Specific Association Between Alcohol Intake, High Grade of Di€Erentiation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Deregulated Gene Expression Pathways in Myelodysplastic Syndrome Hematopoietic Stem Cells
Leukemia (2010) 24, 756–764 & 2010 Macmillan Publishers Limited All rights reserved 0887-6924/10 $32.00 www.nature.com/leu ORIGINAL ARTICLE Deregulated gene expression pathways in myelodysplastic syndrome hematopoietic stem cells A Pellagatti1, M Cazzola2, A Giagounidis3, J Perry1, L Malcovati2, MG Della Porta2,MJa¨dersten4, S Killick5, A Verma6, CJ Norbury7, E Hellstro¨m-Lindberg4, JS Wainscoat1 and J Boultwood1 1LRF Molecular Haematology Unit, NDCLS, John Radcliffe Hospital, Oxford, UK; 2Department of Hematology Oncology, University of Pavia Medical School, Fondazione IRCCS Policlinico San Matteo, Pavia, Italy; 3Medizinische Klinik II, St Johannes Hospital, Duisburg, Germany; 4Division of Hematology, Department of Medicine, Karolinska Institutet, Stockholm, Sweden; 5Department of Haematology, Royal Bournemouth Hospital, Bournemouth, UK; 6Albert Einstein College of Medicine, Bronx, NY, USA and 7Sir William Dunn School of Pathology, University of Oxford, Oxford, UK To gain insight into the molecular pathogenesis of the the World Health Organization.6,7 Patients with refractory myelodysplastic syndromes (MDS), we performed global gene anemia (RA) with or without ringed sideroblasts, according to expression profiling and pathway analysis on the hemato- poietic stem cells (HSC) of 183 MDS patients as compared with the the French–American–British classification, were subdivided HSC of 17 healthy controls. The most significantly deregulated based on the presence or absence of multilineage dysplasia. In pathways in MDS include interferon signaling, thrombopoietin addition, patients with RA with excess blasts (RAEB) were signaling and the Wnt pathways. Among the most signifi- subdivided into two categories, RAEB1 and RAEB2, based on the cantly deregulated gene pathways in early MDS are immuno- percentage of bone marrow blasts. -

Multiple Changes of Gene Expression and Function Reveal Genomic and Phenotypic Complexity in SLE-Like Disease
RESEARCH ARTICLE Multiple Changes of Gene Expression and Function Reveal Genomic and Phenotypic Complexity in SLE-like Disease Maria Wilbe1☯¤a, Sergey V. Kozyrev2☯, Fabiana H. G. Farias2, Hanna D. Bremer3, Anna Hedlund1, Gerli R. Pielberg2, Eija H. Seppälä4¤b, Ulla Gustafson1, Hannes Lohi4, Örjan Carlborg5, Göran Andersson1, Helene Hansson-Hamlin3‡*, Kerstin Lindblad- Toh2,6‡* 1 Department of Animal Breeding and Genetics, Swedish University of Agricultural Sciences (SLU), Uppsala, Sweden, 2 Science for Life Laboratory, Department of Medical Biochemistry and Microbiology, Uppsala University, Uppsala, Sweden, 3 Department of Clinical Sciences, Swedish University of Agricultural Sciences (SLU), Uppsala, Sweden, 4 Research Programs Unit, Molecular Neurology; Department of Veterinary Biosciences, University of Helsinki and Folkhälsan Research Center, Helsinki, Finland, 5 Department of Clinical Sciences, Division of Computational Genetics, Swedish University of Agricultural Sciences (SLU), Uppsala, Sweden, 6 Broad Institute, Cambridge, Cambridge, Massachusetts, United States of America OPEN ACCESS ☯ These authors contributed equally to this work. ¤a Current address: Science for Life Laboratory, Department of Immunology, Genetics and Pathology, Citation: Wilbe M, Kozyrev SV, Farias FHG, Bremer Uppsala University, Uppsala, Sweden HD, Hedlund A, Pielberg GR, et al. (2015) Multiple ¤b Current address: Blueprint Genetics Ltd, Helsinki, Finland Changes of Gene Expression and Function Reveal ‡ HHH and KLT share senior authorship. Genomic and Phenotypic Complexity in SLE-like * [email protected] (HHH); [email protected] (KLT) Disease. PLoS Genet 11(6): e1005248. doi:10.1371/ journal.pgen.1005248 Editor: Joshua M. Akey, University of Washington, Abstract UNITED STATES Received: September 4, 2014 The complexity of clinical manifestations commonly observed in autoimmune disorders poses a major challenge to genetic studies of such diseases. -

A Discovery Resource of Rare Copy Number Variations in Individuals with Autism Spectrum Disorder
INVESTIGATION A Discovery Resource of Rare Copy Number Variations in Individuals with Autism Spectrum Disorder Aparna Prasad,* Daniele Merico,* Bhooma Thiruvahindrapuram,* John Wei,* Anath C. Lionel,*,† Daisuke Sato,* Jessica Rickaby,* Chao Lu,* Peter Szatmari,‡ Wendy Roberts,§ Bridget A. Fernandez,** Christian R. Marshall,*,†† Eli Hatchwell,‡‡ Peggy S. Eis,‡‡ and Stephen W. Scherer*,†,††,1 *The Centre for Applied Genomics, Program in Genetics and Genome Biology, The Hospital for Sick Children, Toronto M5G 1L7, Canada, †Department of Molecular Genetics, University of Toronto, Toronto M5G 1L7, Canada, ‡Offord Centre for Child Studies, Department of Psychiatry and Behavioural Neurosciences, McMaster University, Hamilton L8P 3B6, § Canada, Autism Research Unit, The Hospital for Sick Children, Toronto M5G 1X8, Canada, **Disciplines of Genetics and Medicine, Memorial University of Newfoundland, St. John’s, Newfoundland A1B 3V6, Canada, ††McLaughlin Centre, University of Toronto, Toronto M5G 1L7, Canada, and ‡‡Population Diagnostics, Inc., Melville, New York 11747 ABSTRACT The identification of rare inherited and de novo copy number variations (CNVs) in human KEYWORDS subjects has proven a productive approach to highlight risk genes for autism spectrum disorder (ASD). A rare variants variety of microarrays are available to detect CNVs, including single-nucleotide polymorphism (SNP) arrays gene copy and comparative genomic hybridization (CGH) arrays. Here, we examine a cohort of 696 unrelated ASD number cases using a high-resolution one-million feature CGH microarray, the majority of which were previously chromosomal genotyped with SNP arrays. Our objective was to discover new CNVs in ASD cases that were not detected abnormalities by SNP microarray analysis and to delineate novel ASD risk loci via combined analysis of CGH and SNP array cytogenetics data sets on the ASD cohort and CGH data on an additional 1000 control samples. -

Application of Microrna Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease
Article Application of microRNA Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease Jennifer L. Major, Rushita A. Bagchi * and Julie Pires da Silva * Department of Medicine, Division of Cardiology, University of Colorado Anschutz Medical Campus, Aurora, CO 80045, USA; [email protected] * Correspondence: [email protected] (R.A.B.); [email protected] (J.P.d.S.) Supplementary Tables Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 www.mdpi.com/journal/mps Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 2 of 25 Table 1. List of all hsa-miRs identified by Human microRNA Disease Database (HMDD; v3.2) analysis. hsa-miRs were identified using the term “genetics” and “circulating” as input in HMDD. Targets CAD hsa-miR-1 Targets IR injury hsa-miR-423 Targets Obesity hsa-miR-499 hsa-miR-146a Circulating Obesity Genetics CAD hsa-miR-423 hsa-miR-146a Circulating CAD hsa-miR-149 hsa-miR-499 Circulating IR Injury hsa-miR-146a Circulating Obesity hsa-miR-122 Genetics Stroke Circulating CAD hsa-miR-122 Circulating Stroke hsa-miR-122 Genetics Obesity Circulating Stroke hsa-miR-26b hsa-miR-17 hsa-miR-223 Targets CAD hsa-miR-340 hsa-miR-34a hsa-miR-92a hsa-miR-126 Circulating Obesity Targets IR injury hsa-miR-21 hsa-miR-423 hsa-miR-126 hsa-miR-143 Targets Obesity hsa-miR-21 hsa-miR-223 hsa-miR-34a hsa-miR-17 Targets CAD hsa-miR-223 hsa-miR-92a hsa-miR-126 Targets IR injury hsa-miR-155 hsa-miR-21 Circulating CAD hsa-miR-126 hsa-miR-145 hsa-miR-21 Targets Obesity hsa-mir-223 hsa-mir-499 hsa-mir-574 Targets IR injury hsa-mir-21 Circulating IR injury Targets Obesity hsa-mir-21 Targets CAD hsa-mir-22 hsa-mir-133a Targets IR injury hsa-mir-155 hsa-mir-21 Circulating Stroke hsa-mir-145 hsa-mir-146b Targets Obesity hsa-mir-21 hsa-mir-29b Methods Protoc. -

35Th International Society for Animal Genetics Conference 7
35th INTERNATIONAL SOCIETY FOR ANIMAL GENETICS CONFERENCE 7. 23.16 – 7.27. 2016 Salt Lake City, Utah ABSTRACT BOOK https://www.asas.org/meetings/isag2016 INVITED SPEAKERS S0100 – S0124 https://www.asas.org/meetings/isag2016 epigenetic modifications, such as DNA methylation, and measuring different proteins and cellular metab- INVITED SPEAKERS: FUNCTIONAL olites. These advancements provide unprecedented ANNOTATION OF ANIMAL opportunities to uncover the genetic architecture GENOMES (FAANG) ASAS-ISAG underlying phenotypic variation. In this context, the JOINT SYMPOSIUM main challenge is to decipher the flow of biological information that lies between the genotypes and phe- notypes under study. In other words, the new challenge S0100 Important lessons from complex genomes. is to integrate multiple sources of molecular infor- T. R. Gingeras* (Cold Spring Harbor Laboratory, mation (i.e., multiple layers of omics data to reveal Functional Genomics, Cold Spring Harbor, NY) the causal biological networks that underlie complex traits). It is important to note that knowledge regarding The ~3 billion base pairs of the human DNA rep- causal relationships among genes and phenotypes can resent a storage devise encoding information for be used to predict the behavior of complex systems, as hundreds of thousands of processes that can go on well as optimize management practices and selection within and outside a human cell. This information is strategies. Here, we describe a multi-step procedure revealed in the RNAs that are composed of 12 billion for inferring causal gene-phenotype networks underly- nucleotides, considering the strandedness and allelic ing complex phenotypes integrating multi-omics data. content of each of the diploid copies of the genome. -

Human Social Genomics in the Multi-Ethnic Study of Atherosclerosis
Getting “Under the Skin”: Human Social Genomics in the Multi-Ethnic Study of Atherosclerosis by Kristen Monét Brown A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Epidemiological Science) in the University of Michigan 2017 Doctoral Committee: Professor Ana V. Diez-Roux, Co-Chair, Drexel University Professor Sharon R. Kardia, Co-Chair Professor Bhramar Mukherjee Assistant Professor Belinda Needham Assistant Professor Jennifer A. Smith © Kristen Monét Brown, 2017 [email protected] ORCID iD: 0000-0002-9955-0568 Dedication I dedicate this dissertation to my grandmother, Gertrude Delores Hampton. Nanny, no one wanted to see me become “Dr. Brown” more than you. I know that you are standing over the bannister of heaven smiling and beaming with pride. I love you more than my words could ever fully express. ii Acknowledgements First, I give honor to God, who is the head of my life. Truly, without Him, none of this would be possible. Countless times throughout this doctoral journey I have relied my favorite scripture, “And we know that all things work together for good, to them that love God, to them who are called according to His purpose (Romans 8:28).” Secondly, I acknowledge my parents, James and Marilyn Brown. From an early age, you two instilled in me the value of education and have been my biggest cheerleaders throughout my entire life. I thank you for your unconditional love, encouragement, sacrifices, and support. I would not be here today without you. I truly thank God that out of the all of the people in the world that He could have chosen to be my parents, that He chose the two of you. -

Robles JTO Supplemental Digital Content 1
Supplementary Materials An Integrated Prognostic Classifier for Stage I Lung Adenocarcinoma based on mRNA, microRNA and DNA Methylation Biomarkers Ana I. Robles1, Eri Arai2, Ewy A. Mathé1, Hirokazu Okayama1, Aaron Schetter1, Derek Brown1, David Petersen3, Elise D. Bowman1, Rintaro Noro1, Judith A. Welsh1, Daniel C. Edelman3, Holly S. Stevenson3, Yonghong Wang3, Naoto Tsuchiya4, Takashi Kohno4, Vidar Skaug5, Steen Mollerup5, Aage Haugen5, Paul S. Meltzer3, Jun Yokota6, Yae Kanai2 and Curtis C. Harris1 Affiliations: 1Laboratory of Human Carcinogenesis, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 2Division of Molecular Pathology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 3Genetics Branch, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 4Division of Genome Biology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 5Department of Chemical and Biological Working Environment, National Institute of Occupational Health, NO-0033 Oslo, Norway. 6Genomics and Epigenomics of Cancer Prediction Program, Institute of Predictive and Personalized Medicine of Cancer (IMPPC), 08916 Badalona (Barcelona), Spain. List of Supplementary Materials Supplementary Materials and Methods Fig. S1. Hierarchical clustering of based on CpG sites differentially-methylated in Stage I ADC compared to non-tumor adjacent tissues. Fig. S2. Confirmatory pyrosequencing analysis of DNA methylation at the HOXA9 locus in Stage I ADC from a subset of the NCI microarray cohort. 1 Fig. S3. Methylation Beta-values for HOXA9 probe cg26521404 in Stage I ADC samples from Japan. Fig. S4. Kaplan-Meier analysis of HOXA9 promoter methylation in a published cohort of Stage I lung ADC (J Clin Oncol 2013;31(32):4140-7). Fig. S5. Kaplan-Meier analysis of a combined prognostic biomarker in Stage I lung ADC. -

The Pdx1 Bound Swi/Snf Chromatin Remodeling Complex Regulates Pancreatic Progenitor Cell Proliferation and Mature Islet Β Cell
Page 1 of 125 Diabetes The Pdx1 bound Swi/Snf chromatin remodeling complex regulates pancreatic progenitor cell proliferation and mature islet β cell function Jason M. Spaeth1,2, Jin-Hua Liu1, Daniel Peters3, Min Guo1, Anna B. Osipovich1, Fardin Mohammadi3, Nilotpal Roy4, Anil Bhushan4, Mark A. Magnuson1, Matthias Hebrok4, Christopher V. E. Wright3, Roland Stein1,5 1 Department of Molecular Physiology and Biophysics, Vanderbilt University, Nashville, TN 2 Present address: Department of Pediatrics, Indiana University School of Medicine, Indianapolis, IN 3 Department of Cell and Developmental Biology, Vanderbilt University, Nashville, TN 4 Diabetes Center, Department of Medicine, UCSF, San Francisco, California 5 Corresponding author: [email protected]; (615)322-7026 1 Diabetes Publish Ahead of Print, published online June 14, 2019 Diabetes Page 2 of 125 Abstract Transcription factors positively and/or negatively impact gene expression by recruiting coregulatory factors, which interact through protein-protein binding. Here we demonstrate that mouse pancreas size and islet β cell function are controlled by the ATP-dependent Swi/Snf chromatin remodeling coregulatory complex that physically associates with Pdx1, a diabetes- linked transcription factor essential to pancreatic morphogenesis and adult islet-cell function and maintenance. Early embryonic deletion of just the Swi/Snf Brg1 ATPase subunit reduced multipotent pancreatic progenitor cell proliferation and resulted in pancreas hypoplasia. In contrast, removal of both Swi/Snf ATPase subunits, Brg1 and Brm, was necessary to compromise adult islet β cell activity, which included whole animal glucose intolerance, hyperglycemia and impaired insulin secretion. Notably, lineage-tracing analysis revealed Swi/Snf-deficient β cells lost the ability to produce the mRNAs for insulin and other key metabolic genes without effecting the expression of many essential islet-enriched transcription factors. -
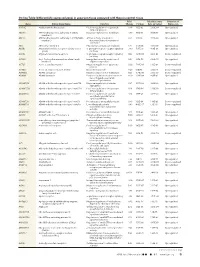
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

HERC3 (NM 014606) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RG214034 HERC3 (NM_014606) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: HERC3 (NM_014606) Human Tagged ORF Clone Tag: TurboGFP Symbol: HERC3 Vector: pCMV6-AC-GFP (PS100010) E. coli Selection: Ampicillin (100 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RG214034 representing NM_014606 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGTTATGTTGGGGATATTGGTCTCTGGGCCAACCTGGTATCAGCACCAACCTGCAGGGAATTGTGGCTG AGCCCCAGGTGTGTGGGTTCATATCTGACAGAAGTGTCAAGGAAGTGGCCTGTGGGGGAAACCACTCTGT GTTCCTGCTGGAAGATGGGGAAGTTTACACATGTGGTTTGAACACCAAGGGGCAACTGGGCCATGAGAGG GAAGGAAACAAGCCAGAACAAATTGGAGCTCTGGCAGATCAGCATATCATTCATGTGGCATGTGGCGAGT CCCACAGTCTGGCCCTCAGTGACCGAGGCCAGCTGTTTTCTTGGGGTGCAGGGAGTGATGGTCAGCTAGG ACTCATGACTACTGAGGATTCTGTGGCAGTGCCCAGGTTAATACAAAAGCTGAACCAGCAAACAATATTA CAAGTTTCCTGTGGCAACTGGCATTGCTTGGCTCTTGCGGCTGATGGCCAGTTCTTCACCTGGGGAAAGA ACAGCCATGGGCAGCTTGGCTTAGGGAAGGAGTTCCCCTCCCAAGCCAGCCCACAGAGGGTGAGGTCCCT GGAGGGGATCCCACTGGCTCAGGTGGCTGCCGGAGGGGCTCACAGCTTTGCCCTGTCTCTCTCAGGAGCT GTTTTTGGCTGGGGGATGAATAATGCCGGGCAGCTAGGGCTCAGTGATGAAAAAGATCGAGAATCTCCAT GCCATGTAAAACTCTTACGCACGCAAAAAGTTGTCTATATTAGTTGTGGAGAAGAACACACAGCAGTTCT CACAAAGAGTGGAGGTGTGTTTACCTTTGGCGCTGGTTCCTGTGGGCAACTTGGACACGACTCCATGAAT GATGAGGTTAACCCTAGAAGAGTTCTAGAGCTGATGGGTAGTGAAGTAACTCAAATTGCTTGTGGCAGAC -

HERC5, a HECT E3 Ubiquitin Ligase Tightly Regulated in LPS Activated Endothelial Cells
Research Article 4749 HERC5, a HECT E3 ubiquitin ligase tightly regulated in LPS activated endothelial cells Renate Kroismayr, Ulrike Baranyi*, Christian Stehlik‡, Andrea Dorfleutner‡, Bernd R. Binder§ and Joachim Lipp Department of Vascular Biology and Thrombosis Research, Medical University of Vienna and BMT, Bio-Molecular Therapeutics, Schwarzspanierstrasse 17, 1090 Vienna, Austria *Present address: Department of Surgery, Vienna General Hospital, Medical University of Vienna, Währinger Güttel 18-20, 1090 Vienna, Austria ‡Present address: Mary Babb Randolph Cancer Center and Department of Microbiology, Immunology, and Cell Biology, West Virginia University School of Medicine, PO Box 9300, Morgantown, WV 26506, USA §Author for correspondence (e-mail: [email protected]) Accepted 27 May 2004 Journal of Cell Science 117, 4749-4756 Published by The Company of Biologists 2004 doi:10.1242/jcs.01338 Summary By differential screening we isolated genes upregulated mRNA levels suggests that the initial drop in HERC5 in inflammatory cytokine-stimulated human skin protein is balanced by increased protein synthesis due to microvascular endothelial cells. One of these cDNAs upregulation of HERC5 mRNA. This leads to recovery of encoded RCC1 (regulator of chromosome condensation 1)- HERC5 protein levels within 12 hours of LPS stimulation like repeats and a HECT (homologous to E6-AP C- and points at a tight control of HERC5 protein. To analyze terminus) domain, representing a member of the HERC functional activity of this putative member of the ubiquitin- (HECT and RCC1 domain protein) family of ubiquitin conjugating pathway we performed in vitro assays with ligases. The mRNA level of this member, HERC5, is different ubiquitin-conjugating enzymes. -

Molecular Sciences High-Resolution Chromosome Ideogram Representation of Currently Recognized Genes for Autism Spectrum Disorder
Int. J. Mol. Sci. 2015, 16, 6464-6495; doi:10.3390/ijms16036464 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Article High-Resolution Chromosome Ideogram Representation of Currently Recognized Genes for Autism Spectrum Disorders Merlin G. Butler *, Syed K. Rafi † and Ann M. Manzardo † Departments of Psychiatry & Behavioral Sciences and Pediatrics, University of Kansas Medical Center, Kansas City, KS 66160, USA; E-Mails: [email protected] (S.K.R.); [email protected] (A.M.M.) † These authors contributed to this work equally. * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +1-913-588-1873; Fax: +1-913-588-1305. Academic Editor: William Chi-shing Cho Received: 23 January 2015 / Accepted: 16 March 2015 / Published: 20 March 2015 Abstract: Recently, autism-related research has focused on the identification of various genes and disturbed pathways causing the genetically heterogeneous group of autism spectrum disorders (ASD). The list of autism-related genes has significantly increased due to better awareness with advances in genetic technology and expanding searchable genomic databases. We compiled a master list of known and clinically relevant autism spectrum disorder genes identified with supporting evidence from peer-reviewed medical literature sources by searching key words related to autism and genetics and from authoritative autism-related public access websites, such as the Simons Foundation Autism Research Institute autism genomic database dedicated to gene discovery and characterization. Our list consists of 792 genes arranged in alphabetical order in tabular form with gene symbols placed on high-resolution human chromosome ideograms, thereby enabling clinical and laboratory geneticists and genetic counsellors to access convenient visual images of the location and distribution of ASD genes.