Human Ipsc-Derived Hypertrophic Chondrocytes Reveal a Mutation-Specific Unfolded Protein Response in Chondrodysplasias
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Transcriptome Response of High- and Low-Light-Adapted Prochlorococcus Strains to Changing Iron Availability
Transcriptome response of high- and low-light-adapted Prochlorococcus strains to changing iron availability The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Thompson, Anne W et al. “Transcriptome Response of High- and Low-light-adapted Prochlorococcus Strains to Changing Iron Availability.” ISME Journal (2011), 1-15. As Published http://dx.doi.org/10.1038/ismej.2011.49 Publisher Nature Publishing Group Version Author's final manuscript Citable link http://hdl.handle.net/1721.1/64705 Terms of Use Creative Commons Attribution-Noncommercial-Share Alike 3.0 Detailed Terms http://creativecommons.org/licenses/by-nc-sa/3.0/ Title: Transcriptome response of high- and low-light adapted Prochlorococcus strains to changing iron availability Running title: Prochlorococcus response to iron stress 5 Contributors: Anne W. Thompson1, Katherine Huang1, Mak A. Saito* 2, Sallie W. Chisholm* 1, 3 10 1 MIT Department of Civil and Environmental Engineering 2 Woods Hole Oceanographic Institution – Department of Marine Chemistry and Geochemistry 3 MIT Department of Biology 15 * To whom correspondence should be addressed: E-mail: [email protected] and [email protected] Subject Category: Microbial population and community ecology 20 Abstract Prochlorococcus contributes significantly to ocean primary productivity. The link between primary productivity and iron in specific ocean regions is well established and iron-limitation of Prochlorococcus cell division rates in these regions has been 25 demonstrated. However, the extent of ecotypic variation in iron metabolism among Prochlorococcus and the molecular basis for differences is not understood. Here, we examine the growth and transcriptional response of Prochlorococcus strains, MED4 and MIT9313, to changing iron concentrations. -

Seq2pathway Vignette
seq2pathway Vignette Bin Wang, Xinan Holly Yang, Arjun Kinstlick May 19, 2021 Contents 1 Abstract 1 2 Package Installation 2 3 runseq2pathway 2 4 Two main functions 3 4.1 seq2gene . .3 4.1.1 seq2gene flowchart . .3 4.1.2 runseq2gene inputs/parameters . .5 4.1.3 runseq2gene outputs . .8 4.2 gene2pathway . 10 4.2.1 gene2pathway flowchart . 11 4.2.2 gene2pathway test inputs/parameters . 11 4.2.3 gene2pathway test outputs . 12 5 Examples 13 5.1 ChIP-seq data analysis . 13 5.1.1 Map ChIP-seq enriched peaks to genes using runseq2gene .................... 13 5.1.2 Discover enriched GO terms using gene2pathway_test with gene scores . 15 5.1.3 Discover enriched GO terms using Fisher's Exact test without gene scores . 17 5.1.4 Add description for genes . 20 5.2 RNA-seq data analysis . 20 6 R environment session 23 1 Abstract Seq2pathway is a novel computational tool to analyze functional gene-sets (including signaling pathways) using variable next-generation sequencing data[1]. Integral to this tool are the \seq2gene" and \gene2pathway" components in series that infer a quantitative pathway-level profile for each sample. The seq2gene function assigns phenotype-associated significance of genomic regions to gene-level scores, where the significance could be p-values of SNPs or point mutations, protein-binding affinity, or transcriptional expression level. The seq2gene function has the feasibility to assign non-exon regions to a range of neighboring genes besides the nearest one, thus facilitating the study of functional non-coding elements[2]. Then the gene2pathway summarizes gene-level measurements to pathway-level scores, comparing the quantity of significance for gene members within a pathway with those outside a pathway. -

Transcriptomic Analysis of the Aquaporin (AQP) Gene Family
Pancreatology 19 (2019) 436e442 Contents lists available at ScienceDirect Pancreatology journal homepage: www.elsevier.com/locate/pan Transcriptomic analysis of the Aquaporin (AQP) gene family interactome identifies a molecular panel of four prognostic markers in patients with pancreatic ductal adenocarcinoma Dimitrios E. Magouliotis a, b, Vasiliki S. Tasiopoulou c, Konstantinos Dimas d, * Nikos Sakellaridis d, Konstantina A. Svokos e, Alexis A. Svokos f, Dimitris Zacharoulis b, a Division of Surgery and Interventional Science, Faculty of Medical Sciences, UCL, London, UK b Department of Surgery, University of Thessaly, Biopolis, Larissa, Greece c Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece d Department of Pharmacology, Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece e The Warren Alpert Medical School of Brown University, Providence, RI, USA f Riverside Regional Medical Center, Newport News, VA, USA article info abstract Article history: Background: This study aimed to assess the differential gene expression of aquaporin (AQP) gene family Received 14 October 2018 interactome in pancreatic ductal adenocarcinoma (PDAC) using data mining techniques to identify novel Received in revised form candidate genes intervening in the pathogenicity of PDAC. 29 January 2019 Method: Transcriptome data mining techniques were used in order to construct the interactome of the Accepted 9 February 2019 AQP gene family and to determine which genes members are differentially expressed in PDAC as Available online 11 February 2019 compared to controls. The same techniques were used in order to evaluate the potential prognostic role of the differentially expressed genes. Keywords: PDAC Results: Transcriptome microarray data of four GEO datasets were incorporated, including 142 primary Aquaporin tumor samples and 104 normal pancreatic tissue samples. -

A Minimum-Labeling Approach for Reconstructing Protein Networks Across Multiple Conditions
A Minimum-Labeling Approach for Reconstructing Protein Networks across Multiple Conditions Arnon Mazza1, Irit Gat-Viks2, Hesso Farhan3, and Roded Sharan1 1 Blavatnik School of Computer Science, Tel Aviv University, Tel Aviv 69978, Israel. Email: [email protected]. 2 Dept. of Cell Research and Immunology, Tel Aviv University, Tel Aviv 69978, Israel. 3 Biotechnology Institute Thurgau, University of Konstanz, Unterseestrasse 47, CH-8280 Kreuzlingen, Switzerland. Abstract. The sheer amounts of biological data that are generated in recent years have driven the development of network analysis tools to fa- cilitate the interpretation and representation of these data. A fundamen- tal challenge in this domain is the reconstruction of a protein-protein sub- network that underlies a process of interest from a genome-wide screen of associated genes. Despite intense work in this area, current algorith- mic approaches are largely limited to analyzing a single screen and are, thus, unable to account for information on condition-specific genes, or reveal the dynamics (over time or condition) of the process in question. Here we propose a novel formulation for network reconstruction from multiple-condition data and devise an efficient integer program solution for it. We apply our algorithm to analyze the response to influenza in- fection in humans over time as well as to analyze a pair of ER export related screens in humans. By comparing to an extant, single-condition tool we demonstrate the power of our new approach in integrating data from multiple conditions in a compact and coherent manner, capturing the dynamics of the underlying processes. 1 Introduction With the increasing availability of high-throughput data, network biol- arXiv:1307.7803v1 [q-bio.QM] 30 Jul 2013 ogy has become the method of choice for filtering, interpreting and rep- resenting these data. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Conserved and Novel Properties of Clathrin-Mediated Endocytosis in Dictyostelium Discoideum" (2012)
Rockefeller University Digital Commons @ RU Student Theses and Dissertations 2012 Conserved and Novel Properties of Clathrin- Mediated Endocytosis in Dictyostelium Discoideum Laura Macro Follow this and additional works at: http://digitalcommons.rockefeller.edu/ student_theses_and_dissertations Part of the Life Sciences Commons Recommended Citation Macro, Laura, "Conserved and Novel Properties of Clathrin-Mediated Endocytosis in Dictyostelium Discoideum" (2012). Student Theses and Dissertations. Paper 163. This Thesis is brought to you for free and open access by Digital Commons @ RU. It has been accepted for inclusion in Student Theses and Dissertations by an authorized administrator of Digital Commons @ RU. For more information, please contact [email protected]. CONSERVED AND NOVEL PROPERTIES OF CLATHRIN- MEDIATED ENDOCYTOSIS IN DICTYOSTELIUM DISCOIDEUM A Thesis Presented to the Faculty of The Rockefeller University in Partial Fulfillment of the Requirements for the degree of Doctor of Philosophy by Laura Macro June 2012 © Copyright by Laura Macro 2012 CONSERVED AND NOVEL PROPERTIES OF CLATHRIN- MEDIATED ENDOCYTOSIS IN DICTYOSTELIUM DISCOIDEUM Laura Macro, Ph.D. The Rockefeller University 2012 The protein clathrin mediates one of the major pathways of endocytosis from the extracellular milieu and plasma membrane. Clathrin functions with a network of interacting accessory proteins, one of which is the adaptor complex AP-2, to co-ordinate vesicle formation. Disruption of genes involved in clathrin-mediated endocytosis causes embryonic lethality in multicellular animals suggesting that clathrin-mediated endocytosis is a fundamental cellular process. However, loss of clathrin-mediated endocytosis genes in single cell eukaryotes, such as S.cerevisiae (yeast), does not cause lethality, suggesting that clathrin may convey specific advantages for multicellularity. -

Supplementary Methods
Supplementary methods Human lung tissues and tissue microarray (TMA) All human tissues were obtained from the Lung Cancer Specialized Program of Research Excellence (SPORE) Tissue Bank at the M.D. Anderson Cancer Center (Houston, TX). A collection of 26 lung adenocarcinomas and 24 non-tumoral paired tissues were snap-frozen and preserved in liquid nitrogen for total RNA extraction. For each tissue sample, the percentage of malignant tissue was calculated and the cellular composition of specimens was determined by histological examination (I.I.W.) following Hematoxylin-Eosin (H&E) staining. All malignant samples retained contained more than 50% tumor cells. Specimens resected from NSCLC stages I-IV patients who had no prior chemotherapy or radiotherapy were used for TMA analysis by immunohistochemistry. Patients who had smoked at least 100 cigarettes in their lifetime were defined as smokers. Samples were fixed in formalin, embedded in paraffin, stained with H&E, and reviewed by an experienced pathologist (I.I.W.). The 413 tissue specimens collected from 283 patients included 62 normal bronchial epithelia, 61 bronchial hyperplasias (Hyp), 15 squamous metaplasias (SqM), 9 squamous dysplasias (Dys), 26 carcinomas in situ (CIS), as well as 98 squamous cell carcinomas (SCC) and 141 adenocarcinomas. Normal bronchial epithelia, hyperplasia, squamous metaplasia, dysplasia, CIS, and SCC were considered to represent different steps in the development of SCCs. All tumors and lesions were classified according to the World Health Organization (WHO) 2004 criteria. The TMAs were prepared with a manual tissue arrayer (Advanced Tissue Arrayer ATA100, Chemicon International, Temecula, CA) using 1-mm-diameter cores in triplicate for tumors and 1.5 to 2-mm cores for normal epithelial and premalignant lesions. -

Downloaded from the Projected by No Less Than Six of the Servers Mentioned Above
ORIGINAL RESEARCH published: 11 June 2021 doi: 10.3389/fmolb.2021.687576 A Comprehensive Analysis of the Downregulation of miRNA-1827 and Its Prognostic Significance by Targeting SPTBN2 and BCL2L1 in Ovarian Cancer Penghui Feng 1†, Zhitong Ge 2†, Zaixin Guo 1, Lin Lin 1,3 and Qi Yu 1* 1Department of Obstetrics and Gynecology, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China, 2Department of Ultrasound, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China, 3Department of Obstetrics and Gynecology, The Maternal and Child Health Hospital of Guangxi Zhuang Autonomous Region, Beijing, China Background: Previous studies demonstrated that miRNA-1827 could repress various Edited by: cancers on proliferation, angiogenesis, and metastasis. However, little attention has been Hossain Shekhar, paid to its role in ovarian cancer as a novel biomarker or intervention target, especially its University of Dhaka, Bangladesh clinical significance and underlying regulatory network. Reviewed by: Udhaya Kumar S., Methods: A meta-analysis of six microarrays was adopted here to determine the Vellore Institute of Technology, India fi Sanjay Mishra, expression trend of miRNA-1827, and was further validated by gene expression pro le The Ohio State University, data and cellular experiments. We explored the functional annotations through enrichment United States analysis for the differentially expressed genes targeted by miRNA-1827. Subsequently, we *Correspondence: identified two hub genes, SPTBN2 and BCL2L1, based on interaction analysis using two Qi Yu [email protected] online archive tools, miRWALK (it consolidates the resources of 12 miRNA-focused †These authors have contributed servers) and Gene Expression Profiling Interactive Analysis (GEPIA). -
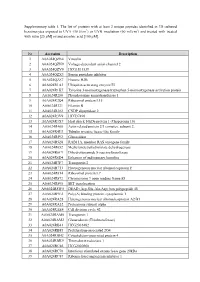
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31 -

Host Cell Factors Necessary for Influenza a Infection: Meta-Analysis of Genome Wide Studies
Host Cell Factors Necessary for Influenza A Infection: Meta-Analysis of Genome Wide Studies Juliana S. Capitanio and Richard W. Wozniak Department of Cell Biology, Faculty of Medicine and Dentistry, University of Alberta Abstract: The Influenza A virus belongs to the Orthomyxoviridae family. Influenza virus infection occurs yearly in all countries of the world. It usually kills between 250,000 and 500,000 people and causes severe illness in millions more. Over the last century alone we have seen 3 global influenza pandemics. The great human and financial cost of this disease has made it the second most studied virus today, behind HIV. Recently, several genome-wide RNA interference studies have focused on identifying host molecules that participate in Influen- za infection. We used nine of these studies for this meta-analysis. Even though the overlap among genes identified in multiple screens was small, network analysis indicates that similar protein complexes and biological functions of the host were present. As a result, several host gene complexes important for the Influenza virus life cycle were identified. The biological function and the relevance of each identified protein complex in the Influenza virus life cycle is further detailed in this paper. Background and PA bound to the viral genome via nucleoprotein (NP). The viral core is enveloped by a lipid membrane derived from Influenza virus the host cell. The viral protein M1 underlies the membrane and anchors NEP/NS2. Hemagglutinin (HA), neuraminidase Viruses are the simplest life form on earth. They parasite host (NA), and M2 proteins are inserted into the envelope, facing organisms and subvert the host cellular machinery for differ- the viral exterior. -

Produktinformation
Produktinformation Diagnostik & molekulare Diagnostik Laborgeräte & Service Zellkultur & Verbrauchsmaterial Forschungsprodukte & Biochemikalien Weitere Information auf den folgenden Seiten! See the following pages for more information! Lieferung & Zahlungsart Lieferung: frei Haus Bestellung auf Rechnung SZABO-SCANDIC Lieferung: € 10,- HandelsgmbH & Co KG Erstbestellung Vorauskassa Quellenstraße 110, A-1100 Wien T. +43(0)1 489 3961-0 Zuschläge F. +43(0)1 489 3961-7 [email protected] • Mindermengenzuschlag www.szabo-scandic.com • Trockeneiszuschlag • Gefahrgutzuschlag linkedin.com/company/szaboscandic • Expressversand facebook.com/szaboscandic SAN TA C RUZ BI OTEC HNOL OG Y, INC . COPG siRNA (h): sc-37254 BACKGROUND STORAGE AND RESUSPENSION Membrane and vesicular trafficking in the early secretory pathway are medi - Store lyophilized siRNA duplex at -20° C with desiccant. Stable for at least ated by non-clathrin COP (coat protein) I-coated vesicles. COPI-coated vesi cles one year from the date of shipment. Once resuspended, store at -20° C, mediate retrograde transport from the Golgi back to the ER and intra-Golgi avoid contact with RNAses and repeated freeze thaw cycles. transport. The cytosolic precursor of the COPI coat, the heptameric coatomer Resuspend lyophilized siRNA duplex in 330 µl of the RNAse-free water complex, is composed of two subcomplexes. The first consists of the COPB, pro vided. Resuspension of the siRNA duplex in 330 µl of RNAse-free water COPG, COPD and COPZ subunits (also known as -, -, - and -COP, respec - β γ δ ζ makes a 10 µM solution in a 10 µM Tris-HCl, pH 8.0, 20 mM NaCl, 1 mM tively), which are distantly homologous to AP Clathrin adaptor subunits. -

Supplemental Data 5-21-18
Supplemental Methods and Data Androgen receptor polyglutamine expansion drives age-dependent quality control defects and muscle dysfunction Samir R. Nath1,2,3, Zhigang Yu1, Theresa A. Gipson4, Gregory B. Marsh5, Eriko Yoshidome1, Diane M. Robins6, Sokol V. Todi5, David E. Housman4, Andrew P. Lieberman1 1 Department of Pathology, University of Michigan Medical School, Ann Arbor, MI 48109 2 Medical Scientist Training Program, University of Michigan Medical School, Ann Arbor, MI 48109 3 Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, MI 48109 4 Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, MA 02139 5 Department of Pharmacology, Wayne State University School of Medicine, Detroit, MI 48201 6 Department of Human Genetics, University of Michigan Medical School, Ann Arbor, MI 48109 Supplemental Methods qPCR For Drosophila samples, total RNA was extracted from adult fly heads using TRIzol. Fifteen heads were used per sample. Extracted RNA was treated with TURBO DNAse (Ambion) to eliminate contaminating DNA, and reverse transcription was carried out as indicated above. RNA levels were quantified using StepOnePlus Real-Time PCR System with Fast SYBR Green Master Mix (Applied Biosystems). rp49 was used as the internal control. Each round of qRT-PCR was conducted in technical triplicates. A total of three independent repeats was conducted. Fly histology: For histological preparation (75, 76), wings and proboscises of adult flies were removed and bodies were fixed overnight in 2% glutaraldehyde/2% paraformaldehyde in Tris- buffered saline with 0.1% Triton X-100, rotating at 4˚C. Fixed bodies were subsequently dehydrated by using a series of 30%, 50%, 75%, and 100% ethanol/propylene oxide.