gamma 1 PRKCD Protein Kinase C, Delta ADIPOQ Adiponectin, C1Q And Collagen Domain Containing BMP2 Bone Morphogenetic Protein 2 LIF Leukemia Inhibitory Factor TGFB2 -

Genetic and Neurodevelopmental Spectrum Of
Cognitive and behavioural genetics J Med Genet: first published as 10.1136/jmedgenet-2015-103451 on 17 March 2016. Downloaded from ORIGINAL ARTICLE Genetic and neurodevelopmental spectrum of SYNGAP1-associated intellectual disability and epilepsy Cyril Mignot,1,2,3 Celina von Stülpnagel,4,5 Caroline Nava,1,6 Dorothée Ville,7 Damien Sanlaville,8,9,10 Gaetan Lesca,8,9,10 Agnès Rastetter,6 Benoit Gachet,6 Yannick Marie,6 G Christoph Korenke,11 Ingo Borggraefe,12 Dorota Hoffmann-Zacharska,13 Elżbieta Szczepanik,14 Mariola Rudzka-Dybała,14 Uluç Yiş,15 Hande Çağlayan,16 Arnaud Isapof,17 Isabelle Marey,1 Eleni Panagiotakaki,18 Christian Korff,19 Eva Rossier,20 Angelika Riess,21 Stefanie Beck-Woedl,21 Anita Rauch,22 Christiane Zweier,23 Juliane Hoyer,23 André Reis,23 Mikhail Mironov,24 Maria Bobylova,24 Konstantin Mukhin,24 Laura Hernandez-Hernandez,25 Bridget Maher,25 Sanjay Sisodiya,25 Marius Kuhn,26 Dieter Glaeser,26 Sarah Weckhuysen,6,27 Candace T Myers,28 Heather C Mefford,28 Konstanze Hörtnagel,29 Saskia Biskup,29 EuroEPINOMICS-RES MAE working group, Johannes R Lemke,30 Delphine Héron,1,2,3,4 Gerhard Kluger,4,5 Christel Depienne1,6 ▸ Additional material is ABSTRACT INTRODUCTION published online only. To view Objective We aimed to delineate the neurodevelopmental The human SYNGAP1 gene on chromosome please visit the journal online (http://dx.doi.org/10.1136/ spectrum associated with SYNGAP1 mutations and to 6p21.3 encodes the synaptic RAS-GTPase-activating jmedgenet-2015-103451). investigate genotype–phenotype correlations. protein 1, a protein of the post-synaptic density Methods We sequenced the exome or screened the exons (PSD) of glutamatergic neurons.12SYNGAP1 inter- For numbered affiliations see end of article. -

Supplementary Materials
Supplementary Suppl. Figure 1: MAPK signalling pathway of A: NCI-H2502, B: NCI-H2452, C: MSTO-211H and D: MRC-5. Suppl. Figure 2: Cell cycle pathway of A: NCI-H2502, B: NCI-H2452, C: MSTO-211H and D: MRC- 5. Suppl. Figure 3: Cancer pathways of A: NCI-H2502, B: NCI-H2452, C: MSTO-211H and D: MRC-5. Suppl. Figure 4: Phosphorylation level of A: ARAF, B: EPHA1, C: EPHA2, D: EPHA7 in all cell lines. For each cell line, phosphorylation levels are depicted before (Medium) and after cisplatin treatment (Cis). Suppl. Figure 5: Phosphorylation Level of A: KIT, B: PTPN11, C: PIK3R1, D: PTPN6 in all cell lines. For each cell line, phosphorylation levels are depicted before (Medium) and after cisplatin treatment (Cis). Suppl. Figure 6: Phosphorylation Level of A: KDR, B: EFS, C: AKT1, D: PTK2B/FAK2 in all cell lines. For each cell line, phosphorylation levels are depicted before (Medium) and after cisplatin treatment (Cis). Suppl. Figure 7: Scoreplots and volcanoplots of PTK upstream kinase analysis: A: Scoreplot of PTK- Upstream kinase analysis for NCI-H2052 cells. B: Volcanoplot of PTK-Upstream kinase analysis for NCI-H2052 cells. C: Scoreplot of PTK-Upstream kinase analysis for NCI-H2452 cells. D: Volcanoplot of PTK-Upstream kinase analysis for NCI-H2452 cells. E: Scoreplot of PTK-Upstream kinase analysis for MSTO-211H cells. F: Volcanoplot of PTK-Upstream kinase analysis for MSTO- 211H cells. G: Scoreplot of PTK-Upstream kinase analysis for MRC-5cells. H: Volcanoplot of PTK- Upstream kinase analysis for MRC-5 cells. Suppl. Figure 8: Scoreplots and volcanoplots of STK upstream kinase analysis: A: Scoreplot of STK- Upstream kinase analysis for NCI-H2052 cells. -

Blood Vitronectin Is a Major Activator of LIF and IL-6 in the Brain Through Integrin–FAK and Upar Signaling Matthew P
© 2018. Published by The Company of Biologists Ltd | Journal of Cell Science (2018) 131, jcs202580. doi:10.1242/jcs.202580 RESEARCH ARTICLE Blood vitronectin is a major activator of LIF and IL-6 in the brain through integrin–FAK and uPAR signaling Matthew P. Keasey1, Cuihong Jia1, Lylyan F. Pimentel1,2, Richard R. Sante1, Chiharu Lovins1 and Theo Hagg1,* ABSTRACT Microglia and astrocytes express the VTN receptors αvβ3 and α β We defined how blood-derived vitronectin (VTN) rapidly and potently v 5 integrin (Herrera-Molina et al., 2012; Kang et al., 2008; activates leukemia inhibitory factor (LIF) and pro-inflammatory Milner, 2009; Welser-Alves et al., 2011). Microglia and astrocytes, interleukin 6 (IL-6) in vitro and after vascular injury in the brain. as well as endothelial cells, are major producers of pro- α in vitro Treatment with VTN (but not fibrinogen, fibronectin, laminin-111 or inflammatory cytokines, such as IL-6 and TNF , and collagen-I) substantially increased LIF and IL-6 within 4 h in after traumatic or ischemic injury to the brain (Banner et al., 1997; C6-astroglioma cells, while VTN−/− mouse plasma was less effective Erta et al., 2012; Lau and Yu, 2001) or upon self-induction by IL-6 than that from wild-type mice. LIF and IL-6 were induced by (Van Wagoner and Benveniste, 1999). IL-6 is a major regulator of a intracerebral injection of recombinant human (rh)VTN in mice, but variety of inflammatory disorders and a target for therapies (Hunter induction seen upon intracerebral hemorrhage was less in VTN−/− and Jones, 2015). -

PRKACA Mediates Resistance to HER2-Targeted Therapy in Breast Cancer Cells and Restores Anti-Apoptotic Signaling
Oncogene (2015) 34, 2061–2071 © 2015 Macmillan Publishers Limited All rights reserved 0950-9232/15 www.nature.com/onc ORIGINAL ARTICLE PRKACA mediates resistance to HER2-targeted therapy in breast cancer cells and restores anti-apoptotic signaling SE Moody1,2,3, AC Schinzel1, S Singh1, F Izzo1, MR Strickland1, L Luo1,2, SR Thomas3, JS Boehm3, SY Kim4, ZC Wang5,6 and WC Hahn1,2,3 Targeting HER2 with antibodies or small molecule inhibitors in HER2-positive breast cancer leads to improved survival, but resistance is a common clinical problem. To uncover novel mechanisms of resistance to anti-HER2 therapy in breast cancer, we performed a kinase open reading frame screen to identify genes that rescue HER2-amplified breast cancer cells from HER2 inhibition or suppression. In addition to multiple members of the MAPK (mitogen-activated protein kinase) and PI3K (phosphoinositide 3-kinase) signaling pathways, we discovered that expression of the survival kinases PRKACA and PIM1 rescued cells from anti-HER2 therapy. Furthermore, we observed elevated PRKACA expression in trastuzumab-resistant breast cancer samples, indicating that this pathway is activated in breast cancers that are clinically resistant to trastuzumab-containing therapy. We found that neither PRKACA nor PIM1 restored MAPK or PI3K activation after lapatinib or trastuzumab treatment, but rather inactivated the pro-apoptotic protein BAD, the BCl-2-associated death promoter, thereby permitting survival signaling through BCL- XL. Pharmacological blockade of BCL-XL/BCL-2 partially abrogated the rescue effects conferred by PRKACA and PIM1, and sensitized cells to lapatinib treatment. These observations suggest that combined targeting of HER2 and the BCL-XL/BCL-2 anti-apoptotic pathway may increase responses to anti-HER2 therapy in breast cancer and decrease the emergence of resistant disease. -

Supplementary Table 1. in Vitro Side Effect Profiling Study for LDN/OSU-0212320. Neurotransmitter Related Steroids
Supplementary Table 1. In vitro side effect profiling study for LDN/OSU-0212320. Percent Inhibition Receptor 10 µM Neurotransmitter Related Adenosine, Non-selective 7.29% Adrenergic, Alpha 1, Non-selective 24.98% Adrenergic, Alpha 2, Non-selective 27.18% Adrenergic, Beta, Non-selective -20.94% Dopamine Transporter 8.69% Dopamine, D1 (h) 8.48% Dopamine, D2s (h) 4.06% GABA A, Agonist Site -16.15% GABA A, BDZ, alpha 1 site 12.73% GABA-B 13.60% Glutamate, AMPA Site (Ionotropic) 12.06% Glutamate, Kainate Site (Ionotropic) -1.03% Glutamate, NMDA Agonist Site (Ionotropic) 0.12% Glutamate, NMDA, Glycine (Stry-insens Site) 9.84% (Ionotropic) Glycine, Strychnine-sensitive 0.99% Histamine, H1 -5.54% Histamine, H2 16.54% Histamine, H3 4.80% Melatonin, Non-selective -5.54% Muscarinic, M1 (hr) -1.88% Muscarinic, M2 (h) 0.82% Muscarinic, Non-selective, Central 29.04% Muscarinic, Non-selective, Peripheral 0.29% Nicotinic, Neuronal (-BnTx insensitive) 7.85% Norepinephrine Transporter 2.87% Opioid, Non-selective -0.09% Opioid, Orphanin, ORL1 (h) 11.55% Serotonin Transporter -3.02% Serotonin, Non-selective 26.33% Sigma, Non-Selective 10.19% Steroids Estrogen 11.16% 1 Percent Inhibition Receptor 10 µM Testosterone (cytosolic) (h) 12.50% Ion Channels Calcium Channel, Type L (Dihydropyridine Site) 43.18% Calcium Channel, Type N 4.15% Potassium Channel, ATP-Sensitive -4.05% Potassium Channel, Ca2+ Act., VI 17.80% Potassium Channel, I(Kr) (hERG) (h) -6.44% Sodium, Site 2 -0.39% Second Messengers Nitric Oxide, NOS (Neuronal-Binding) -17.09% Prostaglandins Leukotriene, -

Gene Expression Profiling of Chronic Myeloid Leukemia with Variant T(9;22)
Albano et al. Molecular Cancer 2013, 12:36 http://www.molecular-cancer.com/content/12/1/36 SHORT COMMUNICATION Open Access Gene expression profiling of chronic myeloid leukemia with variant t(9;22) reveals a different signature from cases with classic translocation Francesco Albano*, Antonella Zagaria, Luisa Anelli, Nicoletta Coccaro, Luciana Impera, Crescenzio Francesco Minervini, Angela Minervini, Antonella Russo Rossi, Giuseppina Tota, Paola Casieri and Giorgina Specchia Abstract Background: The t(9;22)(q34;q11) generating the BCR/ABL1 fusion gene represents the cytogenetic hallmark of chronic myeloid leukemia (CML). About 5–10% of CML cases show variant translocations with the involvement of other chromosomes in addition to chromosomes 9 and 22. The molecular bases of biological differences between CML patients with classic and variant t(9;22) have never been clarified. Findings: In this study, we performed gene expression microarray analysis to compare CML patients bearing variant rearrangements and those with classic t(9;22)(q34;q11). We identified 59 differentially expressed genes significantly associated with the two analyzed groups. The role of specific candidate genes such as TRIB1 (tribbles homolog 1), PTK2B (protein tyrosine kinase 2 beta), and C5AR1 (complement component 5a receptor 1) is discussed. Conclusions: Our results reveal that in CML cases with variant t(9;22) there is an enhancement of the MAPK pathway deregulation and show that kinases are a common target of molecular alterations in hematological disorders. Keywords: Chronic myeloid leukemia, Variant t(9;22) rearrangements, Gene expression profiling, Protein kinases, Cellular pathways Background BCR has been reported in CML cases with variant trans- Chronic myeloid leukemia (CML) is a myeloprolife- locations with a greater frequency (30-40%) than in rative disorder derived from hematopoietic stem cell cases with classic t(9;22) (10-18%) [6,7]. -
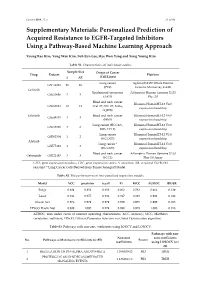
Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach
Cancers 2019, 11, x S1 of S9 Supplementary Materials: Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach Young Rae Kim, Yong Wan Kim, Suh Eun Lee, Hye Won Yang and Sung Young Kim Table S1. Characteristics of individual studies. Sample Size Origin of Cancer Drug Dataset Platform S AR (Cell Lines) Lung cancer Agilent-014850 Whole Human GSE34228 26 26 (PC9) Genome Microarray 4x44K Gefitinib Epidermoid carcinoma Affymetrix Human Genome U133 GSE10696 3 3 (A431) Plus 2.0 Head and neck cancer Illumina HumanHT-12 V4.0 GSE62061 12 12 (Cal-27, SSC-25, FaDu, expression beadchip SQ20B) Erlotinib Head and neck cancer Illumina HumanHT-12 V4.0 GSE49135 3 3 (HN5) expression beadchip Lung cancer (HCC827, Illumina HumanHT-12 V3.0 GSE38310 3 6 ER3, T15-2) expression beadchip Lung cancer Illumina HumanHT-12 V3.0 GSE62504 1 2 (HCC827) expression beadchip Afatinib Lung cancer * Illumina HumanHT-12 V4.0 GSE75468 1 3 (HCC827) expression beadchip Head and neck cancer Affymetrix Human Genome U133 Cetuximab GSE21483 3 3 (SCC1) Plus 2.0 Array GEO, gene expression omnibus; GSE, gene expression series; S, sensitive; AR, acquired EGFR-TKI resistant; * Lung Cancer Cells Derived from Tumor Xenograft Model. Table S2. The performances of four penalized regression models. Model ACC precision recall F1 MCC AUROC BRIER Ridge 0.889 0.852 0.958 0.902 0.782 0.964 0.129 Lasso 0.944 0.957 0.938 0.947 0.889 0.991 0.042 Elastic Net 0.978 0.979 0.979 0.979 0.955 0.999 0.023 EPSGO Elastic Net 0.989 1.000 0.979 0.989 0.978 1.000 0.018 AUROC, area under curve of receiver operating characteristic; ACC, accuracy; MCC, Matthews correlation coefficient; EPSGO, Efficient Parameter Selection via Global Optimization algorithm. -
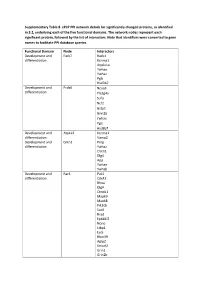
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Artificial Intelligence-Based Computational Framework for Drug
Tsuji et al. Alzheimer’s Research & Therapy (2021) 13:92 https://doi.org/10.1186/s13195-021-00826-3 RESEARCH Open Access Artificial intelligence-based computational framework for drug-target prioritization and inference of novel repositionable drugs for Alzheimer’s disease Shingo Tsuji1*† , Takeshi Hase2,3,4,5†, Ayako Yachie-Kinoshita2,4, Taiko Nishino2,SamikGhosh2, Masataka Kikuchi6, Kazuro Shimokawa7, Hiroyuki Aburatani1, Hiroaki Kitano2 and Hiroshi Tanaka3 Abstract Background: Identifying novel therapeutic targets is crucial for the successful development of drugs. However, the cost to experimentally identify therapeutic targets is huge and only approximately 400 genes are targets for FDA-approved drugs. As a result, it is inevitable to develop powerful computational tools that can identify potential novel therapeutic targets. Fortunately, the human protein-protein interaction network (PIN) could be a useful resource to achieve this objective. Methods: In this study, we developed a deep learning-based computational framework that extracts low-dimensional representations of high-dimensional PIN data. Our computational framework uses latent features and state-of-the-art machine learning techniques to infer potential drug target genes. Results: We applied our computational framework to prioritize novel putative target genes for Alzheimer’s disease and successfully identified key genes that may serve as novel therapeutic targets (e.g., DLG4, EGFR, RAC1, SYK, PTK2B, SOCS1). Furthermore, based on these putative targets, we could infer repositionable -

47421 SAP102 (A7R8L) Rabbit Mab
Revision 1 C 0 2 - t SAP102 (A7R8L) Rabbit mAb a e r o t S Orders: 877-616-CELL (2355) [email protected] 1 Support: 877-678-TECH (8324) 2 4 Web: [email protected] 7 www.cellsignal.com 4 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source/Isotype: UniProt ID: Entrez-Gene Id: WB, IP H M R Endogenous 102 Rabbit Q92796 1741 Product Usage Information 1. Cuthbert, P.C. et al. (2007) J Neurosci 27, 2673-82. 2. Makino, K. et al. (1997) Oncogene 14, 2425-33. Application Dilution 3. Masuko, N. et al. (1999) J Biol Chem 274, 5782-90. 4. Lau, L.F. et al. (1996) J Biol Chem 271, 21622-8. Western Blotting 1:1000 5. Yan, J. et al. (2005) Mol Psychiatry 10, 329-32. Immunoprecipitation 1:50 Storage Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA, 50% glycerol and less than 0.02% sodium azide. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity SAP102 (A7R8L) Rabbit mAb recognizes endogenous levels of total SAP102 protein. Species Reactivity: Human, Mouse, Rat Source / Purification Monoclonal antibody is produced by immunizing animals with a synthetic peptide corresponding to residues near the carboxy terminus of human SAP102 protein. Background Synapse-associated protein 102 (SAP102, DLG3) belongs to the membrane-associated guanylate kinase (MAGUK) protein family and is a homolog of the Drosophila disc large (dlg) tumor suppressor protein. SAP102 consists of three PDZ domains, a Src homology 3 (SH3) domain, and a guanylate kinase (GK) domain (1).