Downloaded from the Projected by No Less Than Six of the Servers Mentioned Above
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Seq2pathway Vignette
seq2pathway Vignette Bin Wang, Xinan Holly Yang, Arjun Kinstlick May 19, 2021 Contents 1 Abstract 1 2 Package Installation 2 3 runseq2pathway 2 4 Two main functions 3 4.1 seq2gene . .3 4.1.1 seq2gene flowchart . .3 4.1.2 runseq2gene inputs/parameters . .5 4.1.3 runseq2gene outputs . .8 4.2 gene2pathway . 10 4.2.1 gene2pathway flowchart . 11 4.2.2 gene2pathway test inputs/parameters . 11 4.2.3 gene2pathway test outputs . 12 5 Examples 13 5.1 ChIP-seq data analysis . 13 5.1.1 Map ChIP-seq enriched peaks to genes using runseq2gene .................... 13 5.1.2 Discover enriched GO terms using gene2pathway_test with gene scores . 15 5.1.3 Discover enriched GO terms using Fisher's Exact test without gene scores . 17 5.1.4 Add description for genes . 20 5.2 RNA-seq data analysis . 20 6 R environment session 23 1 Abstract Seq2pathway is a novel computational tool to analyze functional gene-sets (including signaling pathways) using variable next-generation sequencing data[1]. Integral to this tool are the \seq2gene" and \gene2pathway" components in series that infer a quantitative pathway-level profile for each sample. The seq2gene function assigns phenotype-associated significance of genomic regions to gene-level scores, where the significance could be p-values of SNPs or point mutations, protein-binding affinity, or transcriptional expression level. The seq2gene function has the feasibility to assign non-exon regions to a range of neighboring genes besides the nearest one, thus facilitating the study of functional non-coding elements[2]. Then the gene2pathway summarizes gene-level measurements to pathway-level scores, comparing the quantity of significance for gene members within a pathway with those outside a pathway. -

Transcriptomic Analysis of the Aquaporin (AQP) Gene Family
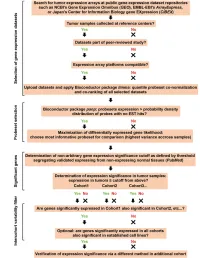
Pancreatology 19 (2019) 436e442 Contents lists available at ScienceDirect Pancreatology journal homepage: www.elsevier.com/locate/pan Transcriptomic analysis of the Aquaporin (AQP) gene family interactome identifies a molecular panel of four prognostic markers in patients with pancreatic ductal adenocarcinoma Dimitrios E. Magouliotis a, b, Vasiliki S. Tasiopoulou c, Konstantinos Dimas d, * Nikos Sakellaridis d, Konstantina A. Svokos e, Alexis A. Svokos f, Dimitris Zacharoulis b, a Division of Surgery and Interventional Science, Faculty of Medical Sciences, UCL, London, UK b Department of Surgery, University of Thessaly, Biopolis, Larissa, Greece c Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece d Department of Pharmacology, Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece e The Warren Alpert Medical School of Brown University, Providence, RI, USA f Riverside Regional Medical Center, Newport News, VA, USA article info abstract Article history: Background: This study aimed to assess the differential gene expression of aquaporin (AQP) gene family Received 14 October 2018 interactome in pancreatic ductal adenocarcinoma (PDAC) using data mining techniques to identify novel Received in revised form candidate genes intervening in the pathogenicity of PDAC. 29 January 2019 Method: Transcriptome data mining techniques were used in order to construct the interactome of the Accepted 9 February 2019 AQP gene family and to determine which genes members are differentially expressed in PDAC as Available online 11 February 2019 compared to controls. The same techniques were used in order to evaluate the potential prognostic role of the differentially expressed genes. Keywords: PDAC Results: Transcriptome microarray data of four GEO datasets were incorporated, including 142 primary Aquaporin tumor samples and 104 normal pancreatic tissue samples. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Mapping Transmembrane Binding Partners for E-Cadherin Ectodomains
SUPPLEMENTARY INFORMATION TITLE: Mapping transmembrane binding partners for E-cadherin ectodomains. AUTHORS: Omer Shafraz 1, Bin Xie 2, Soichiro Yamada 1, Sanjeevi Sivasankar 1, 2, * AFFILIATION: 1 Department of Biomedical Engineering, 2 Biophysics Graduate Group, University of California, Davis, CA 95616. *CORRESPONDING AUTHOR: Sanjeevi Sivasankar, Tel: (530)-754-0840, Email: [email protected] Figure S1: Western blots a. EC-BioID, WT and Ecad-KO cell lysates stained for Ecad and tubulin. b. HRP-streptavidin staining of biotinylated proteins eluted from streptavidin coated magnetic beads incubated with cell lysates of EC-BioID with (+) and without (-) exogenous biotin. c. C-BioID, WT and Ecad-KO cell lysates stained for Ecad and tubulin. d. HRP-streptavidin staining of biotinylated proteins eluted from streptavidin coated magnetic beads incubated with cell lysates of C-BioID with (+) and without (-) exogenous biotin. (+) Biotin (-) Biotin Sample 1 Sample 2 Sample 3 Sample 4 Sample 1 Sample 2 Sample 3 Sample 4 Percent Percent Percent Percent Percent Percent Percent Percent Gene ID Coverage Coverage Coverage Coverage Coverage Coverage Coverage Coverage CDH1 29.6 31.4 41.1 36.5 10.8 6.7 28.8 29.1 DSG2 26 14.6 45 37 0.8 1.9 1.6 18.7 CXADR 30.2 26.2 32.7 27.1 0.0 0.0 0.0 6.9 EFNB1 24.3 30.6 24 30.3 0.0 0.0 0.0 0.0 ITGA2 16.5 22.2 30.1 33.4 1.1 1.1 5.2 7.2 CDH3 21.8 9.7 20.6 25.3 1.3 1.3 0.0 0.0 ITGB1 11.8 16.7 23.9 20.3 0.0 2.9 8.5 5.8 DSC3 9.7 7.5 11.5 13.3 0.0 0.0 2.6 0.0 EPHA2 23.2 31.6 31.6 30.5 0.8 0.0 0.0 5.7 ITGB4 21.8 27.8 33.1 30.7 0.0 1.2 3.9 4.4 ITGB3 23.5 22.2 26.8 24.7 0.0 0.0 5.2 9.1 CDH6 22.8 18.1 28.6 24.3 0.0 0.0 0.0 9.1 CDH17 8.8 12.4 20.7 18.4 0.0 0.0 0.0 0.0 ITGB6 12.7 10.4 14 17.1 0.0 0.0 0.0 1.7 EPHB4 11.4 8.1 14.2 16.3 0.0 0.0 0.0 0.0 ITGB8 5 10 15 17.6 0.0 0.0 0.0 0.0 ITGB5 6.2 9.5 15.2 13.8 0.0 0.0 0.0 0.0 EPHB2 8.5 4.8 9.8 12.1 0.0 0.0 0.0 0.0 CDH24 5.9 7.2 8.3 9 0.0 0.0 0.0 0.0 Table S1: EC-BioID transmembrane protein hits. -

Metabolic and Inflammatory Proteins Differentially Expressed in Platelets
ics om & B te i ro o P in f f o o r l m a Journal of a n t r i Flores-Nascimento et al., J Proteomics Bioinform 2014, 7:1 c u s o J DOI: 10.4172/jpb.1000298 ISSN: 0974-276X Proteomics & Bioinformatics Research Article Article OpenOpen Access Access Metabolic and Inflammatory Proteins Differentially Expressed in Platelets from Unprovoked Deep Vein Thrombosis Patients Mariane C Flores-Nascimento1*, Karina Kleinfelder-Fontanesi2, Thiago Matos de Araújo3, Gabriel Forato Anhê3, Rodrigo Secolin4, Fernanda Andrade Orsi1, Erich Vinicius de Paula1 and Joyce M Annichino-Bizzacchi1 1Hematology-Hemotherapy Center, University of Campinas (UNICAMP), Campinas, SP, Brazil 2Biology Institute, University of Campinas (UNICAMP), Campinas, SP, Brazil 3Department of Pharmacology, University of Campinas (UNICAMP), Campinas, SP, Brazil 4Medical Genetic Department, University of Campinas (UNICAMP), Campinas, SP, Brazil Abstract Deep vein thrombosis (DVT) is multi-causal disease associated to high morbidity and mortality due to complications, especially from pulmonary embolism. New factors of relevance on the DVT pathophysiology enhance our understanding of disease mechanisms and can be translated into improved management of patients, prevention of recurrence and development of new therapies. In this context, the precise role of platelets in the pathogenesis of DVT is not completely understood. Our objectives were to acquire and to analyze the whole platelet protein profile of samples from 3 DVT patients and to compare them to results obtained from 1 sibling and 1 neighbor from each patient (in order to minimize genetic and environmental interferences). These patients presented unprovoked and recurrent episodes of proximal DVT as well as a family history of DVT. -

Human Ipsc-Derived Hypertrophic Chondrocytes Reveal a Mutation-Specific Unfolded Protein Response in Chondrodysplasias
bioRxiv preprint doi: https://doi.org/10.1101/2020.05.19.103960; this version posted May 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Human iPSC-derived hypertrophic chondrocytes reveal a mutation-specific unfolded protein response in chondrodysplasias Yann Pretemer1, Shunsuke Kawai1,2,3, Makoto Watanabe1,4, Sanae Nagata1, Megumi Nishio2, Sakura Tamaki2,5, Cantas Alev6, Jing-Yi Xue7, Zheng Wang7,8, Kenichi Fukiage9,10, Masako Tsukanaka9, Tohru Futami9, Shiro Ikegawa7, Junya Toguchida1,2,3,5* 1Department of Cell Growth and Differentiation, Center for iPS Cell Research and Application, Kyoto University, Kyoto, Japan. 2Institute for Frontier Life and Medical Sciences, Kyoto University, Kyoto, Japan. 3Department of Orthopaedic Surgery, Graduate School of Medicine, Kyoto University, Kyoto, Japan. 4Life Science Research Center, Technology Research Laboratory, Shimadzu Corporation, Kyoto, Japan. 5Institute for Advancement of Clinical and Translational Sciences, Kyoto University Hospital, Kyoto University, Kyoto, Japan. 6Institute for the Advanced Study of Human Biology, Kyoto University, Kyoto, Japan. 7Laboratory for Bone and Joint Diseases, RIKEN Center for Integrative Medical Sciences, Tokyo, Japan. 8McKusick-Zhang Center for Genetic Medicine and State Key Laboratory of Medical Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, China. 9Department of Pediatric Orthopaedics, Shiga Medical Center for Children, Moriyama, Japan. 10Department of Orthopaedic Surgery, Bobath Memorial Hospital, Osaka, Japan. *e-mail: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.05.19.103960; this version posted May 19, 2020. -

Supplemental Table S-2. Genes That Were Differentially Regulated by 2-Fold Or Greater Between Male and Female Tendons
Supplemental Table S-2. Genes that were differentially regulated by 2-fold or greater between male and female tendons. Log Fold Affymetrix Change Male vs Probe ID Entrez ID Symbol Description Female 17543785 213742 Xist inactive X specific transcripts -5.47 17426022 100041687 Mup8 major urinary protein 8 -3.73 17426062 100048885 Gm2083 major urinary protein LOC100048885 -3.53 17426043 100048885 Gm2083 major urinary protein LOC100048885 -3.50 17297591 NA NA NA -3.50 17426011 100048885 Gm2083 major urinary protein LOC100048885 -3.46 17425990 100048885 Gm2083 major urinary protein LOC100048885 -3.46 17425915 100048885 Gm2083 major urinary protein LOC100048885 -3.46 17425941 100048885 Gm2083 major urinary protein LOC100048885 -3.40 17517360 66402 Sln sarcolipin -3.18 17426032 100048885 Gm2083 major urinary protein LOC100048885 -3.12 17346069 240120 Zfp119b zinc finger protein 119b -3.07 17426000 100048885 Gm2083 major urinary protein LOC100048885 -3.03 17425900 100048885 Gm2083 major urinary protein LOC100048885 -2.91 17425888 100048885 Gm2083 major urinary protein LOC100048885 -2.85 17315002 70152 Mettl7a1 methyltransferase like 7A1 -2.62 17248348 NA NA NA -2.41 17524209 NA NA NA -2.39 17443890 24135 Zfp68 zinc finger protein 68 -2.38 17238834 NA NA NA -2.33 17343251 108121 U2af1 U2 small nuclear ribonucleoprotein auxiliary factor (U2AF) 1 -2.32 17406421 99889 Arfip1 ADP-ribosylation factor interacting protein 1 -2.30 17497612 93747 Echs1 enoyl Coenzyme A hydratase, short chain, 1, mitochondrial -2.22 17238892 NA NA NA -2.22 17238934 64009 -

Murine Perinatal Beta Cell Proliferation and the Differentiation of Human Stem Cell Derived Insulin Expressing Cells Require NEUROD1
Page 1 of 105 Diabetes Murine perinatal beta cell proliferation and the differentiation of human stem cell derived insulin expressing cells require NEUROD1 Anthony I. Romer,1,2 Ruth A. Singer1,3, Lina Sui2, Dieter Egli,2* and Lori Sussel1,4* 1Department of Genetics and Development, Columbia University, New York, NY 10032, USA 2Department of Pediatrics, Columbia University, New York, NY 10032, USA 3Integrated Program in Cellular, Molecular and Biomedical Studies, Columbia University, New York, NY 10032, USA 4Department of Pediatrics, University of Colorado Denver School of Medicine, Denver, CO 80045, USA *Co-Corresponding Authors Dieter Egli 1150 St. Nicholas Avenue New York, NY 10032 [email protected] Lori Sussel 1775 Aurora Ct. Aurora, CO 80045 [email protected] Word Count: Abstract= 149; Body= 4773 Total Paper Figures= 7, Total Supplemental Tables= 4, Total Supplemental Figures= 5 Diabetes Publish Ahead of Print, published online September 13, 2019 Diabetes Page 2 of 105 Abstract Inactivation of the β cell transcription factor NEUROD1 causes diabetes in mice and humans. In this study, we uncovered novel functions of Neurod1 during murine islet cell development and during the differentiation of human embryonic stem cells (HESCs) into insulin-producing cells. In mice, we determined that Neurod1 is required for perinatal proliferation of alpha and beta cells. Surprisingly, apoptosis only makes a minor contribution to beta cell loss when Neurod1 is deleted. Inactivation of NEUROD1 in HESCs severely impaired their differentiation from pancreatic progenitors into insulin expressing (HESC-beta) cells; however survival or proliferation was not affected at the time points analyzed. NEUROD1 was also required in HESC-beta cells for the full activation of an essential beta cell transcription factor network. -
Supplementary Data
SUPPLEMENTAL INFORMATION A study restricted to chemokine receptors as well as a genome-wide transcript analysis uncovered CXCR4 as preferentially expressed in Ewing's sarcoma (Ewing's sarcoma) cells of metastatic origin (Figure 4). Transcriptome analyses showed that in addition to CXCR4, genes known to support cell motility and invasion topped the list of genes preferentially expressed in metastasis-derived cells (Figure 4D). These included kynurenine 3-monooxygenase (KMO), galectin-1 (LGALS1), gastrin-releasing peptide (GRP), procollagen C-endopeptidase enhancer (PCOLCE), and ephrin receptor B (EPHB3). KMO, a key enzyme of tryptophan catabolism, has not been linked to metastasis. Tryptophan and its catabolites, however, are involved in immune evasion by tumors, a process that can assist in tumor progression and metastasis (1). LGALS1, GRP, PCOLCE and EPHB3 have been linked to tumor progression and metastasis of several cancers (2-4). Top genes preferentially expressed in L-EDCL included genes that suppress cell motility and/or potentiate cell adhesion such as plakophilin 1 (PKP1), neuropeptide Y (NPY), or the metastasis suppressor TXNIP (5-7) (Figure 4D). Overall, L-EDCL were enriched in gene sets geared at optimizing nutrient transport and usage (Figure 4D; Supplementary Table 3), a state that may support the early stages of tumor growth. Once tumor growth outpaces nutrient and oxygen supplies, gene expression programs are usually switched to hypoxic response and neoangiogenesis, which ultimately lead to tumor egress and metastasis. Accordingly, gene sets involved in extracellular matrix remodeling, MAPK signaling, and response to hypoxia were up-regulated in M-EDCL (Figure 4D; Supplementary Table 4), consistent with their association to metastasis in other cancers (8, 9). -

ZNF384-Related Fusion Genes Define a Subgroup of Childhood B-Cell
Acute Lymphoblastic Leukemia SUPPLEMENTARY APPENDIX ZNF384 -related fusion genes define a subgroup of childhood B-cell precursor acute lymphoblastic leukemia with a characteristic immunotype Shinsuke Hirabayashi, 1,2 Kentaro Ohki, 1 Kazuhiko Nakabayashi, 3 Hitoshi Ichikawa, 4 Yukihide Momozawa, 5 Kohji Oka - mura, 6 Akinori Yaguchi, 1,7 Kazuki Terada, 1 Yuya Saito, 1,8 Ai Yoshimi, 1,9 Hiroko Ogata-Kawata, 3 Hiromi Sakamoto, 4 Moto - hiro Kato, 1,10 Junya Fujimura, 7 Moeko Hino, 11 Akitoshi Kinoshita, 12 Harumi Kakuda, 13 Hidemitsu Kurosawa, 14 Keisuke Kato, 9 Ryosuke Kajiwara, 15 Koichi Moriwaki, 16 Tsuyoshi Morimoto, 17 Kozue Nakamura, 18 Yasushi Noguchi, 19 Tomoo Osumi, 1,20 Kazuo Sakashita, 21 Junko Takita, 22 Yuki Yuza, 8 Koich Matsuda, 23 Teruhiko Yoshida, 4 Kenji Matsumoto, 24 Kenichiro Hata, 3 Michiaki Kubo, 5 Yoichi Matsubara, 25 Takashi Fukushima, 26 Katsuyoshi Koh, 27 Atsushi Manabe, 2 Akira Ohara 28 and Nobutaka Kiyokawa 1 for the Tokyo Children’s Cancer Study Group (TCCSG) 1Department of Pediatric Hematology and Oncology Research, National Research Institute for Child Health and Development, Seta - gaya-ku, Tokyo; 2Department of Pediatrics, St. Luke's International Hospital, Chuo-ku, Tokyo; 3Department of Maternal-Fetal Biology, National Research Institute for Child Health and Development, Setagaya-ku, Tokyo; 4Division of Genetics, National Cancer Center Re - search Institute, Chuo-ku, Tokyo; 5Laboratory for Genotyping Development, Center for Integrative Medical Sciences (IMS), RIKEN, Yoko - hama-shi, Kanagawa; 6Department -

Supporting Information
Supporting Information Shtutman et al. 10.1073/pnas.1103842108 SI Methods chronic myelogenous leukemia; and CCRF-CEM, acute lympho- Cell Lines. Tumor cell lines were obtained from American Type blastic leukemia. cDNA was prepared with the SMART cDNA Culture Collection. BJ normal foreskin fibroblasts immortalized synthesis kit (Clontech Laboratories). Normalization of the cDNA with human telomerase reverse (line BJ1-hTERT) were obtained mixtures was carried out by Evrogen using duplex-specific nuclease from Clontech Laboratories, BJ-ELN, BJ-ELB, and BJ-ELR cells (8, 9). Normalized cDNA mixture was fragmented as described (1) were kindly provided by W. Hahn (Massachusetts General (10), ligated with adapters generated by annealing one long and one Hospital, Boston, MA). Human prostate epithelial cells were short strand containing AgeI or SphI restriction sites, and then was fi fi obtained from LifeLine Cell Technologies. MCF7 cells were PCR-ampli ed using the following adapter-speci c PCR primers: maintained in DMEM supplemented with 0.1 mmol/L of non- AgeI long adapter: AAACGTGGAGAATAAGCCGAATC- essential amino acids, 1 mmol/L of sodium pyruvate, 0.01 mg/mL CACCGGTACAATGGATGGATGG of bovine insulin (Sigma), and 10% FBS (HyClone) or 10% FC2 AgeI short adapter: CCATCCATCCATTGTACCGGTG (Invitrogen). MDA-MB231, HeLa, SW480, HCT116, HT1080, SphI long adapter: AAATCCTAAACCGACCGAGTATC- ACHN, K562, SK-MEL-28, SK-MEL-103, SK-MEL-19, SK-MEL- CACGAGCATGCCTAACTAACTA 29, SK-ML-147, T24, and U251 cell lines were grown in DMEM SphI short adapter: TAGTTAGTTAGGCATGCTCGTG with 10% FC2. HL60 cells were grown in DMEM with 20% FBS, AgeI amplified primer: CGTGGAGAATAAGCCGAATC and CCRF-CEM cells were grown in RPMI with 10% FBS. -

COPZ1 Antibody (N-Term) Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP13127A
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 COPZ1 Antibody (N-term) Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP13127A Specification COPZ1 Antibody (N-term) - Product Information Application WB,E Primary Accession P61923 Other Accession P61924, P35604, NP_057141.1 Reactivity Human, Mouse Predicted Bovine Host Rabbit Clonality Polyclonal Isotype Rabbit Ig Calculated MW 20198 Antigen Region 24-53 COPZ1 Antibody (N-term) - Additional Information COPZ1 Antibody (N-term) (Cat. #AP13127a) western blot analysis in L929 cell line lysates (35ug/lane).This demonstrates the COPZ1 Gene ID 22818 antibody detected the COPZ1 protein (arrow). Other Names Coatomer subunit zeta-1, Zeta-1-coat protein, Zeta-1 COP, COPZ1, COPZ Target/Specificity This COPZ1 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 24-53 amino acids from the N-terminal region of human COPZ1. Dilution WB~~1:1000 Format Purified polyclonal antibody supplied in PBS COPZ1 Antibody (N-term) (Cat. #AP13127a) with 0.09% (W/V) sodium azide. This western blot analysis in mouse brain tissue antibody is purified through a protein A lysates (35ug/lane).This demonstrates the column, followed by peptide affinity COPZ1 antibody detected the COPZ1 protein purification. (arrow). Storage Maintain refrigerated at 2-8°C for up to 2 COPZ1 Antibody (N-term) - Background weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw The coatomer is a cytosolic protein complex cycles. that binds to dilysine motifs and reversibly associates with Golgi non-clathrin-coated Precautions vesicles, which further mediate biosynthetic Page 1/2 10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 COPZ1 Antibody (N-term) is for research protein transport from the ER, via the Golgi up use only and not for use in diagnostic or to the trans Golgi network.